
The c4 antisense RNA is a non-coding RNA used by certain phages that infect bacteria. It was initially identified in the P1 and P7 phages of E. coli. The identification of c4 antisense RNAs solved the mystery of the mechanism for regulation of the ant gene, which is an anti-repressor.
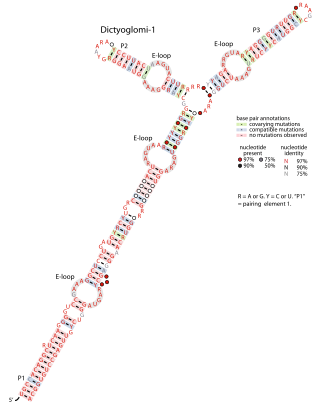
The Dictyoglomi-1 RNA motif is a conserved RNA structure that was discovered via bioinformatics. Only four instances of the RNA were detected, and all are in the bacterial phylum Dictyoglomota, whose members have not been extensively studied. The RNA might have a cis-regulatory role, but the evidence is ambiguous. Because of the few instances of Dictyoglomi-1 RNAs known, it is also unknown whether the RNA structure might extend further in the 5′ or 3′ direction, or in both directions.

The Downstream-peptide motif refers to a conserved RNA structure identified by bioinformatics in the cyanobacterial genera Synechococcus and Prochlorococcus and one phage that infects such bacteria. It was also detected in marine samples of DNA from uncultivated bacteria, which are presumably other species of cyanobacteria.

The glutamine riboswitch is a conserved RNA structure that was predicted by bioinformatics. It is present in a variety of lineages of cyanobacteria, as well as some phages that infect cyanobacteria. It is also found in DNA extracted from uncultivated bacteria living in the ocean that are presumably species of cyanobacteria.
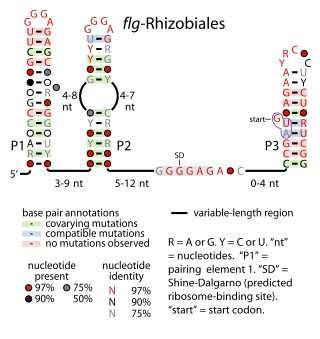
The flg-Rhizobiales RNA motif is an RNA structure that is conserved in certain bacteria. All known flg-Rhizobiales RNAs are located in the presumptive 5' untranslated regions of operons that contain genes whose functions relate to the creation of flagellar basal bodies. The flg-Rhizobiales RNAs are restricted to the Hyphomicrobiales, an order of alphaproteobacteria, although only some Rhizobiales bacterial are predicted to use flg-Rhizobiales RNAs. The exact function of these RNAs is unknown, although it is hypothesized that they have a cis-regulatory function in controlling expression of the downstream operons.

The Gut-1 RNA motif is a conserved RNA structure identified by bioinformatics. These RNAs are present in environmental sequences, and as of 2010 are not known to be present in any species that has been grown under laboratory conditions. Gut-1 RNA is exclusively found in DNA from uncultivated bacteria present in samples from the human gut.

The gyrA RNA motif is a conserved RNA structure identified by bioinformatics. The RNAs are present in multiple species of bacteria within the order Pseudomonadales. This order contains the genus Pseudomonas, which includes the opportunistic human pathogen Pseudomonas aeruginosa and Pseudomonas syringae, a plant pathogen.

The Lacto-rpoB RNA motif is a conserved RNA structure identified by bioinformatics. It has been detected only in lactic acid bacteria, and is always located in the presumed 5' untranslated regions of rpoB genes. These genes encode a subunit of RNA polymerase, and it is hypothesized that Lacto-rpoB RNA participate in the regulation of these genes.
The wcaG RNA motif is an RNA structure conserved in some bacteria that was detected by bioinformatics. wcaG RNAs are found in certain phages that infect cyanobacteria. Most known wcaG RNAs were found in sequences of DNA extracted from uncultivated marine bacteria. wcaG RNAs might function as cis-regulatory elements, in view of their consistent location in the possible 5' untranslated regions of genes. It was suggested the wcaG RNAs might further function as riboswitches.

The msiK RNA motif describes a conserved RNA structure discovered using bioinformatics. The RNA is always found in the presumed 5' untranslated regions of genes annotated as msiK, and is therefore hypothesized to be an RNA-based cis-regulatory element that regulates these genes.

PhotoRC RNA motifs refer to conserved RNA structures that are associated with genes acting in the photosynthetic reaction centre of photosynthetic bacteria. Two such RNA classes were identified and called the PhotoRC-I and PhotoRC-II motifs. PhotoRC-I RNAs were detected in the genomes of some cyanobacteria. Although no PhotoRC-II RNA has been detected in cyanobacteria, one is found in the genome of a purified phage that infects cyanobacteria. Both PhotoRC-I and PhotoRC-II RNAs are present in sequences derived from DNA that was extracted from uncultivated marine bacteria.
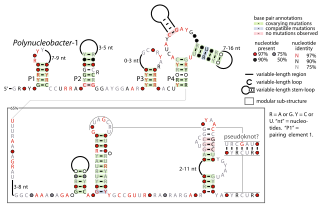
The Polynucleobacter-1 RNA motif is a conserved RNA structure that was identified by bioinformatics. The RNA structure is predominantly located in genome sequences derived from DNA extracted from uncultivated marine samples. However it was also predicted in the genome of Polynucleobacter species QLW-P1DMWA-1, a kind of betaproteobacteria. The RNAs are often located near to a conserved gene that might be homologous to a gene found in a phage that infects cyanobacteria. However, it is unknown if the RNA is used by phages.
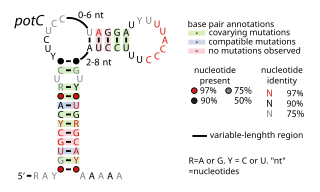
The potC RNA motif is a conserved RNA structure discovered using bioinformatics. The RNA is detected only in genome sequences derived from DNA that was extracted from uncultivated marine bacteria. Thus, this RNA is present in environmental samples, but not yet found in any cultivated organism. potC RNAs are located in the presumed 5' untranslated regions of genes predicted to encode either membrane transport proteins or peroxiredoxins. Therefore, it was hypothesized that potC RNAs are cis-regulatory elements, but their detailed function is unknown.

The psaA RNA motif describes a class of RNAs with a common secondary structure. psaA RNAs are exclusively found in locations that presumably correspond to the 5' untranslated regions of operons formed of psaA and psaB genes. For this reason, it was hypothesized that psaA RNAs function as cis-regulatory elements of these genes. The psaAB genes encode proteins that form subunits in the photosystem I structure used for photosynthesis. psaA RNAs have been detected only in cyanobacteria, which is consistent with their association with photosynthesis.

The Pseudomon-Rho RNA motif refers to a conserved RNA structure that was discovered using bioinformatics. The RNAs that conform to this motif are found in species within the genus Pseudomonas, as well as the related Azotobacter vinelandii. They are consistently located in what could be the 5' untranslated regions of genes that encode the Rho factor protein, and this arrangement in bacteria suggested that Pseudomon-Rho RNAs might be cis-regulatory elements that regulate concentrations of the Rho protein.

The rmf RNA motif is a conserved RNA structure that was originally detected using bioinformatics. rmf RNAs are consistently foundwithin species classified into the genus Pseudomonas, and is located potentially in the 5′ untranslated regions of rmf genes. These genes encodes the ribosome modulation factor protein, which affects the translation of genes by modifying ribosome structure in response to stress such as starvation. This ribosome modulation is a part of the stringent response in bacteria. The likely biological role of rmf RNAs is ambiguous. Since the RNA could be in the 5′ UTRs of protein-coding genes, it was hypothesized that it functions as a cis-regulatory element. This hypothesis is bolstered by the observation that ribosome modulation factor binds ribosomal RNA, and many cis-regulatory RNAs called ribosomal protein leaders participate in a feedback regulation mechanism by binding to proteins that normally bind to ribosomal RNA. However, since rmf RNAs are not very close to the rmf genes, they might function as non-coding RNAs.
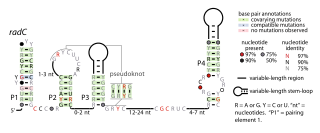
The radC RNA motif is a conserved RNA structure identified by bioinformatics. The radC RNA motif is found in certain bacteria where it is consistent located in the presumed 5' untranslated regions of genes whose encoded proteins bind DNA are interact with other proteins that bind DNA. These proteins include integrases, methyltransferases that might methylate DNA, proteins that inhibit restriction enzymes and radC genes. Although radC genes were thought to encode DNA repair proteins, this conclusion was based on mutation data that was later shown to affect a different gene. However, it is still possible that radC genes play some DNA-related role. No radC RNAs have been detected in any purified phage whose sequence was available as of 2010, although integrases are often used by phages.

The Termite-flg RNA motif is a conserved RNA structure identified by bioinformatics. Genomic sequences corresponding to Termite-flg RNAs have been identified only in uncultivated bacteria present in the termite hindgut. As of 2010 it has not been identified in the DNA of any cultivated species, and is thus an example of RNAs present in environmental samples.

The Whalefall-1 RNA motif refers to a conserved RNA structure that was discovered using bioinformatics. Structurally, the motif consists of two stem-loops, the second of which is often terminated by a CUUG tetraloop, which is an energetically favorable RNA sequence. Whalefall-1 RNAs are found only in DNA extracted from uncultivated bacteria found on whale fall, i.e., a whale carcass. As of 2010, Whalefall-1 RNAs have not been detected in any known, cultivated species of bacteria, and are thus one of several RNAs present in environmental samples.
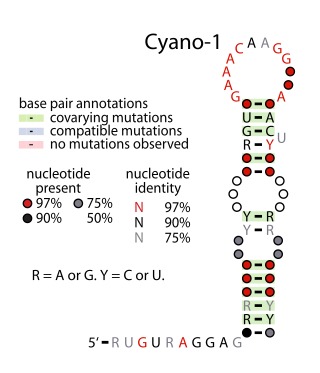
Yfr2 is a family of non-coding RNAs. Members of the Yrf2 family have been identified in almost all studied species of cyanobacteria. The family was identified through a bioinformatics screen of published cyanobacterial genomes, having previously been grouped in a family of Yfr2–5.



















