Related Research Articles

Micro ribonucleic acid are small, single-stranded, non-coding RNA molecules containing 21–23 nucleotides. Found in plants, animals, and even some viruses, miRNAs are involved in RNA silencing and post-transcriptional regulation of gene expression. miRNAs base-pair to complementary sequences in messenger RNA (mRNA) molecules, then silence said mRNA molecules by one or more of the following processes:

The miR-8 microRNA precursor, is a short non-coding RNA gene involved in gene regulation. miR-8 in Drosophila melanogaster is expressed from the 3' arm of related precursor hairpins, along with miR-200, miR-236, miR-429 and human and mouse homolog miR-141. Members of this precursor family have now been predicted or experimentally confirmed in a wide range of species. The bounds of the precursors are predicted based on conservation and base pairing and are not generally known.

The miR-9 microRNA, is a short non-coding RNA gene involved in gene regulation. The mature ~21nt miRNAs are processed from hairpin precursor sequences by the Dicer enzyme. The dominant mature miRNA sequence is processed from the 5' arm of the mir-9 precursor, and from the 3' arm of the mir-79 precursor. The mature products are thought to have regulatory roles through complementarity to mRNA. In vertebrates, miR-9 is highly expressed in the brain, and is suggested to regulate neuronal differentiation. A number of specific targets of miR-9 have been proposed, including the transcription factor REST and its partner CoREST.

The mir-10 microRNA precursor is a short non-coding RNA gene involved in gene regulation. It is part of an RNA gene family which contains mir-10, mir-51, mir-57, mir-99 and mir-100. mir-10, mir-99 and mir-100 have now been predicted or experimentally confirmed in a wide range of species. miR-51 and miR-57 have currently only been identified in the nematode Caenorhabditis elegans.

The mir-2 microRNA family includes the microRNA genes mir-2 and mir-13. Mir-2 is widespread in invertebrates, and it is the largest family of microRNAs in the model species Drosophila melanogaster. MicroRNAs from this family are produced from the 3' arm of the precursor hairpin. Leaman et al. showed that the miR-2 family regulates cell survival by translational repression of proapoptotic factors. Based on computational prediction of targets, a role in neural development and maintenance has been suggested.

The mir-6 microRNA precursor is a precursor microRNA specific to Drosophila species. In Drosophila melanogaster there are three mir-6 paralogs called dme-mir-6-1, dme-mir-6-2, dme-mir-6-3, which are clustered together in the genome. The extents of these hairpin precursors are estimated based on hairpin prediction. Each precursor is generated following the cleavage of a longer primary transcript in the nucleus, and is exported in the cytoplasm. In the cytoplasm, precursors are further processed by the enzyme Dicer, generating ~22 nucleotide products from each arm of the hairpin. The products generated from the 3' arm of each mir-6 precursor have identical sequences. Both 5' and 3' mature products are experimentally validated. Experimental data suggests that the mature products of mir-6 hairpins are expressed in the early embryo of Drosophila and target apoptotic genes such as hid, grim and rpr.
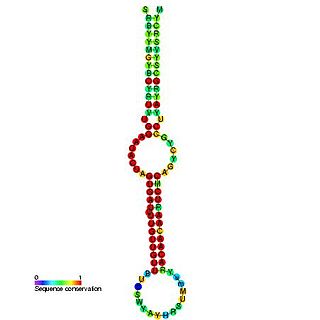
This family represents the microRNA (miRNA) precursor mir-7. This miRNA has been predicted or experimentally confirmed in a wide range of species. miRNAs are transcribed as ~70 nucleotide precursors and subsequently processed by the Dicer enzyme to give a ~22 nucleotide product. In this case the mature sequence comes from the 5' arm of the precursor. The extents of the hairpin precursors are not generally known and are estimated based on hairpin prediction. The involvement of Dicer in miRNA processing suggests a relationship with the phenomenon of RNA interference.
Post-transcriptional regulation is the control of gene expression at the RNA level. It occurs once the RNA polymerase has been attached to the gene's promoter and is synthesizing the nucleotide sequence. Therefore, as the name indicates, it occurs between the transcription phase and the translation phase of gene expression. These controls are critical for the regulation of many genes across human tissues. It also plays a big role in cell physiology, being implicated in pathologies such as cancer and neurodegenerative diseases.

In molecular biology mir-126 is a short non-coding RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several pre- and post-transcription mechanisms.

In molecular biology, miR-184 microRNA is a short non-coding RNA molecule. MicroRNAs (miRNAs) function as posttranscriptional regulators of expression levels of other genes by several mechanisms. Several targets for miR-184 have been described, including that of mediators of neurological development, apoptosis and it has been suggested that miR-184 plays an essential role in development.

In molecular biology, mir-433 is a short non-coding RNA molecule. MicroRNAs (miRNAs) function as posttranscriptional regulators of expression levels of other genes by several mechanisms. They play roles in development, metabolism and carcinogenesis.
In molecular biology mir-346 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.
In molecular biology mir-350 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.
In molecular biology mir-11 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms. There is an evidence to suggest that miR-11 plays a role in apoptosis.
In molecular biology mir-14 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.
In molecular biology mir-202 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms. The pre-miR-202 in the mouse genome is located fully within an exon, whereas in human it lies across a splice junction. This implies that human miR-202 is exposed to a negative regulation by splicing, whereas murine miR-202 is not.
In molecular biology mir-278 microRNA is a short RNA molecule belonging to a class of molecules referred to as microRNAs. These function to regulate the expression levels of other genes by several mechanisms, primarily binding to their target at its 3'UTR.
mir-279 is a short RNA molecule found in Drosophila melanogaster that belongs to a class of molecules known as microRNAs. microRNAs are ~22nt-long non-coding RNAs that post-transcriptionally regulate the expression of genes, often by binding to the 3' untranslated region of mRNA, targeting the transcript for degradation. miR-279 has diverse tissue-specific functions in the fly, influencing developmental processes related to neurogenesis and oogenesis, as well as behavioral processes related to circadian rhythms. The varied roles of mir-279, both in the developing and adult fly, highlight the utility of microRNAs in regulating unique biological processes.
mir-615 microRNA is a short non-coding RNA molecule belonging both to the family of microRNAs and to that of small interfering RNAs (siRNAs). MicroRNAs function to regulate the expression levels of other genes by several mechanisms, whilst siRNAs are involved primarily with the RNA interference (RNAi) pathway. siRNAs have been linked through some members to the regulation of cancer cell growth, specifically in prostate adenocarcinoma.

MiR-212 is a short non-coding RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms, generally reducing protein levels through the cleavage of mRNAs or the repression of their translation. Several targets for miR-132 have been described, including mediators of neurological development, synaptic transmission, inflammation and angiogenesis.
References
- ↑ Xiong H, Qian J, He T, Li F (January 2009). "Independent transcription of miR-281 in the intron of ODA in Drosophila melanogaster". Biochemical and Biophysical Research Communications. 378 (4): 883–9. doi:10.1016/j.bbrc.2008.12.010. PMID 19073139.