
In molecular biology, the RNase E 5′ UTR element is a cis-acting element located in the 5′ UTR of ribonuclease (RNase) E messenger RNA (mRNA).

The Bacteroid-trp RNA motif is a conserved RNA element detected by bioinformatics. It is found in the phylum Bacteroidota in the apparent 5' untranslated regions of genes that encode enzymes used in the synthesis of the amino acid tryptophan. A short open reading frame is found within the motif that encodes at least two tryptophan codons. Similar motifs have been identified regulating tryptophan genes in Pseudomonadota, but not in Bacteroidota. However, the Bacteroid-trp RNA motif likely operates via the same mechanism of attenuation.

The Bacteroides-1 RNA motif is a conserved RNA structure identified in bacteria within the genus Bacteroides. The RNAs are typically found downstream of of genes that participate in the synthesis of exopolysaccharides of unknown types. It is possible that Bacteroides-1 RNAs regulate the upstream genes, but since this mode of regulation is unusual in bacteria, it is more likely that the structure functions as a non-coding RNA.
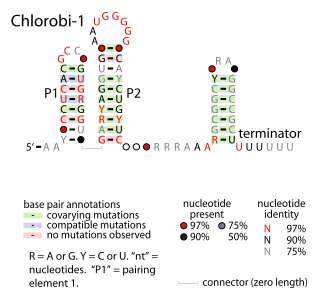
The Chlorobi-1 RNA motif is a conserved RNA secondary structure identified by bioinformatics. It is predicted to be used only by Chlorobiota, a phylum of bacteria. The motif consists of two stem-loops that are followed by an apparent rho-independent transcription terminator. The motif is presumed to function as an independently transcribed non-coding RNA.
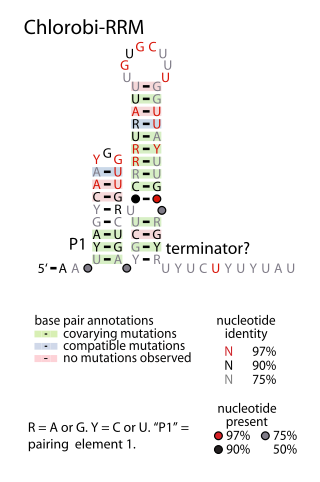
The Chlorobi-RRM RNA motif is a conserved RNA structure identified by bioinformatics. It is found within bacteria in the phylum Chlorobiota, and is exclusively detected in the presumed 5' untranslated regions of genes that encode putative RNA-binding proteins. Since many RNA-binding proteins regulate their own expression in a feedback mechanism by binding or acting up their 5' UTR, it was proposed that the Chlorobi-RRM is a component in an analogous feedback mechanism. Structurally, the motif consists of two stem-loops, the second of which might function as a rho-independent transcription terminator.

The Downstream-peptide motif refers to a conserved RNA structure identified by bioinformatics in the cyanobacterial genera Synechococcus and Prochlorococcus and one phage that infects such bacteria. It was also detected in marine samples of DNA from uncultivated bacteria, which are presumably other species of cyanobacteria.

The L17 downstream element RNA motif is a conserved RNA structure identified in bacteria by bioinformatics. All known L17 downstream elements were detected immediately downstream of genes encoding the L17 subunit of the ribosome, and therefore might be in the 3' untranslated regions of these genes. The element is found in a variety of lactic acid bacteria and in the genus Listeria.
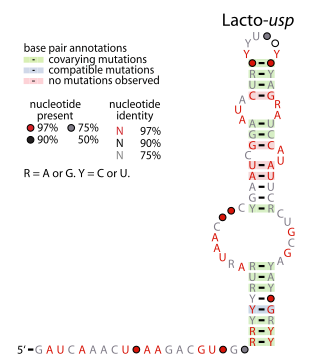
The Lacto-usp RNA motif is a conserved RNA structure identified in bacteria by bioinformatics. Lacto-usp RNAs are found exclusively in lactic acid bacteria, and exclusively in the possible 5′ untranslated regions of operons that contain a hypothetical gene and a usp gene. The usp gene encodes the universal stress protein. It was proposed that the Lacto-usp might correspond to the 6S RNA of the relevant species, because four of five of these species lack a predicted 6S RNA, and 6S RNAs commonly occur in 5′ UTRs of usp genes. However, given that the Lacto-usp RNA motif is much shorter than the standard 6S RNA structure, the function of Lacto-usp RNAs remains unclear.

The livK RNA motif describes a conserved RNA structure that was discovered using bioinformatics. The livK motif is detected only in the species Pseudomonas syringae. It is found in the potential 5' untranslated regions of livK genes and downstream livM and livH genes, as well as the 5' UTRs of amidase genes. The liv genes are predicted to be transporters of branched-chain amino acids, i.e., leucine, isoleucine or valine. The specific reaction catalyzed the amidase genes is not predicted.
The Lnt RNA motif refers to a conserved RNA structure found in certain bacteria. Specifically, Lnt RNAs are known only in species within the phylum Chlorobiota, and are located in the possible 5' untranslated regions of genes that are annotated as encoding apolipoprotein N-acyltransferase enzymes. There is some doubt as to whether the indicated motif is transcribed as RNA, or whether its reverse complement is transcribed. If the reverse complement is transcribed it would potentially in 5' UTRs of genes encoding bacteriochlorophyll A, and would be close to the start codon of those genes.

The Moco-II RNA motif is a conserved RNA structure identified by bioinformatics. However, only 8 examples of the RNA motif are known. The RNAs are potentially in the 5' untranslated regions of genes related to molybdenum cofactor (Moco), specifically a gene that encodes a molybdenum-binding domain and a nitrate reductase, which uses Moco as a cofactor. Thus the RNA might be involved in the regulation of genes based on Moco levels. Reliable predictions of Moco-II RNAs are restricted to deltaproteobacteria, but a Moco-II RNA might be present in a betaproteobacterial species. The Moco RNA motif is another RNA that is associated with Moco, and its complex secondary structure and genetic experiments have led to proposals that it is a riboswitch. However, the simpler structure of the Moco-II RNA motif is less typical of riboswitches. Moco-II RNAs are typically followed by a predicted rho-independent transcription terminator.

PhotoRC RNA motifs refer to conserved RNA structures that are associated with genes acting in the photosynthetic reaction centre of photosynthetic bacteria. Two such RNA classes were identified and called the PhotoRC-I and PhotoRC-II motifs. PhotoRC-I RNAs were detected in the genomes of some cyanobacteria. Although no PhotoRC-II RNA has been detected in cyanobacteria, one is found in the genome of a purified phage that infects cyanobacteria. Both PhotoRC-I and PhotoRC-II RNAs are present in sequences derived from DNA that was extracted from uncultivated marine bacteria.
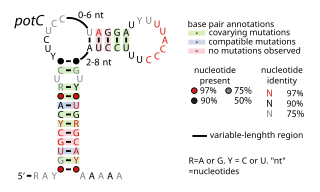
The potC RNA motif is a conserved RNA structure discovered using bioinformatics. The RNA is detected only in genome sequences derived from DNA that was extracted from uncultivated marine bacteria. Thus, this RNA is present in environmental samples, but not yet found in any cultivated organism. potC RNAs are located in the presumed 5' untranslated regions of genes predicted to encode either membrane transport proteins or peroxiredoxins. Therefore, it was hypothesized that potC RNAs are cis-regulatory elements, but their detailed function is unknown.
The Pseudomon-groES RNA motif is a conserved RNA structure identified in certain bacteria using bioinformatics. It is found in most species within the family Pseudomonadaceae, and is consistently located in the 5' untranslated regions of operons that contain groES genes. RNA transcripts of the groES genes in Pseudomonas aeruginosa where shown experimentally to be initiated at one of two start sites, from promoters called "P1" and "P2". The Pseudomon-groES RNA is in the 5' UTR of transcripts initiated from the P1 site, but is truncated in P2 transcripts. groES genes are involved in the cellular response to heat shock, but it is not thought that the Pseudomonas-groES RNA motif is involved in heat shock regulation. However, it is thought that the motif might regulate groES genes in response to other stimuli.

The rmf RNA motif is a conserved RNA structure that was originally detected using bioinformatics. rmf RNAs are consistently foundwithin species classified into the genus Pseudomonas, and is located potentially in the 5′ untranslated regions of rmf genes. These genes encodes the ribosome modulation factor protein, which affects the translation of genes by modifying ribosome structure in response to stress such as starvation. This ribosome modulation is a part of the stringent response in bacteria. The likely biological role of rmf RNAs is ambiguous. Since the RNA could be in the 5′ UTRs of protein-coding genes, it was hypothesized that it functions as a cis-regulatory element. This hypothesis is bolstered by the observation that ribosome modulation factor binds ribosomal RNA, and many cis-regulatory RNAs called ribosomal protein leaders participate in a feedback regulation mechanism by binding to proteins that normally bind to ribosomal RNA. However, since rmf RNAs are not very close to the rmf genes, they might function as non-coding RNAs.
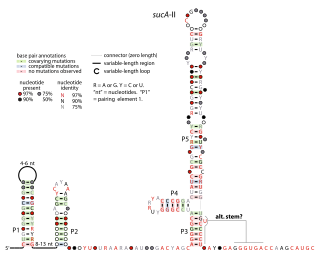
The sucA-II RNA motif is a conserved RNA structure identified by bioinformatics. It is consistently found in the presumed 5' untranslated regions of sucA genes, which encode Oxoglutarate dehydrogenase enzymes that participate in the citric acid cycle. Given this arrangement, sucA-II RNAs might regulate the downstream sucA gene. This genetic arrangement is similar to the previously reported sucA RNA motif. However, sucA-II RNAs are found only in bacteria classified within the genus Pseudomonas, whereas the previously reported motif is found only in betaproteobacteria.

The sucC RNA motif is a conserved RNA structure discovered using bioinformatics. sucC RNAs are found in the genus Pseudomonas, and are consistently found in possible 5' untranslated regions of sucC genes. These genes encode Succinyl coenzyme A synthetase, and are hypothesised to be regulated by the sucC RNAs. sucC genes participate in the citric acid cycle, and another gene involved in the citric acid cycle, sucA, is also predicted to be regulated by a conserved RNA structure.

The traJ-II RNA motif is a conserved RNA structure discovered in bacteria by using bioinformatics. traJ-II RNAs appear to be in the 5' untranslated regions of protein-coding genes called traJ, which functions in the process of bacterial conjugation. A previously identified motif known as TraJ 5' UTR is also found upstream of traJ genes functions as the target of FinP antisense RNAs, so it is possible that traJ-II RNAs play a similar role as targets of an antisense RNA. However, some sequence features within the traJ-II RNA motif suggest that the biological RNA might be transcribed from the reverse-complement strand. Thus is it unclear whether traJ-II function as cis-regulatory elements. traJ-II RNAs are found in a variety of Pseudomonadota.

The yjdF RNA motif is a conserved RNA structure identified using bioinformatics. Most yjdF RNAs are located in bacteria classified within the phylum Bacillota. A yjdF RNA is found in the presumed 5' untranslated region of the yjdF gene in Bacillus subtilis, and almost all yjdF RNAs are found in the 5' UTRs of homologs of this gene. The function of the yjdF gene is unknown, but the protein that it is predicted to encode is classified by the Pfam Database as DUF2992.

















