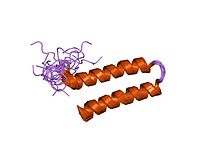
The alpha helix (α-helix) is a common motif in the secondary structure of proteins and is a right hand-helix conformation in which every backbone N−H group hydrogen bonds to the backbone C=O group of the amino acid located three or four residues earlier along the protein sequence.
DnaG is a bacterial DNA primase and is encoded by the dnaG gene. The enzyme DnaG, and any other DNA primase, synthesizes short strands of RNA known as oligonucleotides during DNA replication. These oligonucleotides are known as primers because they act as a starting point for DNA synthesis. DnaG catalyzes the synthesis of oligonucleotides that are 10 to 60 nucleotides long, however most of the oligonucleotides synthesized are 11 nucleotides. These RNA oligonucleotides serve as primers, or starting points, for DNA synthesis by bacterial DNA polymerase III. DnaG is important in bacterial DNA replication because DNA polymerase cannot initiate the synthesis of a DNA strand, but can only add nucleotides to a preexisting strand. DnaG synthesizes a single RNA primer at the origin of replication. This primer serves to prime leading strand DNA synthesis. For the other parental strand, the lagging strand, DnaG synthesizes an RNA primer every few kilobases (kb). These primers serve as substrates for the synthesis of Okazaki fragments.
A coiled coil is a structural motif in proteins in which 2–7 alpha-helices are coiled together like the strands of a rope. Many coiled coil-type proteins are involved in important biological functions, such as the regulation of gene expression — e.g., transcription factors. Notable examples are the oncoproteins c-Fos and c-jun, as well as the muscle protein tropomyosin.
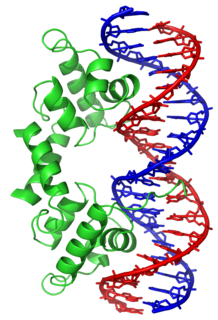
In proteins, the helix-turn-helix (HTH) is a major structural motif capable of binding DNA. Each monomer incorporates two α helices, joined by a short strand of amino acids, that bind to the major groove of DNA. The HTH motif occurs in many proteins that regulate gene expression. It should not be confused with the helix-loop-helix motif.

A leucine zipper is a common three-dimensional structural motif in proteins. They were first described by Landschulz and collaborators in 1988 when they found that an enhancer binding protein had a very characteristic 30-amino acid segment and the display of these amino acid sequences on an idealized alpha helix revealed a periodic repetition of leucine residues at every seventh position over a distance covering eight helical turns. The polypeptide segments containing these periodic arrays of leucine residues were proposed to exist in an alpha-helical conformation and the leucine side chains from one alpha helix interdigitate with those from the alpha helix of a second polypeptide, facilitating dimerization.

The death-effector domain (DED) is a protein interaction domain found only in eukaryotes that regulates a variety of cellular signalling pathways. The DED domain is found in inactive procaspases and proteins that regulate caspase activation in the apoptosis cascade such as FAS-associating death domain-containing protein (FADD). FADD recruits procaspase 8 and procaspase 10 into a death induced signaling complex (DISC). This recruitment is mediated by a homotypic interaction between the procaspase DED and a second DED that is death effector domain in an adaptor protein that is directly associated with activated TNF receptors. Complex formation allows proteolytic activation of procaspase into the active caspase form which results in the initiation of apoptosis. Structurally the DED domain are a subclass of protein motif known as the death fold and contains 6 alpha helices, that closely resemble the structure of the Death domain (DD).

The M1 protein is a matrix protein of the influenza virus. It forms a coat inside the viral envelope. This is a bifunctional membrane/RNA-binding protein that mediates the encapsidation of RNA-nucleoprotein cores into the membrane envelope. It is therefore required that M1 binds both membrane and RNA simultaneously.
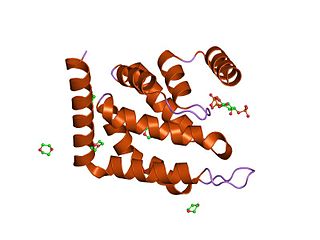
The epsin N-terminal homology (ENTH) domain is a structural domain that is found in proteins involved in endocytosis and cytoskeletal machinery.

DNA repair protein XRCC4 also known as X-ray repair cross-complementing protein 4 or XRCC4 is a protein that in humans is encoded by the XRCC4 gene. In addition to humans, the XRCC4 protein is also expressed in many other metazoans, fungi and in plants. The X-ray repair cross-complementing protein 4 is one of several core proteins involved in the non-homologous end joining (NHEJ) pathway to repair DNA double strand breaks (DSBs).
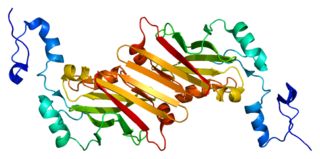
E3 ubiquitin-protein ligase SIAH1 is an enzyme that in humans is encoded by the SIAH1 gene.

Histidine kinases (HK) are multifunctional, and in non-animal kingdoms, typically transmembrane, proteins of the transferase class of enzymes that play a role in signal transduction across the cellular membrane. The vast majority of HKs are homodimers that exhibit autokinase, phosphotransfer, and phosphatase activity. HKs can act as cellular receptors for signaling molecules in a way analogous to tyrosine kinase receptors (RTK). Multifunctional receptor molecules such as HKs and RTKs typically have portions on the outside of the cell that bind to hormone- or growth factor-like molecules, portions that span the cell membrane, and portions within the cell that contain the enzymatic activity. In addition to kinase activity, the intracellular domains typically have regions that bind to a secondary effector molecule or complex of molecules that further propagate signal transduction within the cell. Distinct from other classes of protein kinases, HKs are usually parts of a two-component signal transduction mechanisms in which HK transfers a phosphate group from ATP to a histidine residue within the kinase, and then to an aspartate residue on the receiver domain of a response regulator protein. More recently, the widespread existence of protein histidine phosphorylation distinct from that of two-component histidine kinases has been recognised in human cells. In marked contrast to Ser, Thr and Tyr phosphorylation, the analysis of phosphorylated Histidine using standard biochemical and mass spectrometric approaches is much more challenging, and special procedures and separation techniques are required for their preservation alongside classical Ser, Thr and Tyr phosphorylation on proteins isolated from human cells.

E3 ubiquitin-protein ligase SIAH2 is an enzyme that in humans is encoded by the SIAH2 gene.
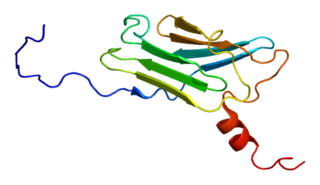
Calcyclin-binding protein is a protein that in humans is encoded by the CACYBP gene.

The tetratricopeptide repeat (TPR) is a structural motif. It consists of a degenerate 34 amino acid tandem repeat identified in a wide variety of proteins. It is found in tandem arrays of 3–16 motifs, which form scaffolds to mediate protein–protein interactions and often the assembly of multiprotein complexes. These alpha-helix pair repeats usually fold together to produce a single, linear solenoid domain called a TPR domain. Proteins with such domains include the anaphase-promoting complex (APC) subunits cdc16, cdc23 and cdc27, the NADPH oxidase subunit p67-phox, hsp90-binding immunophilins, transcription factors, the protein kinase R (PKR), the major receptor for peroxisomal matrix protein import PEX5, protein arginine methyltransferase 9 (PRMT9), and mitochondrial import proteins.
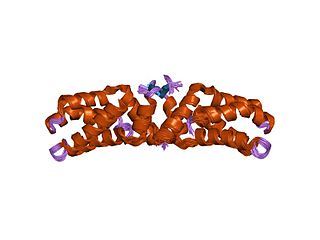
The L27 domain is a protein domain that is found in receptor targeting proteins Lin-2 and Lin-7, as well as some protein kinases and human MPP2 protein. The L27 domain is a protein interaction module that exists in a large family of scaffold proteins, functioning as an organisation centre of large protein assemblies required for the establishment and maintenance of cell polarity. L27 domains form specific heterotetrameric complexes, in which each domain contains three alpha-helices. The L27_2 domain is a protein-protein interaction domain capable of organising scaffold proteins into supramolecular assemblies by formation of heteromeric L27_2 domain complexes. L27_2 domain-mediated protein assemblies have been shown to play essential roles in cellular processes including asymmetric cell division, establishment and maintenance of cell polarity, and clustering of receptors and ion channels. Members of this family form specific heterotetrameric complexes, in which each domain contains three alpha-helices. The two N-terminal helices of each L27_2 domain pack together to form a tight, four-helix bundle in the heterodimer, whilst the third helix of each L27_2 domain forms another four-helix bundle that assembles the two units of the heterodimer into a tetramer.

Bacterial glutathione transferases are part of a superfamily of enzymes that play a crucial role in cellular detoxification. The primary role of GSTs is to catalyze the conjugation of glutathione (GSH) with the electrophilic centers of a wide variety of molecules. The most commonly known substrates of GSTs are xenobiotic synthetic chemicals. There are also classes of GSTs that utilize glutathione as a cofactor rather than a substrate. Often these GSTs are involved in reduction of reactive oxidative species toxic to the bacterium. Conjugation with glutathione receptors reders toxic substances more soluble, and therefore more readily exocytosed from the cell.
The Thiol-activated Cholesterol-dependent Cytolysin(CDC) family is a member of the MACPF superfamily. Cholesterol dependent cytolysins are a family of β-barrel pore-forming exotoxins that are secreted by gram-positive bacteria. CDCs are secreted as water-soluble monomers of 50-70 kDa, that when bound to the target cell, form a circular homo-oligomeric complex containing as many as 40 monomers. Through multiple conformational changes, the β-barrel transmembrane structure is formed and inserted into the target cell membrane. The presence of cholesterol in the target membrane is required for pore formation, though the presence of cholesterol is not required by all CDCs for binding. For example, Intermedilysin secreted by Streptococcus intermedius will bind only to target membranes containing a specific protein receptor, independent of the presence of cholesterol, but cholesterol is required by intermedilysin for pore formation. While the lipid environment of cholesterol in the membrane can affect toxin binding, the exact molecular mechanism that cholesterol regulates the cytolytic activity of the CDC is not fully understood.

S-adenosylmethionine synthetase is an enzyme that creates S-adenosylmethionine by reacting methionine and ATP.
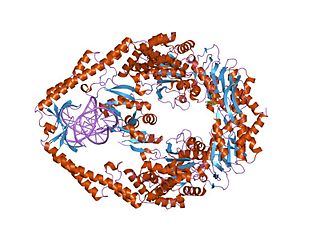
MutS is a mismatch DNA repair protein, originally described in Escherichia coli.

In molecular biology, the cyclase-associated protein family (CAP) is a family of highly conserved actin-binding proteins present in a wide range of organisms including yeast, flies, plants, and mammals. CAPs are multifunctional proteins that contain several structural domains. CAP is involved in species-specific signalling pathways. In Drosophila, CAP functions in Hedgehog-mediated eye development and in establishing oocyte polarity. In Dictyostelium discoideum, CAP is involved in microfilament reorganisation near the plasma membrane in a PIP2-regulated manner and is required to perpetuate the cAMP relay signal to organise fruitbody formation. In plants, CAP is involved in plant signalling pathways required for co-ordinated organ expansion. In yeast, CAP is involved in adenylate cyclase activation, as well as in vesicle trafficking and endocytosis. In both yeast and mammals, CAPs appear to be involved in recycling G-actin monomers from ADF/cofilins for subsequent rounds of filament assembly. In mammals, there are two different CAPs that share 64% amino acid identity.