
Beta-lactamases (β-lactamases) are enzymes produced by bacteria that provide multi-resistance to beta-lactam antibiotics such as penicillins, cephalosporins, cephamycins, monobactams and carbapenems (ertapenem), although carbapenems are relatively resistant to beta-lactamase. Beta-lactamase provides antibiotic resistance by breaking the antibiotics' structure. These antibiotics all have a common element in their molecular structure: a four-atom ring known as a beta-lactam (β-lactam) ring. Through hydrolysis, the enzyme lactamase breaks the β-lactam ring open, deactivating the molecule's antibacterial properties.
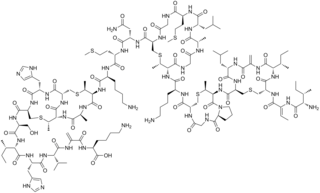
Nisin is a polycyclic antibacterial peptide produced by the bacterium Lactococcus lactis that is used as a food preservative. It has 34 amino acid residues, including the uncommon amino acids lanthionine (Lan), methyllanthionine (MeLan), didehydroalanine (Dha), and didehydroaminobutyric acid (Dhb). These unusual amino acids are introduced by posttranslational modification of the precursor peptide. In these reactions a ribosomally synthesized 57-mer is converted to the final peptide. The unsaturated amino acids originate from serine and threonine, and the enzyme-catalysed addition of cysteine residues to the didehydro amino acids result in the multiple (5) thioether bridges.
Lantibiotics are a class of polycyclic peptide antibiotics that contain the characteristic thioether amino acids lanthionine or methyllanthionine, as well as the unsaturated amino acids dehydroalanine, and 2-aminoisobutyric acid. They belong to ribosomally synthesized and post-translationally modified peptides.
The periplasm is a concentrated gel-like matrix in the space between the inner cytoplasmic membrane and the bacterial outer membrane called the periplasmic space in gram-negative bacteria. Using cryo-electron microscopy it has been found that a much smaller periplasmic space is also present in gram-positive bacteria, between cell wall and the plasma membrane. The periplasm may constitute up to 40% of the total cell volume of gram-negative bacteria, but is a much smaller percentage in gram-positive bacteria.

Polymyxins are antibiotics. Polymyxins B and E are used in the treatment of Gram-negative bacterial infections. They work mostly by breaking up the bacterial cell membrane. They are part of a broader class of molecules called nonribosomal peptides.

Antimicrobial peptides (AMPs), also called host defence peptides (HDPs) are part of the innate immune response found among all classes of life. Fundamental differences exist between prokaryotic and eukaryotic cells that may represent targets for antimicrobial peptides. These peptides are potent, broad spectrum antimicrobials which demonstrate potential as novel therapeutic agents. Antimicrobial peptides have been demonstrated to kill Gram negative and Gram positive bacteria, enveloped viruses, fungi and even transformed or cancerous cells. Unlike the majority of conventional antibiotics it appears that antimicrobial peptides frequently destabilize biological membranes, can form transmembrane channels, and may also have the ability to enhance immunity by functioning as immunomodulators.

Microcins are very small bacteriocins, composed of relatively few amino acids. For this reason, they are distinct from their larger protein cousins. The classic example is microcin V, of Escherichia coli. Subtilosin A is another bacteriocin from Bacillus subtilis. The peptide has a cyclized backbone and forms three cross-links between the sulphurs of Cys13, Cys7 and Cys4 and the alpha-positions of Phe22, Thr28 and Phe31.

Pseudomonas aeruginosa is a common encapsulated, Gram-negative, aerobic–facultatively anaerobic, rod-shaped bacterium that can cause disease in plants and animals, including humans. A species of considerable medical importance, P. aeruginosa is a multidrug resistant pathogen recognized for its ubiquity, its intrinsically advanced antibiotic resistance mechanisms, and its association with serious illnesses – hospital-acquired infections such as ventilator-associated pneumonia and various sepsis syndromes. P. aeruginosa is able to selectively inhibit various antibiotics from penetrating its outer membrane - and has high resistance to several antibiotics. According to the World Health Organization P. aeruginosa poses one of the greatest threats to humans in terms of antibiotic resistance.
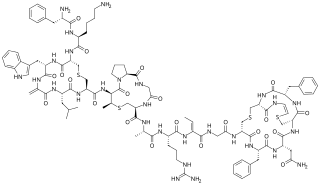
Mutacin 1140 is a bacteriocin produced by Streptococcus mutans. It has activity against a broad spectrum of Gram-positive bacteria. It is a member of the class of compounds known as lantibiotics.

A colicin is a type of bacteriocin produced by and toxic to some strains of Escherichia coli. Colicins are released into the environment to reduce competition from other bacterial strains. Colicins bind to outer membrane receptors, using them to translocate to the cytoplasm or cytoplasmic membrane, where they exert their cytotoxic effect, including depolarisation of the cytoplasmic membrane, DNase activity, RNase activity, or inhibition of murein synthesis.

Class II bacteriocins are a class of small peptides that inhibit the growth of various bacteria.
Sakacins are bacteriocins produced by Lactobacillus sakei. They are often clustered with the other lactic acid bacteriocins. The best known sakacins are sakacin A, G, K, P, and Q. In particular, sakacin A and P have been well characterized.

ATP-dependent Clp protease proteolytic subunit (ClpP) is an enzyme that in humans is encoded by the CLPP gene. This protein is an essential component to form the protein complex of Clp protease.

Rhamnolipids are a class of glycolipid produced by Pseudomonas aeruginosa, amongst other organisms, frequently cited as bacterial surfactants. They have a glycosyl head group, in this case a rhamnose moiety, and a 3-(hydroxyalkanoyloxy)alkanoic acid (HAA) fatty acid tail, such as 3-hydroxydecanoic acid.
Bacterial morphological plasticity refers to changes in the shape and size that bacterial cells undergo when they encounter stressful environments. Although bacteria have evolved complex molecular strategies to maintain their shape, many are able to alter their shape as a survival strategy in response to protist predators, antibiotics, the immune response, and other threats.
Ribosomally synthesized and post-translationally modified peptides (RiPPs), also known as ribosomal natural products, are a diverse class of natural products of ribosomal origin. Consisting of more than 20 sub-classes, RiPPs are produced by a variety of organisms, including prokaryotes, eukaryotes, and archaea, and they possess a wide range of biological functions.

Resistance-nodulation-division (RND) family transporters are a category of bacterial efflux pumps, especially identified in Gram-negative bacteria and located in the cytoplasmic membrane, that actively transport substrates. The RND superfamily includes seven families: the heavy metal efflux (HME), the hydrophobe/amphiphile efflux-1, the nodulation factor exporter family (NFE), the SecDF protein-secretion accessory protein family, the hydrophobe/amphiphile efflux-2 family, the eukaryotic sterol homeostasis family, and the hydrophobe/amphiphile efflux-3 family. These RND systems are involved in maintaining homeostasis of the cell, removal of toxic compounds, and export of virulence determinants. They have a broad substrate spectrum and can lead to the diminished activity of unrelated drug classes if over-expressed. The first reports of drug resistant bacterial infections were reported in the 1940s after the first mass production of antibiotics. Most of the RND superfamily transport systems are made of large polypeptide chains. RND proteins exist primarily in gram-negative bacteria but can also be found in gram-positive bacteria, archaea, and eukaryotes.
Rhs toxins belong to the polymorphic toxin category of bacterial exotoxins. Rhs proteins are widespread and can be produced by both Gram-negative and Gram-positive bacteria. Rhs toxins are very large proteins of usually more than 1,500 aminoacids with variable C-terminal toxic domains. Their toxic activity can either target eukaryotes or other bacteria.

Bacterial secretion systems are protein complexes present on the cell membranes of bacteria for secretion of substances. Specifically, they are the cellular devices used by pathogenic bacteria to secrete their virulence factors to invade the host cells. They can be classified into different types based on their specific structure, composition and activity. Generally, proteins can be secreted through two different processes. One process is a one-step mechanism in which proteins from the cytoplasm of bacteria are transported and delivered directly through the cell membrane into the host cell. Another involves a two-step activity in which the proteins are first transported out of the inner cell membrane, then deposited in the periplasm, and finally through the outer cell membrane into the host cell.
Pyocins are bacteriocins produced by bacteria belonging to the Pseudomonas genus. François Jacob described the first pyocin in 1954. Pyocins can be divided into three distinct classes: S-type, R-type, and F-type pyocins. S-type pyocins are colicin-like bacteriocins as R-type and F-type pyocins belong to tailocins.














