Related Research Articles

The immune system is a network of biological systems that protects an organism from diseases. It detects and responds to a wide variety of pathogens, from viruses to parasitic worms, as well as cancer cells and objects such as wood splinters, distinguishing them from the organism's own healthy tissue. Many species have two major subsystems of the immune system. The innate immune system provides a preconfigured response to broad groups of situations and stimuli. The adaptive immune system provides a tailored response to each stimulus by learning to recognize molecules it has previously encountered. Both use molecules and cells to perform their functions.

Inflammation is part of the biological response of body tissues to harmful stimuli, such as pathogens, damaged cells, or irritants. The five cardinal signs are heat, pain, redness, swelling, and loss of function.

Macrophages are a type of white blood cell of the innate immune system that engulf and digest pathogens, such as cancer cells, microbes, cellular debris, and foreign substances, which do not have proteins that are specific to healthy body cells on their surface. This process is called phagocytosis, which acts to defend the host against infection and injury.

Toll-like receptors (TLRs) are a class of proteins that play a key role in the innate immune system. They are single-spanning receptors usually expressed on sentinel cells such as macrophages and dendritic cells, that recognize structurally conserved molecules derived from microbes. Once these microbes have reached physical barriers such as the skin or intestinal tract mucosa, they are recognized by TLRs, which activate immune cell responses. The TLRs include TLR1, TLR2, TLR3, TLR4, TLR5, TLR6, TLR7, TLR8, TLR9, TLR10, TLR11, TLR12, and TLR13. Humans lack genes for TLR11, TLR12 and TLR13 and mice lack a functional gene for TLR10. The receptors TLR1, TLR2, TLR4, TLR5, TLR6, and TLR10 are located on the cell membrane, whereas TLR3, TLR7, TLR8, and TLR9 are located in intracellular vesicles.
Pathogen-associated molecular patterns (PAMPs) are small molecular motifs conserved within a class of microbes, but not present in the host. They are recognized by toll-like receptors (TLRs) and other pattern recognition receptors (PRRs) in both plants and animals. This allows the innate immune system to recognize pathogens and thus, protect the host from infection.
Pattern recognition receptors (PRRs) play a crucial role in the proper function of the innate immune system. PRRs are germline-encoded host sensors, which detect molecules typical for the pathogens. They are proteins expressed mainly by cells of the innate immune system, such as dendritic cells, macrophages, monocytes, neutrophils, as well as by epithelial cells, to identify two classes of molecules: pathogen-associated molecular patterns (PAMPs), which are associated with microbial pathogens, and damage-associated molecular patterns (DAMPs), which are associated with components of host's cells that are released during cell damage or death. They are also called primitive pattern recognition receptors because they evolved before other parts of the immune system, particularly before adaptive immunity. PRRs also mediate the initiation of antigen-specific adaptive immune response and release of inflammatory cytokines.

The innate immune system or nonspecific immune system is one of the two main immunity strategies in vertebrates. The innate immune system is an alternate defense strategy and is the dominant immune system response found in plants, fungi, prokaryotes, and invertebrates.

The S100 proteins are a family of low molecular-weight proteins found in vertebrates characterized by two calcium-binding sites that have helix-loop-helix ("EF-hand-type") conformation. At least 21 different S100 proteins are known. They are encoded by a family of genes whose symbols use the S100 prefix, for example, S100A1, S100A2, S100A3. They are also considered as damage-associated molecular pattern molecules (DAMPs), and knockdown of aryl hydrocarbon receptor downregulates the expression of S100 proteins in THP-1 cells.
Pyroptosis is a highly inflammatory form of lytic programmed cell death that occurs most frequently upon infection with intracellular pathogens and is likely to form part of the antimicrobial response. This process promotes the rapid clearance of various bacterial, viral, fungal and protozoan infections by removing intracellular replication niches and enhancing the host's defensive responses. Pyroptosis can take place in immune cells and is also reported to occur in keratinocytes and some epithelial cells.

Toll-like receptor 4 (TLR4), also designated as CD284, is a key activator of the innate immune response and plays a central role in the fight against bacterial infections. TLR4 is a transmembrane protein of approximately 95 kDa that is encoded by the TLR4 gene.

Toll-like receptor 6 is a protein that in humans is encoded by the TLR6 gene. TLR6 is a transmembrane protein, member of toll-like receptor family, which belongs to the pattern recognition receptor (PRR) family. TLR6 acts in a heterodimer form with toll-like receptor 2 (TLR2). Its ligands include multiple diacyl lipopeptides derived from gram-positive bacteria and mycoplasma and several fungal cell wall saccharides. After dimerizing with TLR2, the NF-κB intracellular signalling pathway is activated, leading to a pro-inflammatory cytokine production and activation of innate immune response. TLR6 has also been designated as CD286.

Toll-like receptor 9 is a protein that in humans is encoded by the TLR9 gene. TLR9 has also been designated as CD289. It is a member of the toll-like receptor (TLR) family. TLR9 is an important receptor expressed in immune system cells including dendritic cells, macrophages, natural killer cells, and other antigen presenting cells. TLR9 is expressed on endosomes internalized from the plasma membrane, binds DNA, and triggers signaling cascades that lead to a pro-inflammatory cytokine response. Cancer, infection, and tissue damage can all modulate TLR9 expression and activation. TLR9 is also an important factor in autoimmune diseases, and there is active research into synthetic TLR9 agonists and antagonists that help regulate autoimmune inflammation.

High mobility group box 1 protein, also known as high-mobility group protein 1 (HMG-1) and amphoterin, is a protein that in humans is encoded by the HMGB1 gene.
Inflammasomes are cytosolic multiprotein complexes of the innate immune system responsible for the activation of inflammatory responses and cell death. They are formed as a result of specific cytosolic pattern recognition receptors (PRRs) sensing microbe-derived pathogen-associated molecular patterns (PAMPs), damage-associated molecular patterns (DAMPs) from the host cell, or homeostatic disruptions. Activation and assembly of the inflammasome promotes the activation of caspase-1, which then proteolytically cleaves pro-inflammatory cytokines, interleukin 1β (IL-1β) and interleukin 18 (IL-18), as well as the pore-forming molecule gasdermin D (GSDMD). The N-terminal GSDMD fragment resulting from this cleavage induces a pro-inflammatory form of programmed cell death distinct from apoptosis, referred to as pyroptosis, which is responsible for the release of mature cytokines. Additionally, inflammasomes can act as integral components of larger cell death-inducing complexes called PANoptosomes, which drive another distinct form of pro-inflammatory cell death called PANoptosis.
NLRP (Nucleotide-binding oligomerization domain, Leucine rich Repeat and Pyrin domain containing), also abbreviated as NALP, is a type of NOD-like receptor. NOD-like receptors are a type of pattern recognition receptor that are found in the cytosol of the cell, recognizing signals of antigens in the cell. NLRP proteins are part of the innate immune system and detect conserved pathogen characteristics, or pathogen-associated molecular patterns, such as such as peptidoglycan, which is found on some bacterial cells. It is thought that NLRP proteins sense danger signals linked to microbial products, initiating the processes associated with the activation of the inflammasome, including K+ efflux and caspase 1 activation. NLRPs are also known to be associated with a number of diseases. Research suggests NLRP proteins may be involved in combating retroviruses in gametes. As of now, there are at least 14 different known NLRP genes in humans, which are named NLRP1 through NLRP14. The genes translate into proteins with differing lengths of leucine-rich repeat domains.

Necroptosis is a programmed form of necrosis, or inflammatory cell death. Conventionally, necrosis is associated with unprogrammed cell death resulting from cellular damage or infiltration by pathogens, in contrast to orderly, programmed cell death via apoptosis. The discovery of necroptosis showed that cells can execute necrosis in a programmed fashion and that apoptosis is not always the preferred form of cell death. Furthermore, the immunogenic nature of necroptosis favors its participation in certain circumstances, such as aiding in defence against pathogens by the immune system. Necroptosis is well defined as a viral defense mechanism, allowing the cell to undergo "cellular suicide" in a caspase-independent fashion in the presence of viral caspase inhibitors to restrict virus replication. In addition to being a response to disease, necroptosis has also been characterized as a component of inflammatory diseases such as Crohn's disease, pancreatitis, and myocardial infarction.
Murine caspase-11, and its human homologs caspase-4 and caspase-5, are mammalian intracellular receptor proteases activated by TLR4 and TLR3 signaling during the innate immune response. Caspase-11, also termed the non-canonical inflammasome, is activated by TLR3/TLR4-TRIF signaling and directly binds cytosolic lipopolysaccharide (LPS), a major structural element of Gram-negative bacterial cell walls. Activation of caspase-11 by LPS is known to cause the activation of other caspase proteins, leading to septic shock, pyroptosis, and often organismal death.
Immunogenic cell death is any type of cell death eliciting an immune response. Both accidental cell death and regulated cell death can result in immune response. Immunogenic cell death contrasts to forms of cell death that do not elicit any response or even mediate immune tolerance.
Seung-Yong Seong is a South Korean immunologist and microbiologist known for his study of innate immune system response and his development of the damage-associated molecular pattern (DAMP) model of immune response initiation in collaboration with Polly Matzinger. Seong is also known for his research on the bacterium Orientia tsutsugamushi and his research on immunological adjuvant when he was a student. Since 2013 he has served as Director of the Wide River Institute of Immunology – Seoul National University in conjunction with his Professor position in the Microbiology and Immunology department of Seoul National University College of Medicine. In 2012, he became Editor in Chief of the World Journal of Immunology.
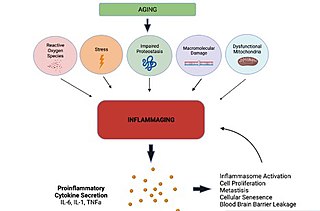
Inflammaging is a chronic, sterile, low-grade inflammation that develops with advanced age, in the absence of overt infection, and may contribute to clinical manifestations of other age-related pathologies. Inflammaging is thought to be caused by a loss of control over systemic inflammation resulting in chronic overstimulation of the innate immune system. Inflammaging is a significant risk factor in mortality and morbidity in aged individuals.
References
- 1 2 Seong SY, Matzinger P (June 2004). "Hydrophobicity: an ancient damage-associated molecular pattern that initiates innate immune responses". Nature Reviews. Immunology. 4 (6): 469–78. doi:10.1038/nri1372. PMID 15173835. S2CID 13336660.
- 1 2 Tang D, Kang R, Coyne CB, Zeh HJ, Lotze MT (September 2012). "PAMPs and DAMPs: signal 0s that spur autophagy and immunity". Immunological Reviews. 249 (1): 158–75. doi:10.1111/j.1600-065X.2012.01146.x. PMC 3662247 . PMID 22889221.
- 1 2 3 4 5 6 7 Roh JS, Sohn DH (August 2018). "Damage-Associated Molecular Patterns in Inflammatory Diseases". Immune Network. 18 (4): e27. doi:10.4110/in.2018.18.e27. PMC 6117512 . PMID 30181915.
- ↑ Roh JS, Sohn DH (August 2018). "Damage-Associated Molecular Patterns in Inflammatory Diseases". Immune Network. 18 (4): e27. doi:10.4110/in.2018.18.e27. PMC 6117512 . PMID 30181915.
- ↑ Chen L, Deng H, Cui H, Fang J, Zuo Z, Deng J, et al. (January 2018). "Inflammatory responses and inflammation-associated diseases in organs". Oncotarget. 9 (6): 7204–7218. doi:10.18632/oncotarget.23208. PMC 5805548 . PMID 29467962.
- ↑ Janeway C (September 1989). "Immunogenicity signals 1,2,3 ... and 0". Immunology Today. 10 (9): 283–6. doi:10.1016/0167-5699(89)90081-9. PMID 2590379.
- 1 2 Rubartelli A, Lotze MT (October 2007). "Inside, outside, upside down: damage-associated molecular-pattern molecules (DAMPs) and redox". Trends in Immunology. 28 (10): 429–36. doi:10.1016/j.it.2007.08.004. PMID 17845865.
- ↑ Land W, Schneeberger H, Schleibner S, Illner WD, Abendroth D, Rutili G, et al. (January 1994). "The beneficial effect of human recombinant superoxide dismutase on acute and chronic rejection events in recipients of cadaveric renal transplants". Transplantation. 57 (2): 211–7. doi: 10.1097/00007890-199401001-00010 . PMID 8310510.
- ↑ Kalogeris T, Baines CP, Krenz M, Korthuis RJ (2012). "Cell biology of ischemia/reperfusion injury". International Review of Cell and Molecular Biology. 298: 229–317. doi:10.1016/B978-0-12-394309-5.00006-7. ISBN 9780123943095. PMC 3904795 . PMID 22878108.
- ↑ Matzinger P (1994). "Tolerance, danger, and the extended family". Annual Review of Immunology. 12: 991–1045. doi:10.1146/annurev.iy.12.040194.005015. PMID 8011301.
- ↑ Panayi GS, Corrigall VM, Henderson B (August 2004). "Stress cytokines: pivotal proteins in immune regulatory networks; Opinion". Current Opinion in Immunology. 16 (4): 531–4. doi:10.1016/j.coi.2004.05.017. PMID 15245751.
- 1 2 Scaffidi P, Misteli T, Bianchi ME (July 2002). "Release of chromatin protein HMGB1 by necrotic cells triggers inflammation". Nature. 418 (6894): 191–5. doi:10.1038/nature00858. PMID 12110890. S2CID 4403741.
- ↑ Scheibner KA, Lutz MA, Boodoo S, Fenton MJ, Powell JD, Horton MR (July 2006). "Hyaluronan fragments act as an endogenous danger signal by engaging TLR2". Journal of Immunology. 177 (2): 1272–81. doi: 10.4049/jimmunol.177.2.1272 . PMID 16818787.
- ↑ Boeynaems JM, Communi D (May 2006). "Modulation of inflammation by extracellular nucleotides". The Journal of Investigative Dermatology. 126 (5): 943–4. doi: 10.1038/sj.jid.5700233 . PMID 16619009.
- ↑ Bours MJ, Swennen EL, Di Virgilio F, Cronstein BN, Dagnelie PC (November 2006). "Adenosine 5'-triphosphate and adenosine as endogenous signaling molecules in immunity and inflammation". Pharmacology & Therapeutics. 112 (2): 358–404. doi:10.1016/j.pharmthera.2005.04.013. PMID 16784779.
- ↑ Shi Y, Evans JE, Rock KL (October 2003). "Molecular identification of a danger signal that alerts the immune system to dying cells". Nature. 425 (6957): 516–21. Bibcode:2003Natur.425..516S. doi:10.1038/nature01991. PMID 14520412. S2CID 2150167.
- ↑ Farkas AM, Kilgore TM, Lotze MT (December 2007). "Detecting DNA: getting and begetting cancer". Current Opinion in Investigational Drugs. 8 (12): 981–6. PMID 18058568.
- ↑ Gardella S, Andrei C, Ferrera D, Lotti LV, Torrisi MR, Bianchi ME, Rubartelli A (October 2002). "The nuclear protein HMGB1 is secreted by monocytes via a non-classical, vesicle-mediated secretory pathway". EMBO Reports. 3 (10): 995–1001. doi:10.1093/embo-reports/kvf198. PMC 1307617 . PMID 12231511.
- ↑ Wang H, Bloom O, Zhang M, Vishnubhakat JM, Ombrellino M, Che J, et al. (July 1999). "HMG-1 as a late mediator of endotoxin lethality in mice". Science. 285 (5425): 248–51. doi:10.1126/science.285.5425.248. PMID 10398600.
- 1 2 Ibrahim ZA, Armour CL, Phipps S, Sukkar MB (December 2013). "RAGE and TLRs: relatives, friends or neighbours?". Molecular Immunology. 56 (4): 739–44. doi:10.1016/j.molimm.2013.07.008. PMID 23954397.
- ↑ Galbiati V, Papale A, Galli CL, Marinovich M, Corsini E (November 2014). "Role of ROS and HMGB1 in contact allergen-induced IL-18 production in human keratinocytes". The Journal of Investigative Dermatology. 134 (11): 2719–2727. doi: 10.1038/jid.2014.203 . PMID 24780928.
- 1 2 Belkaid Y, Hand TW (March 2014). "Role of the microbiota in immunity and inflammation". Cell. 157 (1): 121–41. doi:10.1016/j.cell.2014.03.011. PMC 4056765 . PMID 24679531.
- ↑ Bernard JJ, Cowing-Zitron C, Nakatsuji T, Muehleisen B, Muto J, Borkowski AW, et al. (August 2012). "Ultraviolet radiation damages self noncoding RNA and is detected by TLR3". Nature Medicine. 18 (8): 1286–90. doi:10.1038/nm.2861. PMC 3812946 . PMID 22772463.
- ↑ Diederichs S, Bulk E, Steffen B, Ji P, Tickenbrock L, Lang K, et al. (August 2004). "S100 family members and trypsinogens are predictors of distant metastasis and survival in early-stage non-small cell lung cancer". Cancer Research. 64 (16): 5564–9. doi: 10.1158/0008-5472.CAN-04-2004 . PMID 15313892.
- ↑ Emberley ED, Murphy LC, Watson PH (2004). "S100A7 and the progression of breast cancer". Breast Cancer Research. 6 (4): 153–9. doi: 10.1186/bcr816 . PMC 468668 . PMID 15217486.
- ↑ Emberley ED, Murphy LC, Watson PH (August 2004). "S100 proteins and their influence on pro-survival pathways in cancer". Biochemistry and Cell Biology. 82 (4): 508–15. doi:10.1139/o04-052. PMID 15284904.
- ↑ Lin J, Yang Q, Yan Z, Markowitz J, Wilder PT, Carrier F, Weber DJ (August 2004). "Inhibiting S100B restores p53 levels in primary malignant melanoma cancer cells". The Journal of Biological Chemistry. 279 (32): 34071–7. doi: 10.1074/jbc.M405419200 . PMID 15178678.
- ↑ Marenholz I, Heizmann CW, Fritz G (October 2004). "S100 proteins in mouse and man: from evolution to function and pathology (including an update of the nomenclature)". Biochemical and Biophysical Research Communications. 322 (4): 1111–22. doi:10.1016/j.bbrc.2004.07.096. PMID 15336958.
- 1 2 Maverakis E, Kim K, Shimoda M, Gershwin ME, Patel F, Wilken R, et al. (February 2015). "Glycans in the immune system and The Altered Glycan Theory of Autoimmunity: a critical review". Journal of Autoimmunity. 57: 1–13. doi:10.1016/j.jaut.2014.12.002. PMC 4340844 . PMID 25578468.
- 1 2 Russo MV, McGavern DB (October 2015). "Immune Surveillance of the CNS following Infection and Injury". Trends in Immunology. 36 (10): 637–650. doi:10.1016/j.it.2015.08.002. PMC 4592776 . PMID 26431941.
- ↑ Zeh HJ, Lotze MT (2005). "Addicted to death: invasive cancer and the immune response to unscheduled cell death". Journal of Immunotherapy. 28 (1): 1–9. doi:10.1097/00002371-200501000-00001. PMID 15614039. S2CID 31331291.
- ↑ Kurashima Y, Kiyono H (March 2014). "New era for mucosal mast cells: their roles in inflammation, allergic immune responses and adjuvant development". Experimental & Molecular Medicine. 46 (3): e83. doi:10.1038/emm.2014.7. PMC 3972796 . PMID 24626169.
- ↑ Kurashima Y, Amiya T, Nochi T, Fujisawa K, Haraguchi T, Iba H, et al. (2012). "Extracellular ATP mediates mast cell-dependent intestinal inflammation through P2X7 purinoceptors". Nature Communications. 3: 1034. Bibcode:2012NatCo...3.1034K. doi:10.1038/ncomms2023. PMC 3658010 . PMID 22948816.
- ↑ De Lorenzo G, Ferrari S, Cervone F, Okun E (November 2018). "Extracellular DAMPs in Plants and Mammals: Immunity, Tissue Damage and Repair". Trends in Immunology. 39 (11): 937–950. doi:10.1016/j.it.2018.09.006. PMID 30293747. S2CID 52927468.
- 1 2 Choi HW, Klessig DF (October 2016). "DAMPs, MAMPs, and NAMPs in plant innate immunity". BMC Plant Biology. 16 (1): 232. doi: 10.1186/s12870-016-0921-2 . PMC 5080799 . PMID 27782807.
- ↑ Hou S, Liu Z, Shen H, Wu D (2019-05-22). "Damage-Associated Molecular Pattern-Triggered Immunity in Plants". Frontiers in Plant Science. 10: 646. doi: 10.3389/fpls.2019.00646 . PMC 6547358 . PMID 31191574.
- ↑ Choi HW, Klessig DF (October 2016). "DAMPs, MAMPs, and NAMPs in plant innate immunity". BMC Plant Biology. 16 (1): 232. doi: 10.1186/s12870-016-0921-2 . PMC 5080799 . PMID 27782807.
- 1 2 Foley JF (2015-01-20). "Blocking DAMPs but not PAMPs". Science Signaling. 8 (360): ec13. doi:10.1126/scisignal.aaa6950. S2CID 51601795.
- ↑ Xia C, Braunstein Z, Toomey AC, Zhong J, Rao X (2018). "S100 Proteins As an Important Regulator of Macrophage Inflammation". Frontiers in Immunology. 8: 1908. doi: 10.3389/fimmu.2017.01908 . PMC 5770888 . PMID 29379499.
- ↑ Anders HJ, Schaefer L (July 2014). "Beyond tissue injury-damage-associated molecular patterns, toll-like receptors, and inflammasomes also drive regeneration and fibrosis". Journal of the American Society of Nephrology. 25 (7): 1387–400. doi:10.1681/ASN.2014010117. PMC 4073442 . PMID 24762401.