
Glycolysis is the metabolic pathway that converts glucose into pyruvate and, in most organisms, occurs in the liquid part of cells. The free energy released in this process is used to form the high-energy molecules adenosine triphosphate (ATP) and reduced nicotinamide adenine dinucleotide (NADH). Glycolysis is a sequence of ten reactions catalyzed by enzymes.
A polyphosphate is a salt or ester of polymeric oxyanions formed from tetrahedral PO4 (phosphate) structural units linked together by sharing oxygen atoms. Polyphosphates can adopt linear or a cyclic (also called, ring) structures. In biology, the polyphosphate esters ADP and ATP are involved in energy storage. A variety of polyphosphates find application in mineral sequestration in municipal waters, generally being present at 1 to 5 ppm. GTP, CTP, and UTP are also nucleotides important in the protein synthesis, lipid synthesis, and carbohydrate metabolism, respectively. Polyphosphates are also used as food additives, marked E452.
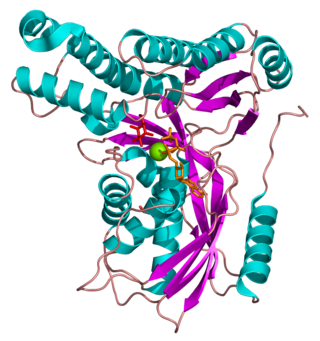
Galactokinase is an enzyme (phosphotransferase) that facilitates the phosphorylation of α-D-galactose to galactose 1-phosphate at the expense of one molecule of ATP. Galactokinase catalyzes the second step of the Leloir pathway, a metabolic pathway found in most organisms for the catabolism of α-D-galactose to glucose 1-phosphate. First isolated from mammalian liver, galactokinase has been studied extensively in yeast, archaea, plants, and humans.
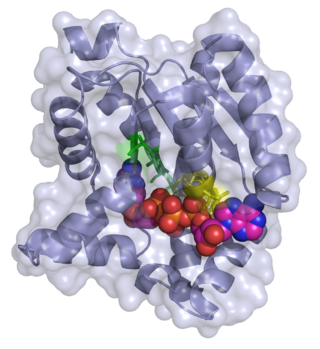
Adenylate kinase is a phosphotransferase enzyme that catalyzes the interconversion of the various adenosine phosphates. By constantly monitoring phosphate nucleotide levels inside the cell, ADK plays an important role in cellular energy homeostasis.

Glycogen phosphorylase is one of the phosphorylase enzymes. Glycogen phosphorylase catalyzes the rate-limiting step in glycogenolysis in animals by releasing glucose-1-phosphate from the terminal alpha-1,4-glycosidic bond. Glycogen phosphorylase is also studied as a model protein regulated by both reversible phosphorylation and allosteric effects.

Inorganic pyrophosphatase is an enzyme that catalyzes the conversion of one ion of pyrophosphate to two phosphate ions. This is a highly exergonic reaction, and therefore can be coupled to unfavorable biochemical transformations in order to drive these transformations to completion. The functionality of this enzyme plays a critical role in lipid metabolism, calcium absorption and bone formation, and DNA synthesis, as well as other biochemical transformations.
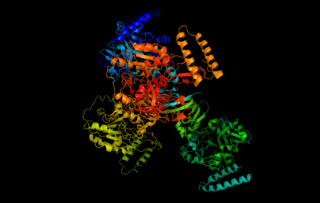
UTP—glucose-1-phosphate uridylyltransferase also known as glucose-1-phosphate uridylyltransferase is an enzyme involved in carbohydrate metabolism. It synthesizes UDP-glucose from glucose-1-phosphate and UTP; i.e.,

Phosphofructokinase (PFK) is a kinase enzyme that phosphorylates fructose 6-phosphate in glycolysis.

Pyridoxine 5′-phosphate oxidase is an enzyme, encoded by the PNPO gene, that catalyzes several reactions in the vitamin B6 metabolism pathway. Pyridoxine 5′-phosphate oxidase catalyzes the final, rate-limiting step in vitamin B6 metabolism, the biosynthesis of pyridoxal 5′-phosphate, the biologically active form of vitamin B6 which acts as an essential cofactor. Pyridoxine 5′-phosphate oxidase is a member of the enzyme class oxidases, or more specifically, oxidoreductases. These enzymes catalyze a simultaneous oxidation-reduction reaction. The substrate oxidase enzymes is hydroxlyated by one oxygen atom of molecular oxygen. Concurrently, the other oxygen atom is reduced to water. Even though molecular oxygen is the electron acceptor in these enzymes' reactions, they are unique because oxygen does not appear in the oxidized product.

Carbamoyl phosphate synthetase catalyzes the ATP-dependent synthesis of carbamoyl phosphate from glutamine or ammonia and bicarbonate. This enzyme catalyzes the reaction of ATP and bicarbonate to produce carboxy phosphate and ADP. Carboxy phosphate reacts with ammonia to give carbamic acid. In turn, carbamic acid reacts with a second ATP to give carbamoyl phosphate plus ADP.
The degradosome is a multiprotein complex present in most bacteria that is involved in the processing of ribosomal RNA and the degradation of messenger RNA and is regulated by Non-coding RNA. It contains the proteins RNA helicase B, RNase E and Polynucleotide phosphorylase.

In enzymology, a phosphoribosylanthranilate isomerase (PRAI) is an enzyme that catalyzes the third step of the synthesis of the amino acid tryptophan.
Choline kinase is an enzyme which catalyzes the first reaction in the choline pathway for phosphatidylcholine (PC) biosynthesis. This reaction involves the transfer of a phosphate group from adenosine triphosphate (ATP) to choline in order to form phosphocholine.

Inositol (1,4,5) trisphosphate 3-kinase (EC 2.7.1.127), abbreviated here as ITP3K, is an enzyme that facilitates a phospho-group transfer from adenosine triphosphate to 1D-myo-inositol 1,4,5-trisphosphate. This enzyme belongs to the family of transferases, specifically those transferring phosphorus-containing groups (phosphotransferases) with an alcohol group as acceptor. The systematic name of this enzyme class is ATP:1D-myo-inositol-1,4,5-trisphosphate 3-phosphotransferase. ITP3K catalyzes the transfer of the gamma-phosphate from ATP to the 3-position of inositol 1,4,5-trisphosphate to form inositol 1,3,4,5-tetrakisphosphate. ITP3K is highly specific for the 1,4,5-isomer of IP3, and it exclusively phosphorylates the 3-OH position, producing Ins(1,3,4,5)P4, also known as inositol tetrakisphosphate or IP4.
In enzymology, a nucleoside-phosphate kinase is an enzyme that catalyzes the chemical reaction
In enzymology, a polyphosphate kinase, or polyphosphate polymerase, is an enzyme that catalyzes the formation of polyphosphate from ATP, with chain lengths of up to a thousand or more orthophosphate moieties.
Obcells are hypothetical proto-organisms or the earliest form of life. The term was first proposed by Thomas Cavalier-Smith in 2001. According to Cavalier-Smith's theory for the origin of the first cell, two cup-shaped obcells or hemicells fused to make a protocell with double-lipid layer envelope, internal genome and ribosomes, protocytosol, and periplasm.
The Arc system is a two-component system found in some bacteria that regulates gene expression in faculatative anaerobes such as Escheria coli. Two-component system means that it has a sensor molecule and a response regulator. Arc is an abbreviation for Anoxic Redox Control system. Arc systems are instrumental in maintaining energy metabolism during transcription of bacteria. The ArcA response regulator looks at growth conditions and expresses genes to best suit the bacteria. The Arc B sensor kinase, which is a tripartite protein, is membrane bound and can autophosphorylate.

GrpE is a bacterial nucleotide exchange factor that is important for regulation of protein folding machinery, as well as the heat shock response. It is a heat-inducible protein and during stress it prevents unfolded proteins from accumulating in the cytoplasm. Accumulation of unfolded proteins in the cytoplasm can lead to cell death.
Sylvy Kornberg née Sylvia Ruth Levy (1917–1986) was an American biochemist who carried out research on DNA replication and polyphosphate synthesis. She discovered and characterized polyphosphate kinase (PPK), an enzyme that helps build long chains of phosphate groups called polyphosphate (PolyP) that play a variety of metabolic and regulatory functions. She worked closely with her husband and research partner, Arthur Kornberg, contributing greatly to the characterization of DNA polymerization that earned him the 1959 Nobel Prize in Physiology or Medicine.














