Related Research Articles

Methionine is an essential amino acid in humans.

Pyridoxal phosphate (PLP, pyridoxal 5'-phosphate, P5P), the active form of vitamin B6, is a coenzyme in a variety of enzymatic reactions. The International Union of Biochemistry and Molecular Biology has catalogued more than 140 PLP-dependent activities, corresponding to ~4% of all classified activities. The versatility of PLP arises from its ability to covalently bind the substrate, and then to act as an electrophilic catalyst, thereby stabilizing different types of carbanionic reaction intermediates.

Cystathionine-β-synthase, also known as CBS, is an enzyme (EC 4.2.1.22) that in humans is encoded by the CBS gene. It catalyzes the first step of the transsulfuration pathway, from homocysteine to cystathionine:
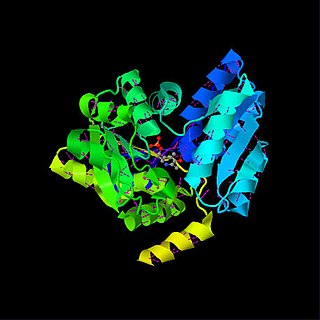
Serine dehydratase or L-serine ammonia lyase (SDH) is in the β-family of pyridoxal phosphate-dependent (PLP) enzymes. SDH is found widely in nature, but its structure and properties vary among species. SDH is found in yeast, bacteria, and the cytoplasm of mammalian hepatocytes. SDH catalyzes the deamination of L-serine to yield pyruvate, with the release of ammonia.

Ornithine aminotransferase (OAT) is an enzyme which is encoded in human by the OAT gene located on chromosome 10.
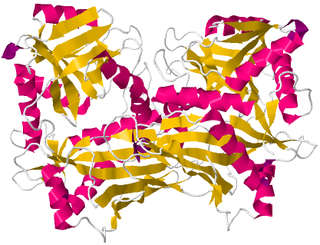
Branched-chain amino acid aminotransferase (BCAT), also known as branched-chain amino acid transaminase, is an aminotransferase enzyme (EC 2.6.1.42) which acts upon branched-chain amino acids (BCAAs). It is encoded by the BCAT2 gene in humans. The BCAT enzyme catalyzes the conversion of BCAAs and α-ketoglutarate into branched chain α-keto acids and glutamate.
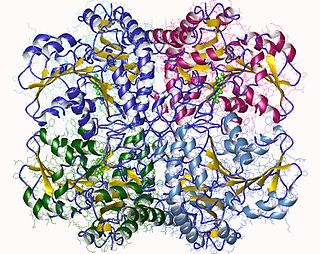
The enzyme cystathionine γ-lyase (EC 4.4.1.1, CTH or CSE; also cystathionase; systematic name L-cystathionine cysteine-lyase (deaminating; 2-oxobutanoate-forming)) breaks down cystathionine into cysteine, 2-oxobutanoate (α-ketobutyrate), and ammonia:
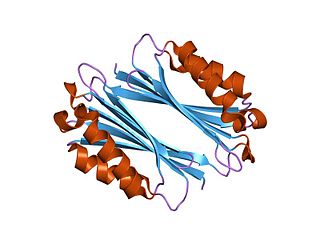
The enzyme adenosylmethionine decarboxylase catalyzes the conversion of S-adenosyl methionine to S-adenosylmethioninamine. Polyamines such as spermidine and spermine are essential for cellular growth under most conditions, being implicated in many cellular processes including DNA, RNA and protein synthesis. S-adenosylmethionine decarboxylase (AdoMetDC) plays an essential regulatory role in the polyamine biosynthetic pathway by generating the n-propylamine residue required for the synthesis of spermidine and spermine from putrescein. Unlike many amino acid decarboxylases AdoMetDC uses a covalently bound pyruvate residue as a cofactor rather than the more common pyridoxal 5'-phosphate. These proteins can be divided into two main groups which show little sequence similarity either to each other, or to other pyruvoyl-dependent amino acid decarboxylases: class I enzymes found in bacteria and archaea, and class II enzymes found in eukaryotes. In both groups the active enzyme is generated by the post-translational autocatalytic cleavage of a precursor protein. This cleavage generates the pyruvate precursor from an internal serine residue and results in the formation of two non-identical subunits termed alpha and beta which form the active enzyme.

The transsulfuration pathway is a metabolic pathway involving the interconversion of cysteine and homocysteine through the intermediate cystathionine. Two transsulfurylation pathways are known: the forward and the reverse.

Cystathionine beta-lyase, also commonly referred to as CBL or β-cystathionase, is an enzyme that primarily catalyzes the following α,β-elimination reaction

The enzyme Acid-Induced Arginine Decarboxylase (AdiA), also commonly referred to as arginine decarboxylase, catalyzes the conversion of L-arginine into agmatine and carbon dioxide. The process consumes a proton in the decarboxylation and employs a pyridoxal-5'-phosphate (PLP) cofactor, similar to other enzymes involved in amino acid metabolism, such as ornithine decarboxylase and glutamine decarboxylase. It is found in bacteria and virus, though most research has so far focused on forms of the enzyme in bacteria. During the AdiA catalyzed decarboxylation of arginine, the necessary proton is consumed from the cell cytoplasm which helps to prevent the over-accumulation of protons inside the cell and serves to increase the intracellular pH. Arginine decarboxylase is part of an enzymatic system in Escherichia coli, Salmonella Typhimurium, and methane-producing bacteria Methanococcus jannaschii that makes these organisms acid resistant and allows them to survive under highly acidic medium.
In enzymology, a cysteine synthase is an enzyme that catalyzes the chemical reaction
In enzymology, a cysteine transaminase is an enzyme that catalyzes the chemical reaction

Aspartate aminotransferase, mitochondrial is an enzyme that in humans is encoded by the GOT2 gene. Glutamic-oxaloacetic transaminase is a pyridoxal phosphate-dependent enzyme which exists in cytoplasmic and inner-membrane mitochondrial forms, GOT1 and GOT2, respectively. GOT plays a role in amino acid metabolism and the urea and Kreb's cycle. Also, GOT2 is a major participant in the malate-aspartate shuttle, which is a passage from the cytosol to the mitochondria. The two enzymes are homodimeric and show close homology. GOT2 has been seen to have a role in cell proliferation, especially in terms of tumor growth.

Cysteine desulfurase, mitochondrial is an enzyme that in humans is encoded by the NFS1 gene.
The Walker A and Walker B motifs are protein sequence motifs, known to have highly conserved three-dimensional structures. These were first reported in ATP-binding proteins by Walker and co-workers in 1982.
In molecular biology, the Cys/Met metabolism PLP-dependent enzyme family is a family of proteins including enzymes involved in cysteine and methionine metabolism which use PLP (pyridoxal-5'-phosphate) as a cofactor.

In molecular biology, group III pyridoxal-dependent decarboxylases are a family of bacterial enzymes comprising ornithine decarboxylase EC 4.1.1.17, lysine decarboxylase EC 4.1.1.18 and arginine decarboxylase EC 4.1.1.19.

In molecular biology, group IV pyridoxal-dependent decarboxylases are a family of enzymes comprising ornithine decarboxylase EC 4.1.1.17, lysine decarboxylase EC 4.1.1.18, arginine decarboxylase EC 4.1.1.19 and diaminopimelate decarboxylaseEC 4.1.1.20. It is also known as the Orn/Lys/Arg decarboxylase class-II family.
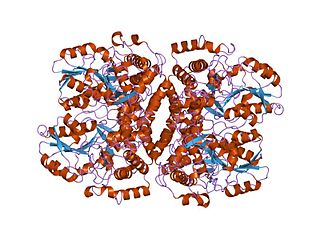

In molecular biology, the group I pyridoxal-dependent decarboxylases, also known as glycine cleavage system P-proteins, are a family of enzymes consisting of glycine cleavage system P-proteins EC 1.4.4.2 from bacterial, mammalian and plant sources. The P protein is part of the glycine decarboxylase multienzyme complex (GDC) also annotated as glycine cleavage system or glycine synthase. The P protein binds the alpha-amino group of glycine through its pyridoxal phosphate cofactor, carbon dioxide is released and the remaining methylamine moiety is then transferred to the lipoamide cofactor of the H protein. GDC consists of four proteins P, H, L and T.
References
- ↑ Ouzounis C, Sander C (May 1993). "Homology of the NifS family of proteins to a new class of pyridoxal phosphate-dependent enzymes". FEBS Letters. 322 (2): 159–64. Bibcode:1993FEBSL.322..159O. doi: 10.1016/0014-5793(93)81559-I . PMID 8482384. S2CID 14587217.