Related Research Articles

A biological membrane, biomembrane or cell membrane is a selectively permeable membrane that separates the interior of a cell from the external environment or creates intracellular compartments by serving as a boundary between one part of the cell and another. Biological membranes, in the form of eukaryotic cell membranes, consist of a phospholipid bilayer with embedded, integral and peripheral proteins used in communication and transportation of chemicals and ions. The bulk of lipids in a cell membrane provides a fluid matrix for proteins to rotate and laterally diffuse for physiological functioning. Proteins are adapted to high membrane fluidity environment of the lipid bilayer with the presence of an annular lipid shell, consisting of lipid molecules bound tightly to the surface of integral membrane proteins. The cell membranes are different from the isolating tissues formed by layers of cells, such as mucous membranes, basement membranes, and serous membranes.

A protease is an enzyme that catalyzes proteolysis, breaking down proteins into smaller polypeptides or single amino acids, and spurring the formation of new protein products. They do this by cleaving the peptide bonds within proteins by hydrolysis, a reaction where water breaks bonds. Proteases are involved in numerous biological pathways, including digestion of ingested proteins, protein catabolism, and cell signaling.
Glycosylation is the reaction in which a carbohydrate, i.e. a glycosyl donor, is attached to a hydroxyl or other functional group of another molecule in order to form a glycoconjugate. In biology, glycosylation usually refers to an enzyme-catalysed reaction, whereas glycation may refer to a non-enzymatic reaction.
An oligosaccharide is a saccharide polymer containing a small number of monosaccharides. Oligosaccharides can have many functions including cell recognition and cell adhesion.
An S-layer is a part of the cell envelope found in almost all archaea, as well as in many types of bacteria. The S-layers of both archaea and bacteria consists of a monomolecular layer composed of only one identical proteins or glycoproteins. This structure is built via self-assembly and encloses the whole cell surface. Thus, the S-layer protein can represent up to 15% of the whole protein content of a cell. S-layer proteins are poorly conserved or not conserved at all, and can differ markedly even between related species. Depending on species, the S-layers have a thickness between 5 and 25 nm and possess identical pores with 2–8 nm in diameter.
A metalloproteinase, or metalloprotease, is any protease enzyme whose catalytic mechanism involves a metal. An example is ADAM12 which plays a significant role in the fusion of muscle cells during embryo development, in a process known as myogenesis.
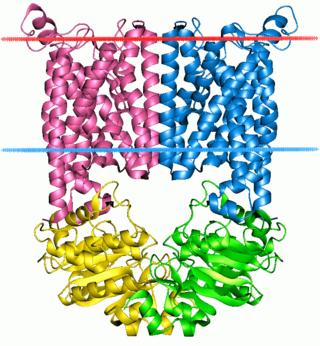
The ATP-binding cassette transporters are a transport system superfamily that is one of the largest and possibly one of the oldest gene families. It is represented in all extant phyla, from prokaryotes to humans. ABC transporters belong to translocases.
Halobacterium is a genus in the family Halobacteriaceae.

Pseudopeptidoglycan is a major cell wall component of some Archaea that differs from bacterial peptidoglycan in chemical structure, but resembles bacterial peptidoglycan in function and physical structure. Pseudopeptidoglycan, in general, is only present in a few methanogenic archaea. The basic components are N-acetylglucosamine and N-acetyltalosaminuronic acid, which are linked by β-1,3-glycosidic bonds.

Ribophorins are dome shaped transmembrane glycoproteins which are located in the membrane of the rough endoplasmic reticulum, but are absent in the membrane of the smooth endoplasmic reticulum. There are two types of ribophorines: ribophorin I and II. These act in the protein complex oligosaccharyltransferase (OST) as two different subunits of the named complex. Ribophorin I and II are only present in eukaryote cells.

Membrane-bound transcription factor site-1 protease, or site-1 protease (S1P) for short, also known as subtilisin/kexin-isozyme 1 (SKI-1), is an enzyme that in humans is encoded by the MBTPS1 gene. S1P cleaves the endoplasmic reticulum loop of sterol regulatory element-binding protein (SREBP) transcription factors.
Archaeocin is the name given to a new type of potentially useful antibiotic that is derived from the Archaea group of organisms. Eight archaeocins have been partially or fully characterized, but hundreds of archaeocins are believed to exist, especially within the haloarchaea. Production of these archaeal proteinaceous antimicrobials is a nearly universal feature of the rod-shaped haloarchaea.

Alphapleolipovirus HHPV1, also known as Haloarcula hispanica pleomorphic virus 1 (HHPV-1) is a virus of the family Pleolipoviridae. It is a double stranded DNA virus that infects the halophilic archaeon Haloarcula hispanica. It has a number of unique features unlike any previously described virus.
The archaellum is a unique structure on the cell surface of many archaea that allows for swimming motility. The archaellum consists of a rigid helical filament that is attached to the cell membrane by a molecular motor. This molecular motor – composed of cytosolic, membrane, and pseudo-periplasmic proteins – is responsible for the assembly of the filament and, once assembled, for its rotation. The rotation of the filament propels archaeal cells in liquid medium, in a manner similar to the propeller of a boat. The bacterial analog of the archaellum is the flagellum, which is also responsible for their swimming motility and can also be compared to a rotating corkscrew. Although the movement of archaella and flagella is sometimes described as "whip-like", this is incorrect, as only cilia from Eukaryotes move in this manner. Indeed, even "flagellum" is a misnomer, as bacterial flagella also work as propeller-like structures.
The Methanosarcinales S-layer Tile Protein (MSTP) is a protein family found almost exclusively in Methanomicrobia members of the order Methanosarcinales. Typically a tandem repeat of two DUF1608 domains are contained in a single MSTP protein chain and these proteins self-assemble into the protective proteinaceous surface layer (S-layer) structure that encompasses the cell. The S-layer, which is found in most Archaea, and in many bacteria, serves many crucial functions including protection from deleterious extracellular substances.
Exosortase refers to a family of integral membrane proteins that occur in Gram-negative bacteria that recognizes and cleaves the carboxyl-terminal sorting signal PEP-CTERM. The name derives from a predicted role analogous to sortase, despite the lack of any detectable sequence homology, and a strong association of exosortase genes with exopolysaccharide or extracellular polymeric substance biosynthesis loci. Many archaea have an archaeosortase, homologous to exosortases rather than to sortases. Archaeosortase A recognizes the signal PGF-CTERM, found at the C-terminus of some archaeal S-layer proteins. Following processing by archaeosortase A, the PGF-CTERM region is gone, and a prenyl-derived lipid anchor is present at the C-terminus instead.
Phosphoglycosyl transferase C (PglC) is an enzyme belonging to a class known as monotopic phosphoglycosyl transferases (PGT). PGTs are required for the synthesis of glycoconjugates on the membrane surface of bacteria. Glycoconjugates, such as glycoproteins, are imperative for bacterial communication as well as host cell interactions between prokaryotic and eukaryotic cells lending to bacteria's pathogenicity.
Sortases are membrane anchored enzyme that sort these surface proteins onto the bacterial cell surface and anchor them to the peptidoglycan. There are different types of sortases and each catalyse the anchoring of different proteins to cell walls.

Haloferax volcanii is a species of organism in the genus Haloferax in the Archaea.
A protein-sorting transpeptidase is an enzyme, such as the sortase SrtA of Staphylococcus aureus, that cleaves one or more target proteins produced by the same cell, as part of a specialized pathway of protein targeting. The typical prokaryotic protein-sorting transpeptidase is characterized as a protease, but does not simply hydrolyze a peptide bond. Instead, the larger, N-terminal portion of the cleaved polypeptide is transferred onto another molecule, such as a precursor of the peptidoglycan cell wall in Gram-positive bacteria.
References
- ↑ Haft, Daniel H.; Samuel H. Payne; Jeremy D. Selengut (January 2012). "Archaeosortases and Exosortases Are Widely Distributed Systems Linking Membrane Transit with Posttranslational Modification". J. Bacteriol. 194 (1): 36–48. doi:10.1128/JB.06026-11. PMC 3256604 . PMID 22037399.
- ↑ Abdul Halim, Mohd Farid; Pfeiffer, Friedhelm; Zou, James; Frisch, Andrew; Haft, Daniel; Wu, Si; Tolić, Nikola; Brewer, Heather; Payne, Samuel H.; Paša-Tolić, Ljiljana; Pohlschroder, Mechthild (Jun 2013). "Haloferax volcanii archaeosortase is required for motility, mating, and C-terminal processing of the S-layer glycoprotein". Mol Microbiol. 88 (6): 1164–75. doi:10.1111/mmi.12248. PMID 23651326. S2CID 5756916.
- ↑ Abdul Halim, MF; Karch, KR; Zhou, Y; Haft, DH; Garcia, BA; Pohlschroder, M (2016). "Permuting the PGF Signature Motif Blocks both Archaeosortase-Dependent C-Terminal Cleavage and Prenyl Lipid Attachment for the Haloferax volcanii S-Layer Glycoprotein". J. Bacteriol. 198 (5): 808–15. doi:10.1128/JB.00849-15. PMC 4810604 . PMID 26712937.
- ↑ Abdul Halim MF, Rodriguez R, Stoltzfus JD, Duggin IG, Pohlschroder M (May 2018). "Conserved residues are critical for Haloferax volcanii archaeosortase catalytic activity: Implications for convergent evolution of the catalytic mechanisms of non-homologous sortases from archaea and bacteria". Molecular Microbiology. 108 (3): 276–287. doi: 10.1111/mmi.13935 . PMID 29465796.
- ↑ Giménez MI, Cerletti M, De Castro RE (2015). "Archaeal membrane-associated proteases: insights on Haloferax volcanii and other haloarchaea". Frontiers in Microbiology. 6: 39. doi: 10.3389/fmicb.2015.00039 . PMC 4343526 . PMID 25774151.