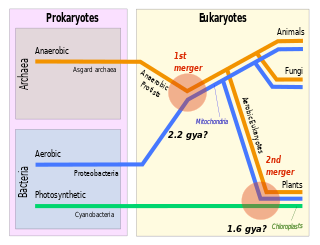
Symbiogenesis is the leading evolutionary theory of the origin of eukaryotic cells from prokaryotic organisms. The theory holds that mitochondria, plastids such as chloroplasts, and possibly other organelles of eukaryotic cells are descended from formerly free-living prokaryotes taken one inside the other in endosymbiosis. Mitochondria appear to be phylogenetically related to Rickettsiales bacteria, while chloroplasts are thought to be related to cyanobacteria.

In biological taxonomy, a domain, also dominion, superkingdom, realm, or empire, is the highest taxonomic rank of all organisms taken together. It was introduced in the three-domain system of taxonomy devised by Carl Woese, Otto Kandler and Mark Wheelis in 1990.
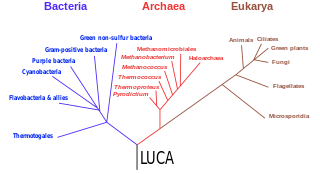
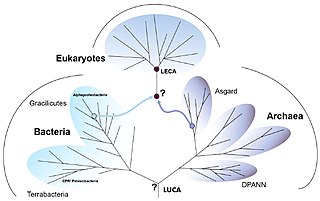
The three-domain system is a taxonomic classification system that groups all cellular life into three domains, namely Archaea, Bacteria and Eukarya, introduced by Carl Woese, Otto Kandler and Mark Wheelis in 1990. The key difference from earlier classifications such as the two-empire system and the five-kingdom classification is the splitting of Archaea from Bacteria as completely different organisms. It has been challenged by the two-domain system that divides organisms into Bacteria and Archaea only, as Eukaryotes are considered as a clade of Archaea.

A unicellular organism, also known as a single-celled organism, is an organism that consists of a single cell, unlike a multicellular organism that consists of multiple cells. Organisms fall into two general categories: prokaryotic organisms and eukaryotic organisms. Most prokaryotes are unicellular and are classified into bacteria and archaea. Many eukaryotes are multicellular, but some are unicellular such as protozoa, unicellular algae, and unicellular fungi. Unicellular organisms are thought to be the oldest form of life, with early protocells possibly emerging 3.5–4.1 billion years ago.

The Thermoproteota are prokaryotes that have been classified as a phylum of the domain Archaea. Initially, the Thermoproteota were thought to be sulfur-dependent extremophiles but recent studies have identified characteristic Thermoproteota environmental rRNA indicating the organisms may be the most abundant archaea in the marine environment. Originally, they were separated from the other archaea based on rRNA sequences; other physiological features, such as lack of histones, have supported this division, although some crenarchaea were found to have histones. Until 2005 all cultured Thermoproteota had been thermophilic or hyperthermophilic organisms, some of which have the ability to grow at up to 113 °C. These organisms stain Gram negative and are morphologically diverse, having rod, cocci, filamentous and oddly-shaped cells. Recent evidence shows that some members of the Thermoproteota are methanogens.

The Korarchaeota is a proposed phylum within the Archaea. The name is derived from the Greek noun koros or kore, meaning young man or young woman, and the Greek adjective archaios which means ancient. They are also known as Xenarchaeota. The name is equivalent to Candidatus Korarchaeota, and they go by the name Xenarchaeota or Xenarchaea as well.
Euryarchaeota is a kingdom of archaea. Euryarchaeota are highly diverse and include methanogens, which produce methane and are often found in intestines; halobacteria, which survive extreme concentrations of salt; and some extremely thermophilic aerobes and anaerobes, which generally live at temperatures between 41 and 122 °C. They are separated from the other archaeans based mainly on rRNA sequences and their unique DNA polymerase.

The last universal common ancestor (LUCA) is the hypothesized common ancestral cell from which the three domains of life, the Bacteria, the Archaea, and the Eukarya originated. The cell had a lipid bilayer; it possessed the genetic code and ribosomes which translated from DNA or RNA to proteins. The LUCA probably existed at latest 3.6 billion years ago, and possibly as early as 4.3 billion years ago or earlier. The nature of this point or stage of divergence remains a topic of research.

A thermoacidophile is an extremophilic microorganism that is both thermophilic and acidophilic; i.e., it can grow under conditions of high temperature and low pH. The large majority of thermoacidophiles are archaea or bacteria, though occasional eukaryotic examples have been reported. Thermoacidophiles can be found in hot springs and solfataric environments, within deep sea vents, or in other environments of geothermal activity. They also occur in polluted environments, such as in acid mine drainage.
Viral eukaryogenesis is the hypothesis that the cell nucleus of eukaryotic life forms evolved from a large DNA virus in a form of endosymbiosis within a methanogenic archaeon or a bacterium. The virus later evolved into the eukaryotic nucleus by acquiring genes from the host genome and eventually usurping its role. The hypothesis was first proposed by Philip Bell in 2001 and was further popularized with the discovery of large, complex DNA viruses that are capable of protein biosynthesis.
Microbial genetics is a subject area within microbiology and genetic engineering. Microbial genetics studies microorganisms for different purposes. The microorganisms that are observed are bacteria and archaea. Some fungi and protozoa are also subjects used to study in this field. The studies of microorganisms involve studies of genotype and expression system. Genotypes are the inherited compositions of an organism. Genetic Engineering is a field of work and study within microbial genetics. The usage of recombinant DNA technology is a process of this work. The process involves creating recombinant DNA molecules through manipulating a DNA sequence. That DNA created is then in contact with a host organism. Cloning is also an example of genetic engineering.

A prokaryote is a single-cell organism whose cell lacks a nucleus and other membrane-bound organelles. The word prokaryote comes from the Ancient Greek πρό (pró), meaning 'before', and κάρυον (káruon), meaning 'nut' or 'kernel'. In the two-empire system arising from the work of Édouard Chatton, prokaryotes were classified within the empire Prokaryota. However in the three-domain system, based upon molecular analysis, prokaryotes are divided into two domains: Bacteria and Archaea. Organisms with nuclei are placed in a third domain: Eukaryota.
The eukaryotes constitute the domain of Eukaryota or Eukarya, organisms whose cells have a membrane-bound nucleus. All animals, plants, fungi, and many unicellular organisms are eukaryotes. They constitute a major group of life forms alongside the two groups of prokaryotes: the Bacteria and the Archaea. Eukaryotes represent a small minority of the number of organisms, but given their generally much larger size, their collective global biomass is much larger than that of prokaryotes.
Evolution of cells refers to the evolutionary origin and subsequent evolutionary development of cells. Cells first emerged at least 3.8 billion years ago approximately 750 million years after Earth was formed.
The eocyte hypothesis in evolutionary biology proposes that the eukaryotes originated from a group of prokaryotes called eocytes. After his team at the University of California, Los Angeles discovered eocytes in 1984, James A. Lake formulated the hypothesis as "eocyte tree" that proposed eukaryotes as part of archaea. Lake hypothesised the tree of life as having only two primary branches: prokaryotes, which include Bacteria and Archaea, and karyotes, that comprise Eukaryotes and eocytes. Parts of this early hypothesis were revived in a newer two-domain system of biological classification which named the primary domains as Archaea and Bacteria.
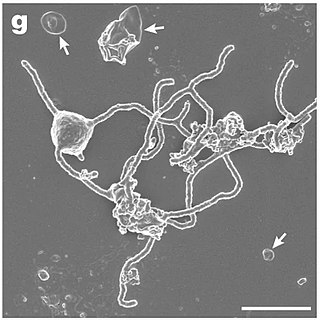
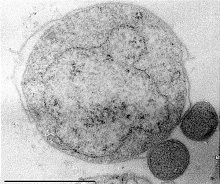
Lokiarchaeota is a proposed phylum of the Archaea. The phylum includes all members of the group previously named Deep Sea Archaeal Group, also known as Marine Benthic Group B. Lokiarchaeota is part of the superphylum Asgard containing the phyla: Lokiarchaeota, Thorarchaeota, Odinarchaeota, Heimdallarchaeota, and Helarchaeota. A phylogenetic analysis disclosed a monophyletic grouping of the Lokiarchaeota with the eukaryotes. The analysis revealed several genes with cell membrane-related functions. The presence of such genes support the hypothesis of an archaeal host for the emergence of the eukaryotes; the eocyte-like scenarios.
Eukaryogenesis, the process which created the eukaryotic cell and lineage, is a milestone in the evolution of life, since eukaryotes include all complex cells and almost all multicellular organisms. The process is widely agreed to have involved symbiogenesis, in which an archeon and a bacterium came together to create the first eukaryotic common ancestor (FECA). This cell had a new level of complexity and capability, with a nucleus, at least one centriole and cilium, facultatively aerobic mitochondria, sex, a dormant cyst with a cell wall of chitin and/or cellulose and peroxisomes. It evolved into a population of single-celled organisms that included the last eukaryotic common ancestor (LECA), gaining capabilities along the way, though the sequence of the steps involved has been disputed, and may not have started with symbiogenesis. In turn, the LECA gave rise to the eukaryotes' crown group, containing the ancestors of animals, fungi, plants, and a diverse range of single-celled organisms.
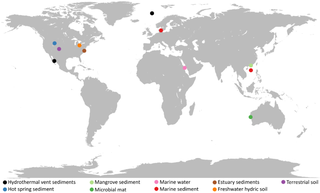
Asgard or Asgardarchaeota is a proposed superphylum belonging to the domain Archaea that contain eukaryotic signature proteins. It appears that the eukaryotes, the domain that contains the animals, plants, and fungi, emerged within the Asgard, in a branch containing the Heimdallarchaeota. This supports the two-domain system of classification over the three-domain system.

Marine prokaryotes are marine bacteria and marine archaea. They are defined by their habitat as prokaryotes that live in marine environments, that is, in the saltwater of seas or oceans or the brackish water of coastal estuaries. All cellular life forms can be divided into prokaryotes and eukaryotes. Eukaryotes are organisms whose cells have a nucleus enclosed within membranes, whereas prokaryotes are the organisms that do not have a nucleus enclosed within a membrane. The three-domain system of classifying life adds another division: the prokaryotes are divided into two domains of life, the microscopic bacteria and the microscopic archaea, while everything else, the eukaryotes, become the third domain.
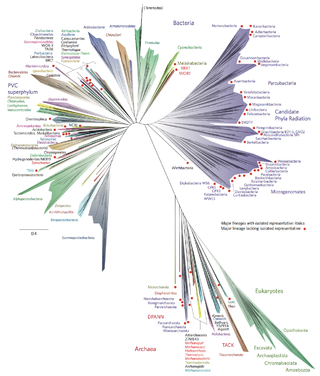
The two-domain system is a biological classification by which all organisms in the tree of life are classified into two domains, Bacteria and Archaea. It emerged from development of knowledge of archaea diversity and challenges the widely accepted three-domain system that classifies life into Bacteria, Archaea, and Eukarya. It was preceded by the eocyte hypothesis of James A. Lake in the 1980s, which was largely superseded by the three-domain system, due to evidence at the time. Better understanding of archaea, especially of their roles in the origin of eukaryotes through symbiogenesis with bacteria, led to the revival of the eocyte hypothesis in the 2000s. The two-domain system became more widely accepted after the discovery of a large group (superphylum) of archaea called Asgard in 2017, which evidence suggests to be the evolutionary root of eukaryotes, thereby making eukaryotes members of the domain Archaea.


























