| Thermoproteota | |
|---|---|
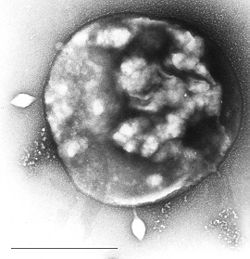 | |
| Archaea Sulfolobus infected with specific virus STSV-1. | |
| Scientific classification | |
| Domain: | Archaea |
| Kingdom: | Thermoproteati |
| Phylum: | Thermoproteota Garrity & Holt 2021 [1] |
| Classes | |
| |
| Synonyms | |
| |
The Thermoproteota are prokaryotes that have been classified as a phylum of the domain Archaea. [3] [4] [5] Initially, the Thermoproteota were thought to be sulfur-dependent extremophiles but recent studies have identified characteristic Thermoproteota environmental rRNA indicating the organisms may be the most abundant archaea in the marine environment. [6] Originally, they were separated from the other archaea based on rRNA sequences; other physiological features, such as lack of histones, have supported this division, although some crenarchaea were found to have histones. [7] Until 2005 all cultured Thermoproteota had been thermophilic or hyperthermophilic organisms, some of which have the ability to grow at up to 113 °C. [8] These organisms stain Gram negative and are morphologically diverse, having rod, cocci, filamentous and oddly-shaped cells. [9] Recent evidence shows that some members of the Thermoproteota are methanogens.
Contents
- Sulfolobus
- Recombinational repair of DNA damage
- Marine species
- Possible connections with eukaryotes
- See also
- References
- Scientific journals
- Scientific handbooks
- External links
Thermoproteota were initially classified as a part of regnum Eocyta in 1984, [10] but this classification has been discarded. The term "eocyte" now applies to either TACK (formerly Crenarchaeota) or to Thermoproteota.