Related Research Articles
Chemistry is the scientific study of the properties and behavior of matter. It is a physical science within the natural sciences that studies the chemical elements that make up matter and compounds made of atoms, molecules and ions: their composition, structure, properties, behavior and the changes they undergo during reactions with other substances. Chemistry also addresses the nature of chemical bonds in chemical compounds.

Computational chemistry is a branch of chemistry that uses computer simulations to assist in solving chemical problems. It uses methods of theoretical chemistry incorporated into computer programs to calculate the structures and properties of molecules, groups of molecules, and solids. The importance of this subject stems from the fact that, with the exception of some relatively recent findings related to the hydrogen molecular ion, achieving an accurate quantum mechanical depiction of chemical systems analytically, or in a closed form, is not feasible. The complexity inherent in the many-body problem exacerbates the challenge of providing detailed descriptions of quantum mechanical systems. While computational results normally complement information obtained by chemical experiments, it can occasionally predict unobserved chemical phenomena.
The following outline is provided as an overview of and topical guide to chemistry:

Physical science is a branch of natural science that studies non-living systems, in contrast to life science. It in turn has many branches, each referred to as a "physical science", together is called the "physical sciences".
Quantum chemistry, also called molecular quantum mechanics, is a branch of physical chemistry focused on the application of quantum mechanics to chemical systems, particularly towards the quantum-mechanical calculation of electronic contributions to physical and chemical properties of molecules, materials, and solutions at the atomic level. These calculations include systematically applied approximations intended to make calculations computationally feasible while still capturing as much information about important contributions to the computed wave functions as well as to observable properties such as structures, spectra, and thermodynamic properties. Quantum chemistry is also concerned with the computation of quantum effects on molecular dynamics and chemical kinetics.

Theoretical chemistry is the branch of chemistry which develops theoretical generalizations that are part of the theoretical arsenal of modern chemistry: for example, the concepts of chemical bonding, chemical reaction, valence, the surface of potential energy, molecular orbitals, orbital interactions, and molecule activation.
Computer science is the study of the theoretical foundations of information and computation and their implementation and application in computer systems. One well known subject classification system for computer science is the ACM Computing Classification System devised by the Association for Computing Machinery.

Permutation City is a 1994 science-fiction novel by Greg Egan that explores many concepts, including quantum ontology, through various philosophical aspects of artificial life and simulated reality. Sections of the story were adapted from Egan's 1992 short story "Dust", which dealt with many of the same philosophical themes. Permutation City won the John W. Campbell Award for the best science-fiction novel of the year in 1995 and was nominated for the Philip K. Dick Award the same year. The novel was also cited in a 2003 Scientific American article on multiverses by Max Tegmark.
Dendral was a project in artificial intelligence (AI) of the 1960s, and the computer software expert system that it produced. Its primary aim was to study hypothesis formation and discovery in science. For that, a specific task in science was chosen: help organic chemists in identifying unknown organic molecules, by analyzing their mass spectra and using knowledge of chemistry. It was done at Stanford University by Edward Feigenbaum, Bruce G. Buchanan, Joshua Lederberg, and Carl Djerassi, along with a team of highly creative research associates and students. It began in 1965 and spans approximately half the history of AI research.
Mathematical and theoretical biology, or biomathematics, is a branch of biology which employs theoretical analysis, mathematical models and abstractions of living organisms to investigate the principles that govern the structure, development and behavior of the systems, as opposed to experimental biology which deals with the conduction of experiments to test scientific theories. The field is sometimes called mathematical biology or biomathematics to stress the mathematical side, or theoretical biology to stress the biological side. Theoretical biology focuses more on the development of theoretical principles for biology while mathematical biology focuses on the use of mathematical tools to study biological systems, even though the two terms are sometimes interchanged.
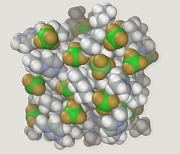
Molecular modelling encompasses all methods, theoretical and computational, used to model or mimic the behaviour of molecules. The methods are used in the fields of computational chemistry, drug design, computational biology and materials science to study molecular systems ranging from small chemical systems to large biological molecules and material assemblies. The simplest calculations can be performed by hand, but inevitably computers are required to perform molecular modelling of any reasonably sized system. The common feature of molecular modelling methods is the atomistic level description of the molecular systems. This may include treating atoms as the smallest individual unit, or explicitly modelling protons and neutrons with its quarks, anti-quarks and gluons and electrons with its photons.
Mathematical chemistry is the area of research engaged in novel applications of mathematics to chemistry; it concerns itself principally with the mathematical modeling of chemical phenomena. Mathematical chemistry has also sometimes been called computer chemistry, but should not be confused with computational chemistry.
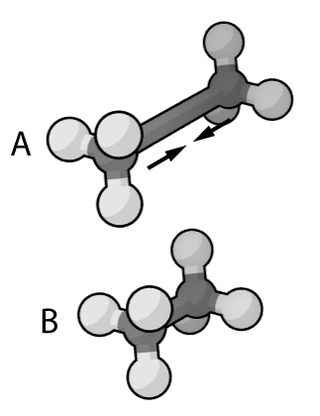
In the context of chemistry, molecular physics, physical chemistry, and molecular modelling, a force field is a computational model that is used to describe the forces between atoms within molecules or between molecules as well as in crystals. Force fields are a variety of interatomic potentials. More precisely, the force field refers to the functional form and parameter sets used to calculate the potential energy of a system on the atomistic level. Force fields are usually used in molecular dynamics or Monte Carlo simulations. The parameters for a chosen energy function may be derived from classical laboratory experiment data, calculations in quantum mechanics, or both. Force fields utilize the same concept as force fields in classical physics, with the main difference being that the force field parameters in chemistry describe the energy landscape on the atomistic level. From a force field, the acting forces on every particle are derived as a gradient of the potential energy with respect to the particle coordinates.
In probability theory, the Gillespie algorithm generates a statistically correct trajectory of a stochastic equation system for which the reaction rates are known. It was created by Joseph L. Doob and others, presented by Dan Gillespie in 1976, and popularized in 1977 in a paper where he uses it to simulate chemical or biochemical systems of reactions efficiently and accurately using limited computational power. As computers have become faster, the algorithm has been used to simulate increasingly complex systems. The algorithm is particularly useful for simulating reactions within cells, where the number of reagents is low and keeping track of every single reaction is computationally feasible. Mathematically, it is a variant of a dynamic Monte Carlo method and similar to the kinetic Monte Carlo methods. It is used heavily in computational systems biology.
Physical organic chemistry, a term coined by Louis Hammett in 1940, refers to a discipline of organic chemistry that focuses on the relationship between chemical structures and reactivity, in particular, applying experimental tools of physical chemistry to the study of organic molecules. Specific focal points of study include the rates of organic reactions, the relative chemical stabilities of the starting materials, reactive intermediates, transition states, and products of chemical reactions, and non-covalent aspects of solvation and molecular interactions that influence chemical reactivity. Such studies provide theoretical and practical frameworks to understand how changes in structure in solution or solid-state contexts impact reaction mechanism and rate for each organic reaction of interest.

The branches of science, also referred to as sciences, scientificfields or scientific disciplines, are commonly divided into three major groups:
Natural computing, also called natural computation, is a terminology introduced to encompass three classes of methods: 1) those that take inspiration from nature for the development of novel problem-solving techniques; 2) those that are based on the use of computers to synthesize natural phenomena; and 3) those that employ natural materials to compute. The main fields of research that compose these three branches are artificial neural networks, evolutionary algorithms, swarm intelligence, artificial immune systems, fractal geometry, artificial life, DNA computing, and quantum computing, among others.
Multi-state modeling of biomolecules refers to a series of techniques used to represent and compute the behaviour of biological molecules or complexes that can adopt a large number of possible functional states.
A chemical graph generator is a software package to generate computer representations of chemical structures adhering to certain boundary conditions. The development of such software packages is a research topic of cheminformatics. Chemical graph generators are used in areas such as virtual library generation in drug design, in molecular design with specified properties, called inverse QSAR/QSPR, as well as in organic synthesis design, retrosynthesis or in systems for computer-assisted structure elucidation (CASE). CASE systems again have regained interest for the structure elucidation of unknowns in computational metabolomics, a current area of computational biology.
References
- 1 2 W. Banzhaf and L. Yamamoto. Artificial Chemistries, MIT Press, 2015.
- ↑ P. Dittrich. Artificial chemistry (AC) In A. R. Meyers (ed.), Computational Complexity: Theory, Techniques, and Applications, pp. 185-203, Springer, 2012.
- 1 2 P. Dittrich, J. Ziegler, and W. Banzhaf. Artificial chemistries — A review. Artificial Life, 7(3):225–275, 2001.
- ↑ J.S.McCaskill. Polymer chemistry on tape: A computational model for emergent genetics. Technical report, MPI for Biophysical Chemistry, 1988.
- ↑ W. Fontana. Algorithmic chemistry. In C. G. Langton, C. Taylor, J. D. Farmer, and S. Rasmussen, editors, Artificial Life II, pages 159–210. Westview Press, 1991.
- ↑ W. Fontana and L. Buss. “The arrival of the fittest”: Toward a theory of biological organization. Bulletin of Mathematical Biology, 56(1):1–64, 1994.
- ↑ P. Dittrich, P. Speroni di Fenizio. Chemical Organization Theory. Bulletin of Mathematical Biology (2007) 69: 1199:1231.
- ↑ P. Speroni di Fenizio. Chemical Organization Theory. PhD thesis, Friedrich Schiller University Jena, 2007.
- ↑ T. Ikegami and T. Hashimoto. Active mutation in self-reproducing networks of machines and tapes. Artificial Life, 2(3):305–318, 1995.
- ↑ T. Ikegami and T.Hashimoto. Replication and diversity in machine-tape coevolutionary systems. In C. G. Langton and K. Shimohara, editors, Artificial Life V, pages 426–433. MIT Press, 1997.
- ↑ H.Suzuki. Models for the conservation of genetic information with string-based artificial chemistry. In W. Banzhaf, J. Ziegler, T. Christaller, P. Dittrich, and J. T. Kim, editors, Advances in Artificial Life, volume 2801 of Lecture Notes in Computer Science, pages 78–88. Springer, 2003.
- ↑ H. Suzuki. A network cell with molecular agents that divides from centrosome signals. Biosystems, 94(1-2):118–125, 2008.
- ↑ Y. Suzuki, J. Takabayashi, and H. Tanaka. Investigation of tritrophic interactions in an ecosystem using abstract chemistry. Artificial Life and Robotics, 6(3):129–132, 2002.
- ↑ Y. Suzuki and H. Tanaka. Modeling p53 signaling pathways by using multiset processing. In G. Ciobanu, G. Pa ̆un, and M. J. Pérez-Jiménez, editors, Applications of Membrane Computing, Natural Computing Series, pages 203–214. Springer, 2006.
- ↑ J.Ziegler and W.Banzhaf. Evolving control metabolisms for a robot. ArtificialLife, 7(2):171–190, 2001.