Related Research Articles
Biophysics is an interdisciplinary science that applies approaches and methods traditionally used in physics to study biological phenomena. Biophysics covers all scales of biological organization, from molecular to organismic and populations. Biophysical research shares significant overlap with biochemistry, molecular biology, physical chemistry, physiology, nanotechnology, bioengineering, computational biology, biomechanics, developmental biology and systems biology.
Chemistry at Harvard Macromolecular Mechanics (CHARMM) is the name of a widely used set of force fields for molecular dynamics, and the name for the molecular dynamics simulation and analysis computer software package associated with them. The CHARMM Development Project involves a worldwide network of developers working with Martin Karplus and his group at Harvard to develop and maintain the CHARMM program. Licenses for this software are available, for a fee, to people and groups working in academia.
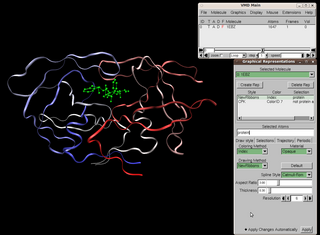
Visual Molecular Dynamics (VMD) is a molecular modelling and visualization computer program. VMD is developed mainly as a tool to view and analyze the results of molecular dynamics simulations. It also includes tools for working with volumetric data, sequence data, and arbitrary graphics objects. Molecular scenes can be exported to external rendering tools such as POV-Ray, RenderMan, Tachyon, Virtual Reality Modeling Language (VRML), and many others. Users can run their own Tcl and Python scripts within VMD as it includes embedded Tcl and Python interpreters. VMD runs on Unix, Apple Mac macOS, and Microsoft Windows. VMD is available to non-commercial users under a distribution-specific license which permits both use of the program and modification of its source code, at no charge.
B. Montgomery "Monte" Pettitt is the Director of the Sealy Center for Structural Biology and Molecular Biophysics at University of Texas Medical Branch in Galveston, Texas, holder of the Robert A. Welch Distinguished Chair in Chemistry, and tenured Professor in the Department of Biochemistry and Molecular Biology, as well as the department of Pharmacology and Toxicology. He is also affiliated with and former director of The W. M. Keck Center at Rice University, and a faculty member of the Structural and Computational Biology and Molecular Biophysics program at Baylor College of Medicine. At the University of Houston, he was the Hugh Roy and Lille Cranz Cullen Distinguished Professor of Chemistry and Robert A. Welch Chair in Chemistry, as well as the director of its Institute for Molecular Design.

The Max Planck Institute for Biophysical Chemistry, also known as the Karl-Friedrich Bonhoeffer Institute, was a research institute of the Max Planck Society, located in Göttingen, Germany. On January 1, 2022, the institute merged with the Max Planck Institute for Experimental Medicine in Göttingen to form the Max Planck Institute for Multidisciplinary Sciences.

Martin Karplus is an Austrian and American theoretical chemist. He is the Director of the Biophysical Chemistry Laboratory, a joint laboratory between the French National Center for Scientific Research and the University of Strasbourg, France. He is also the Theodore William Richards Professor of Chemistry, emeritus at Harvard University. Karplus received the 2013 Nobel Prize in Chemistry, together with Michael Levitt and Arieh Warshel, for "the development of multiscale models for complex chemical systems".
Molecular biophysics is a rapidly evolving interdisciplinary area of research that combines concepts in physics, chemistry, engineering, mathematics and biology. It seeks to understand biomolecular systems and explain biological function in terms of molecular structure, structural organization, and dynamic behaviour at various levels of complexity. This discipline covers topics such as the measurement of molecular forces, molecular associations, allosteric interactions, Brownian motion, and cable theory. Additional areas of study can be found on Outline of Biophysics. The discipline has required development of specialized equipment and procedures capable of imaging and manipulating minute living structures, as well as novel experimental approaches.
Implicit solvation is a method to represent solvent as a continuous medium instead of individual “explicit” solvent molecules, most often used in molecular dynamics simulations and in other applications of molecular mechanics. The method is often applied to estimate free energy of solute-solvent interactions in structural and chemical processes, such as folding or conformational transitions of proteins, DNA, RNA, and polysaccharides, association of biological macromolecules with ligands, or transport of drugs across biological membranes.
Drude particles are model oscillators used to simulate the effects of electronic polarizability in the context of a classical molecular mechanics force field. They are inspired by the Drude model of mobile electrons and are used in the computational study of proteins, nucleic acids, and other biomolecules.
Jeremy Christopher Smith is a British-born computational molecular biophysicist.

Arieh Warshel is an Israeli-American biochemist and biophysicist. He is a pioneer in computational studies on functional properties of biological molecules, Distinguished Professor of Chemistry and Biochemistry, and holds the Dana and David Dornsife Chair in Chemistry at the University of Southern California. He received the 2013 Nobel Prize in Chemistry, together with Michael Levitt and Martin Karplus for "the development of multiscale models for complex chemical systems".
The following outline is provided as an overview of and topical guide to biophysics:
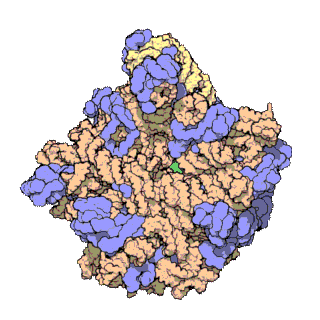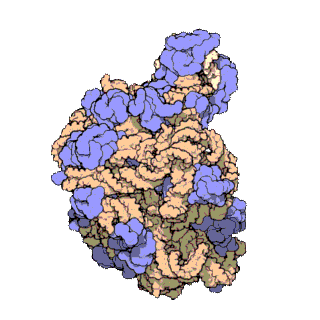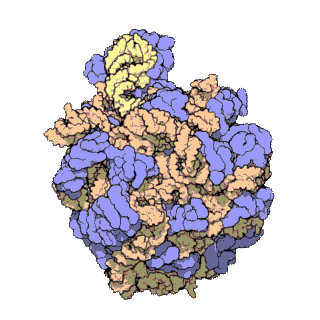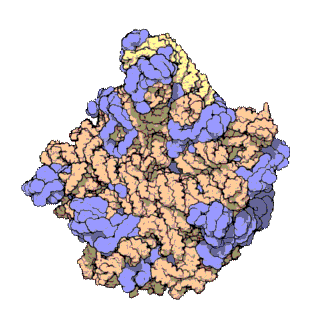
In molecular biology, the term macromolecular assembly (MA) refers to massive chemical structures such as viruses and non-biologic nanoparticles, cellular organelles and membranes and ribosomes, etc. that are complex mixtures of polypeptide, polynucleotide, polysaccharide or other polymeric macromolecules. They are generally of more than one of these types, and the mixtures are defined spatially, and with regard to their underlying chemical composition and structure. Macromolecules are found in living and nonliving things, and are composed of many hundreds or thousands of atoms held together by covalent bonds; they are often characterized by repeating units. Assemblies of these can likewise be biologic or non-biologic, though the MA term is more commonly applied in biology, and the term supramolecular assembly is more often applied in non-biologic contexts. MAs of macromolecules are held in their defined forms by non-covalent intermolecular interactions, and can be in either non-repeating structures, or in repeating linear, circular, spiral, or other patterns. The process by which MAs are formed has been termed molecular self-assembly, a term especially applied in non-biologic contexts. A wide variety of physical/biophysical, chemical/biochemical, and computational methods exist for the study of MA; given the scale of MAs, efforts to elaborate their composition and structure and discern mechanisms underlying their functions are at the forefront of modern structure science.

G. Marius Clore MAE, FRSC, FMedSci, FRS is a British-born, Anglo-American molecular biophysicist and structural biologist. He was born in London, U.K. and is a dual U.S./U.K. Citizen. He is a Member of the National Academy of Sciences, a Fellow of the Royal Society, a Fellow of the Academy of Medical Sciences, a Fellow of the American Academy of Arts and Sciences, a NIH Distinguished Investigator, and the Chief of the Molecular and Structural Biophysics Section in the Laboratory of Chemical Physics of the National Institute of Diabetes and Digestive and Kidney Diseases at the U.S. National Institutes of Health. He is known for his foundational work in three-dimensional protein and nucleic acid structure determination by biomolecular NMR spectroscopy, for advancing experimental approaches to the study of large macromolecules and their complexes by NMR, and for developing NMR-based methods to study rare conformational states in protein-nucleic acid and protein-protein recognition. Clore's discovery of previously undetectable, functionally significant, rare transient states of macromolecules has yielded fundamental new insights into the mechanisms of important biological processes, and in particular the significance of weak interactions and the mechanisms whereby the opposing constraints of speed and specificity are optimized. Further, Clore's work opens up a new era of pharmacology and drug design as it is now possible to target structures and conformations that have been heretofore unseen.
Klaus Schulten was a German-American computational biophysicist and the Swanlund Professor of Physics at the University of Illinois at Urbana-Champaign. Schulten used supercomputing techniques to apply theoretical physics to the fields of biomedicine and bioengineering and dynamically model living systems. His mathematical, theoretical, and technological innovations led to key discoveries about the motion of biological cells, sensory processes in vision, animal navigation, light energy harvesting in photosynthesis, and learning in neural networks.
Kenneth M. Merz Jr. is an American biochemist and molecular biologist currently the Joseph Zichis Chair and a distinguished university professor at Michigan State University and editor-in-chief of American Chemical Society's Journal of Chemical Information and Modeling. A highly cited expert in his field, his research interests are in computational chemistry and biology and computer-aided drug design (CADD). His group has been involved in developing the widely using AMBER suite of programs for simulating chemical and biological systems and the QUICK program for quantum chemical calculations.
Barry H. Honig is an American biochemist, molecular biophysicist, and computational biophysicist, who develops theoretical methods and computer software for "analyzing the structure and function of biological macromolecules."

Syma Khalid is a British Scientist who is a Professor of Computational Microbiology in at the University of Oxford. She was awarded the Suffrage Science award for engineering and physical sciences in 2021.
Alexander D. MacKerell, Jr. is an American biophysicist who is the Grollman-Glick Professor of Pharmaceutical Sciences at the University of Maryland, Baltimore (UMB) and the Director of the Computer-Aided Drug Design (CADD) Center at UMB. He is also the Co-Founder and Chief Scientific Officer of the drug design tech company SilcsBio. In 2022, MacKerell was awarded the prestigious American Chemical Society Award for Computers in Chemical and Pharmaceutical Research.
References
- ↑ "Department of Biochemistry & Molecular Biology". Archived from the original on 2016-01-24.
- ↑ "Faculty members receive named, distinguished service professorships". University of Chicago. January 27, 2015. Retrieved May 11, 2019.
- ↑ "Benoît Roux". University of Chicago. Retrieved March 11, 2019.
- ↑ "Benoît Roux". University of Chicago. Retrieved May 11, 2019.
- ↑ Ion transport in a model gramicidin channel. Structure and thermodynamics. B. Roux and M. Karplus Biophys. J. 1991;59(5):961-981.
- ↑ Theoretical and computational models of ion channels. B. Roux Curr Opin Struct Biol. 2002;12(2):182-189.
- ↑ Merz, Kenneth M.; Roux, Benoît (1996). Biological Membranes: A Molecular Perspective from Computation and Experiment. Birkhäuser. doi:10.1007/978-1-4684-8580-6. ISBN 1468485806. S2CID 30908633.
- ↑ ROUX, BENOIT (2001). Becker, Oren M.; MacKerell Jr., Alexander D.; Roux, Benoit; Watanabe, Masakatsu (eds.). Computational Biochemistry and Biophysics. Taylor Francis. doi:10.1201/9780203903827. ISBN 978-0-203-90382-7.
- ↑ ROUX, BENOIT (2021). COMPUTATIONAL MODELING AND SIMULATIONS OF BIOMOLECULAR SYSTEMS : simulations of... biomolecular systems. [S.l.]: WORLD SCIENTIFIC PUB. doi:10.1142/12173. ISBN 978-981-12-3275-6. OCLC 1225976875.
- ↑ "Special Issue: Membrane Protein Simulations and Free Energy Approaches: In honor of Professor Benoit Roux's 60th birthday". Journal of Computational Chemistry. 41 (5): C1, 379–481. February 15, 2020. Retrieved 30 June 2020.
- ↑ "Past Award Winners". Royal Society of Canada. 21 October 2018. Retrieved May 11, 2019.
- ↑ Christoff, Craig (April 19, 2017). "2017 Fellow of the BSC: Dr. Benoît Roux". Biophysical Society of Canada. Retrieved May 11, 2019.
- ↑ Wood, Matt (2021-09-08). "Benoit Roux, PhD, elected to the Royal Society of Canada". Science Life | Biological Sciences Division Research News. Retrieved 2021-09-13.
- ↑ "Two UChicago scholars elected as 2023 American Association for the Advancement of Science fellows". news.uchicago.edu. 2024-04-18. Retrieved 2024-04-18.