
A DNA virus is a virus that has a genome made of deoxyribonucleic acid (DNA) that is replicated by a DNA polymerase. They can be divided between those that have two strands of DNA in their genome, called double-stranded DNA (dsDNA) viruses, and those that have one strand of DNA in their genome, called single-stranded DNA (ssDNA) viruses. dsDNA viruses primarily belong to two realms: Duplodnaviria and Varidnaviria, and ssDNA viruses are almost exclusively assigned to the realm Monodnaviria, which also includes some dsDNA viruses. Additionally, many DNA viruses are unassigned to higher taxa. Reverse transcribing viruses, which have a DNA genome that is replicated through an RNA intermediate by a reverse transcriptase, are classified into the kingdom Pararnavirae in the realm Riboviria.

Ribonucleic acid (RNA) is a polymeric molecule that is essential for most biological functions, either by performing the function itself or by forming a template for the production of proteins. RNA and deoxyribonucleic acid (DNA) are nucleic acids. The nucleic acids constitute one of the four major macromolecules essential for all known forms of life. RNA is assembled as a chain of nucleotides. Cellular organisms use messenger RNA (mRNA) to convey genetic information that directs synthesis of specific proteins. Many viruses encode their genetic information using an RNA genome.

A retrovirus is a type of virus that inserts a DNA copy of its RNA genome into the DNA of a host cell that it invades, thus changing the genome of that cell. After invading a host cell's cytoplasm, the virus uses its own reverse transcriptase enzyme to produce DNA from its RNA genome, the reverse of the usual pattern, thus retro (backward). The new DNA is then incorporated into the host cell genome by an integrase enzyme, at which point the retroviral DNA is referred to as a provirus. The host cell then treats the viral DNA as part of its own genome, transcribing and translating the viral genes along with the cell's own genes, producing the proteins required to assemble new copies of the virus. Many retroviruses cause serious diseases in humans, other mammals, and birds.
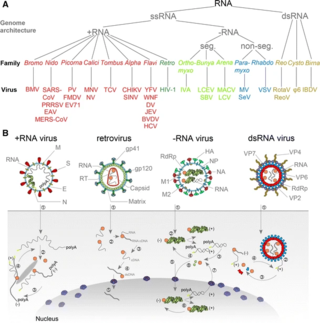
An RNA virus is a virus characterized by a ribonucleic acid (RNA) based genome. The genome can be single-stranded RNA (ssRNA) or double-stranded (dsRNA). Notable human diseases caused by RNA viruses include influenza, SARS, MERS, COVID-19, Dengue virus, hepatitis C, hepatitis E, West Nile fever, Ebola virus disease, rabies, polio, mumps, and measles.

A reverse transcriptase (RT) is an enzyme used to convert RNA genome to DNA, a process termed reverse transcription. Reverse transcriptases are used by viruses such as HIV, COVID-19, and hepatitis B to replicate their genomes, by retrotransposon mobile genetic elements to proliferate within the host genome, and by eukaryotic cells to extend the telomeres at the ends of their linear chromosomes. Contrary to a widely held belief, the process does not violate the flows of genetic information as described by the classical central dogma, as transfers of information from RNA to DNA are explicitly held possible.

Cystovirus is a genus of double-stranded RNA viruses which infects bacteria. It is the only genus in the family Cystoviridae. The name of the group cysto derives from Greek kystis which means bladder or sack. There are seven species in this genus.
Caulimoviridae is a family of viruses infecting plants. There are 94 species in this family, assigned to 11 genera. Viruses belonging to the family Caulimoviridae are termed double-stranded DNA (dsDNA) reverse-transcribing viruses i.e. viruses that contain a reverse transcription stage in their replication cycle. This family contains all plant viruses with a dsDNA genome that have a reverse transcribing phase in their lifecycle.

Polydnaviriformidae ( PDV) is a family of insect viriforms; members are known as polydnaviruses. There are two genera in the family: Bracoform and Ichnoviriform. Polydnaviruses form a symbiotic relationship with parasitoid wasps. Ichnoviriforms (IV) occur in Ichneumonid wasps and Bracoviriforms (BV) in Braconid wasps. The larvae of wasps in both of those groups are themselves parasitic on Lepidoptera, and the polydnaviruses are important in circumventing the immune response of their parasitized hosts. Little or no sequence homology exists between BV and IV, suggesting that the two genera have been evolving independently for a long time.

Gammaretrovirus is a genus in the Retroviridae family. Example species are the murine leukemia virus and the feline leukemia virus. They cause various sarcomas, leukemias and immune deficiencies in mammals, reptiles and birds.
Baltimore classification is a system used to classify viruses based on their manner of messenger RNA (mRNA) synthesis. By organizing viruses based on their manner of mRNA production, it is possible to study viruses that behave similarly as a distinct group. Seven Baltimore groups are described that take into consideration whether the viral genome is made of deoxyribonucleic acid (DNA) or ribonucleic acid (RNA), whether the genome is single- or double-stranded, and whether the sense of a single-stranded RNA genome is positive or negative.

The murine leukemia viruses are retroviruses named for their ability to cause cancer in murine (mouse) hosts. Some MLVs may infect other vertebrates. MLVs include both exogenous and endogenous viruses. Replicating MLVs have a positive sense, single-stranded RNA (ssRNA) genome that replicates through a DNA intermediate via the process of reverse transcription.
In molecular biology and genetics, the sense of a nucleic acid molecule, particularly of a strand of DNA or RNA, refers to the nature of the roles of the strand and its complement in specifying a sequence of amino acids. Depending on the context, sense may have slightly different meanings. For example, the negative-sense strand of DNA is equivalent to the template strand, whereas the positive-sense strand is the non-template strand whose nucleotide sequence is equivalent to the sequence of the mRNA transcript.

Protein kinase RNA-activated also known as protein kinase R (PKR), interferon-induced, double-stranded RNA-activated protein kinase, or eukaryotic translation initiation factor 2-alpha kinase 2 (EIF2AK2) is an enzyme that in humans is encoded by the EIF2AK2 gene on chromosome 2. PKR is a serine/tyrosine kinase that is 551 amino acids long.

RNA-dependent RNA polymerase (RdRp) or RNA replicase is an enzyme that catalyzes the replication of RNA from an RNA template. Specifically, it catalyzes synthesis of the RNA strand complementary to a given RNA template. This is in contrast to typical DNA-dependent RNA polymerases, which all organisms use to catalyze the transcription of RNA from a DNA template.

Double-stranded RNA viruses are a polyphyletic group of viruses that have double-stranded genomes made of ribonucleic acid. The double-stranded genome is used as a template by the viral RNA-dependent RNA polymerase (RdRp) to transcribe a positive-strand RNA functioning as messenger RNA (mRNA) for the host cell's ribosomes, which translate it into viral proteins. The positive-strand RNA can also be replicated by the RdRp to create a new double-stranded viral genome.

The double-stranded RNA-specific adenosine deaminase enzyme family are encoded by the ADAR family genes. ADAR stands for adenosine deaminase acting on RNA. This article focuses on the ADAR proteins; This article details the evolutionary history, structure, function, mechanisms and importance of all proteins within this family.
ICP8, the herpes simplex virus type-1 single-strand DNA-binding protein, is one of seven proteins encoded in the viral genome of HSV-1 that is required for HSV-1 DNA replication. It is able to anneal to single-stranded DNA (ssDNA) as well as melt small fragments of double-stranded DNA (dsDNA); its role is to destabilize duplex DNA during initiation of replication. It differs from helicases because it is ATP- and Mg2+-independent. In cells infected with HSV-1, the DNA in those cells become colocalized with ICP8.
Riboviria is a realm of viruses that includes all viruses that use a homologous RNA-dependent polymerase for replication. It includes RNA viruses that encode an RNA-dependent RNA polymerase, as well as reverse-transcribing viruses that encode an RNA-dependent DNA polymerase. RNA-dependent RNA polymerase (RdRp), also called RNA replicase, produces RNA from RNA. RNA-dependent DNA polymerase (RdDp), also called reverse transcriptase (RT), produces DNA from RNA. These enzymes are essential for replicating the viral genome and transcribing viral genes into messenger RNA (mRNA) for translation of viral proteins.

Ground squirrel hepatitis virus, abbreviated GSHV, is a partially double-stranded DNA virus that is closely related to human Hepatitis B virus (HBV) and Woodchuck hepatitis virus (WHV). It is a member of the family of viruses Hepadnaviridae and the genus Orthohepadnavirus. Like the other members of its family, GSHV has high degree of species and tissue specificity. It was discovered in Beechey ground squirrels, Spermophilus beecheyi, but also infects Arctic ground squirrels, Spermophilus parryi. Commonalities between GSHV and HBV include morphology, DNA polymerase activity in genome repair, cross-reacting viral antigens, and the resulting persistent infection with viral antigen in the blood (antigenemia). As a result, GSHV is used as an experimental model for HBV.

Orthornavirae is a kingdom of viruses that have genomes made of ribonucleic acid (RNA), including genes which encode an RNA-dependent RNA polymerase (RdRp). The RdRp is used to transcribe the viral RNA genome into messenger RNA (mRNA) and to replicate the genome. Viruses in this kingdom share a number of characteristics which promote rapid evolution, including high rates of genetic mutation, recombination, and reassortment.
















