In chemistry, a disulfide refers to a functional group with the structure R−S−S−R′. The linkage is also called an SS-bond or sometimes a disulfide bridge and is usually derived by the coupling of two thiol groups. In biology, disulfide bridges formed between thiol groups in two cysteine residues are an important component of the secondary and tertiary structure of proteins. Persulfide usually refers to R−S−S−H compounds.
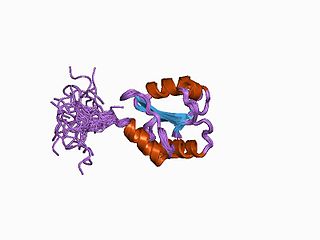
Protein disulfide isomerase, or PDI, is an enzyme in the endoplasmic reticulum (ER) in eukaryotes and the periplasm of bacteria that catalyzes the formation and breakage of disulfide bonds between cysteine residues within proteins as they fold. This allows proteins to quickly find the correct arrangement of disulfide bonds in their fully folded state, and therefore the enzyme acts to catalyze protein folding.

Thioredoxin is a class of small redox proteins known to be present in all organisms. It plays a role in many important biological processes, including redox signaling. In humans, thioredoxins are encoded by TXN and TXN2 genes. Loss-of-function mutation of either of the two human thioredoxin genes is lethal at the four-cell stage of the developing embryo. Although not entirely understood, thioredoxin plays a central role in humans and is increasingly linked to medicine through their response to reactive oxygen species (ROS). In plants, thioredoxins regulate a spectrum of critical functions, ranging from photosynthesis to growth, flowering and the development and germination of seeds. They have also recently been found to play a role in cell-to-cell communication.

Dithiothreitol (DTT) is the common name for a small-molecule redox reagent also known as Cleland's reagent. DTT's formula is C4H10O2S2 and the chemical structure of one of its enantiomers in its reduced form is shown on the right; its oxidized form is a disulfide bonded 6-membered ring (shown below). The reagent is commonly used in its racemic form, as both enantiomers are reactive. Its name derives from the four-carbon sugar, threose. DTT has an epimeric ('sister') compound, dithioerythritol (DTE).
Oxidative protein folding is a process that is responsible for the formation of disulfide bonds between cysteine residues in proteins. The driving force behind this process is a redox reaction, in which electrons pass between several proteins and finally to a terminal electron acceptor.

The thioredoxin fold is a protein fold common to enzymes that catalyze disulfide bond formation and isomerization. The fold is named for the canonical example thioredoxin and is found in both prokaryotic and eukaryotic proteins. It is an example of an alpha/beta protein fold that has oxidoreductase activity. The fold's spatial topology consists of a four-stranded antiparallel beta sheet sandwiched between three alpha helices. The strand topology is 2134 with 3 antiparallel to the rest.

ER oxidoreductin 1 (Ero1) is an oxidoreductase enzyme that catalyses the formation and isomerization of protein disulfide bonds in the endoplasmic reticulum (ER) of eukaryotes. ER Oxidoreductin 1 (Ero1) is a conserved, luminal, glycoprotein that is tightly associated with the ER membrane, and is essential for the oxidation of protein dithiols. Since disulfide bond formation is an oxidative process, the major pathway of its catalysis has evolved to utilise oxidoreductases, which become reduced during the thiol-disulfide exchange reactions that oxidise the cysteine thiol groups of nascent polypeptides. Ero1 is required for the introduction of oxidising equivalents into the ER and their direct transfer to protein disulfide isomerase (PDI), thereby ensuring the correct folding and assembly of proteins that contain disulfide bonds in their native state.

Glutaredoxins are small redox enzymes of approximately one hundred amino-acid residues that use glutathione as a cofactor. In humans this oxidation repair enzyme is also known to participate in many cellular functions, including redox signaling and regulation of glucose metabolism. Glutaredoxins are oxidized by substrates, and reduced non-enzymatically by glutathione. In contrast to thioredoxins, which are reduced by thioredoxin reductase, no oxidoreductase exists that specifically reduces glutaredoxins. Instead, glutaredoxins are reduced by the oxidation of glutathione. Oxidized glutathione is then regenerated by glutathione reductase. Together these components compose the glutathione system.
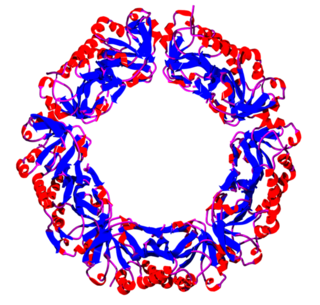
Peroxiredoxins are a ubiquitous family of antioxidant enzymes that also control cytokine-induced peroxide levels and thereby mediate signal transduction in mammalian cells. The family members in humans are PRDX1, PRDX2, PRDX3, PRDX4, PRDX5, and PRDX6. The physiological importance of peroxiredoxins is illustrated by their relative abundance.
Protein disulfide-isomerase A3 (PDIA3), also known as glucose-regulated protein, 58-kD (GRP58), is an isomerase enzyme. This protein localizes to the endoplasmic reticulum (ER) and interacts with lectin chaperones calreticulin and calnexin (CNX) to modulate folding of newly synthesized glycoproteins. It is thought that complexes of lectins and this protein mediate protein folding by promoting formation of disulfide bonds in their glycoprotein substrates.
Betaine reductase is an enzyme that catalyzes the chemical reaction
In enzymology, a glycine reductase (EC 1.21.4.2) is an enzyme that catalyzes the chemical reaction

Isopenicillin N synthase (IPNS) is a non-heme iron protein belongig to the 2-oxoglutarate (2OG)-dependent dioxygenases oxidoreductase family. This enzyme catalyzes the formation of isopenicillin N from δ-(L-α-aminoadipoyl)-L-cysteinyl-D-valine (LLD-ACV).
Adenylyl-sulfate reductase (thioredoxin) is an enzyme that catalyzes the chemical reaction
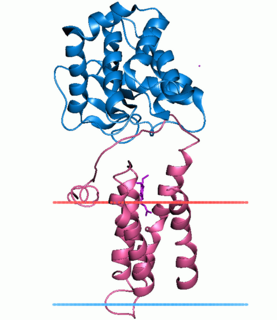
Disulfide bond formation protein B (DsbB) is a protein component of the pathway that leads to disulfide bond formation in periplasmic proteins of Escherichia coli and other bacteria. In Bacillus subtilis it is known as BdbC.

Bacterial glutathione transferases are part of a superfamily of enzymes that play a crucial role in cellular detoxification. The primary role of GSTs is to catalyze the conjugation of glutathione (GSH) with the electrophilic centers of a wide variety of molecules. The most commonly known substrates of GSTs are xenobiotic synthetic chemicals. There are also classes of GSTs that utilize glutathione as a cofactor rather than a substrate. Often these GSTs are involved in reduction of reactive oxidative species toxic to the bacterium. Conjugation with glutathione receptors reders toxic substances more soluble, and therefore more readily exocytosed from the cell.
Thioredoxins are small disulfide-containing redox proteins that have been found in all the kingdoms of living organisms. Thioredoxin serves as a general protein disulfide oxidoreductase. It interacts with a broad range of proteins by a redox mechanism based on reversible oxidation of 2 cysteine thiol groups to a disulfide, accompanied by the transfer of 2 electrons and 2 protons. The net result is the covalent interconversion of a disulfide and a dithiol.

DsbC is a prokaryotic disulfide bond isomerase. The formation of native disulfide bonds play an important role in the proper folding of proteins and stabilize tertiary structures of the protein. DsbC is one of 6 proteins in the Dsb family in prokaryotes. The other proteins are DsbA, DsbB, DsbD, DsbE and DsbG. These enzymes work in tandem with each other to form disulfide bonds during the expression of proteins. DsbC and DsbG act as proofreaders of the disulfide bonds that are formed. They break non-native disulfide bonds that were formed and act as chaperones for the formation of native disulfide bonds. The isomerization of disulfide bonds occurs in the periplasm.

Ferredoxin-thioredoxin reductase EC 1.8.7.2, systematic name ferredoxin:thioredoxin disulfide oxidoreductase, is a [4Fe-4S] protein that plays an important role in the ferredoxin/thioredoxin regulatory chain. It catalyzes the following reaction:
The Disulfide bond oxidoreductase D (DsbD) family is a member of the Lysine Exporter (LysE) Superfamily. A representative list of proteins belonging to the DsbD family can be found in the Transporter Classification Base.













