Related Research Articles
An intron is any nucleotide sequence within a gene that is not expressed or operative in the final RNA product. The word intron is derived from the term intragenic region, i.e., a region inside a gene. The term intron refers to both the DNA sequence within a gene and the corresponding RNA sequence in RNA transcripts. The non-intron sequences that become joined by this RNA processing to form the mature RNA are called exons.

RNA splicing is a process in molecular biology where a newly-made precursor messenger RNA (pre-mRNA) transcript is transformed into a mature messenger RNA (mRNA). It works by removing all the introns and splicing back together exons. For nuclear-encoded genes, splicing occurs in the nucleus either during or immediately after transcription. For those eukaryotic genes that contain introns, splicing is usually needed to create an mRNA molecule that can be translated into protein. For many eukaryotic introns, splicing occurs in a series of reactions which are catalyzed by the spliceosome, a complex of small nuclear ribonucleoproteins (snRNPs). There exist self-splicing introns, that is, ribozymes that can catalyze their own excision from their parent RNA molecule. The process of transcription, splicing and translation is called gene expression, the central dogma of molecular biology.

Alternative splicing, or alternative RNA splicing, or differential splicing, is an alternative splicing process during gene expression that allows a single gene to code for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene. This means the exons are joined in different combinations, leading to different (alternative) mRNA strands. Consequently, the proteins translated from alternatively spliced mRNAs usually contain differences in their amino acid sequence and, often, in their biological functions.

A spliceosome is a large ribonucleoprotein (RNP) complex found primarily within the nucleus of eukaryotic cells. The spliceosome is assembled from small nuclear RNAs (snRNA) and numerous proteins. Small nuclear RNA (snRNA) molecules bind to specific proteins to form a small nuclear ribonucleoprotein complex, which in turn combines with other snRNPs to form a large ribonucleoprotein complex called a spliceosome. The spliceosome removes introns from a transcribed pre-mRNA, a type of primary transcript. This process is generally referred to as splicing. An analogy is a film editor, who selectively cuts out irrelevant or incorrect material from the initial film and sends the cleaned-up version to the director for the final cut.

SR proteins are a conserved family of proteins involved in RNA splicing. SR proteins are named because they contain a protein domain with long repeats of serine and arginine amino acid residues, whose standard abbreviations are "S" and "R" respectively. SR proteins are ~200-600 amino acids in length and composed of two domains, the RNA recognition motif (RRM) region and the RS domain. SR proteins are more commonly found in the nucleus than the cytoplasm, but several SR proteins are known to shuttle between the nucleus and the cytoplasm.

A primary transcript is the single-stranded ribonucleic acid (RNA) product synthesized by transcription of DNA, and processed to yield various mature RNA products such as mRNAs, tRNAs, and rRNAs. The primary transcripts designated to be mRNAs are modified in preparation for translation. For example, a precursor mRNA (pre-mRNA) is a type of primary transcript that becomes a messenger RNA (mRNA) after processing.
RNA-binding proteins are proteins that bind to the double or single stranded RNA in cells and participate in forming ribonucleoprotein complexes. RBPs contain various structural motifs, such as RNA recognition motif (RRM), dsRNA binding domain, zinc finger and others. They are cytoplasmic and nuclear proteins. However, since most mature RNA is exported from the nucleus relatively quickly, most RBPs in the nucleus exist as complexes of protein and pre-mRNA called heterogeneous ribonucleoprotein particles (hnRNPs). RBPs have crucial roles in various cellular processes such as: cellular function, transport and localization. They especially play a major role in post-transcriptional control of RNAs, such as: splicing, polyadenylation, mRNA stabilization, mRNA localization and translation. Eukaryotic cells express diverse RBPs with unique RNA-binding activity and protein–protein interaction. According to the Eukaryotic RBP Database (EuRBPDB), there are 2961 genes encoding RBPs in humans. During evolution, the diversity of RBPs greatly increased with the increase in the number of introns. Diversity enabled eukaryotic cells to utilize RNA exons in various arrangements, giving rise to a unique RNP (ribonucleoprotein) for each RNA. Although RBPs have a crucial role in post-transcriptional regulation in gene expression, relatively few RBPs have been studied systematically.It has now become clear that RNA–RBP interactions play important roles in many biological processes among organisms.
Small nuclear RNA (snRNA) is a class of small RNA molecules that are found within the splicing speckles and Cajal bodies of the cell nucleus in eukaryotic cells. The length of an average snRNA is approximately 150 nucleotides. They are transcribed by either RNA polymerase II or RNA polymerase III. Their primary function is in the processing of pre-messenger RNA (hnRNA) in the nucleus. They have also been shown to aid in the regulation of transcription factors or RNA polymerase II, and maintaining the telomeres.
Gideon Dreyfuss is an American biochemist who is the Isaac Norris Professor of Biochemistry and Biophysics at the University of Pennsylvania School of Medicine and an investigator of the Howard Hughes Medical Institute. He was elected to the National Academy of Sciences in 2012.
Heterogeneous nuclear ribonucleoproteins (hnRNPs) are complexes of RNA and protein present in the cell nucleus during gene transcription and subsequent post-transcriptional modification of the newly synthesized RNA (pre-mRNA). The presence of the proteins bound to a pre-mRNA molecule serves as a signal that the pre-mRNA is not yet fully processed and therefore not ready for export to the cytoplasm. Since most mature RNA is exported from the nucleus relatively quickly, most RNA-binding protein in the nucleus exist as heterogeneous ribonucleoprotein particles. After splicing has occurred, the proteins remain bound to spliced introns and target them for degradation.
An exonic splicing silencer (ESS) is a short region of an exon and is a cis-regulatory element. A set of 103 hexanucleotides known as FAS-hex3 has been shown to be abundant in ESS regions. ESSs inhibit or silence splicing of the pre-mRNA and contribute to constitutive and alternate splicing. To elicit the silencing affect, ESSs recruit proteins that will negatively affect the core splicing machinery.

Survival of motor neuron or survival motor neuron (SMN) is a protein that in humans is encoded by the SMN1 and SMN2 genes.

Heterogeneous nuclear ribonucleoproteins A2/B1 is a protein that in humans is encoded by the HNRNPA2B1 gene.

Splicing factor, arginine/serine-rich 6 is a protein that in humans is encoded by the SFRS6 gene.

Polypyrimidine tract-binding protein 1 is a protein that in humans is encoded by the PTBP1 gene.

Serine/arginine-rich splicing factor 1 (SRSF1) also known as alternative splicing factor 1 (ASF1), pre-mRNA-splicing factor SF2 (SF2) or ASF1/SF2 is a protein that in humans is encoded by the SRSF1 gene. ASF/SF2 is an essential sequence specific splicing factor involved in pre-mRNA splicing. SRSF1 is the gene that codes for ASF/SF2 and is found on chromosome 17. The resulting splicing factor is a protein of approximately 33 kDa. ASF/SF2 is necessary for all splicing reactions to occur, and influences splice site selection in a concentration-dependent manner, resulting in alternative splicing. In addition to being involved in the splicing process, ASF/SF2 also mediates post-splicing activities, such as mRNA nuclear export and translation.

Survival of motor neuron 2 (SMN2) is a gene that encodes the SMN protein in humans.
Splicing regulatory element (SRE) are cis-acting sequences in pre-mRNA, which either enhance or silence (suppress) the splicing of introns, or in general regulates the constitutive or alternative splicing of this pre-mRNA. SREs recruit trans-acting splicing factors to activate or suppress the splice site recognition or spliceosome assembly. The "context dependence" of SREs is categorized into at least two studied groups: (a) the location-dependent activity of SREs: the activity varies with the relative positions of SREs in pre-mRNA; (b) the gene-dependent activity of SREs: the SRE activity observed in one gene is lost when the SRE is moved to another gene.
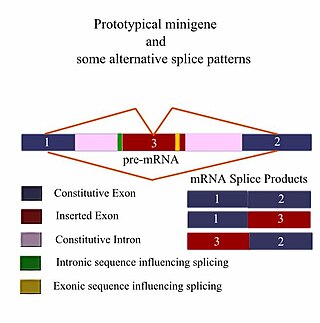
A minigene is a minimal gene fragment that includes an exon and the control regions necessary for the gene to express itself in the same way as a wild type gene fragment. This is a minigene in its most basic sense. More complex minigenes can be constructed containing multiple exons and intron(s). Minigenes provide a valuable tool for researchers evaluating splicing patterns both in vivo and in vitro biochemically assessed experiments. Specifically, minigenes are used as splice reporter vectors and act as a probe to determine which factors are important in splicing outcomes. They can be constructed to test the way both cis-regulatory elements and trans-regulatory elements affect gene expression.

Prp8 refers to both the Prp8 protein and Prp8 gene. Prp8's name originates from its involvement in pre-mRNA processing. The Prp8 protein is a large, highly conserved, and unique protein that resides in the catalytic core of the spliceosome and has been found to have a central role in molecular rearrangements that occur there. Prp8 protein is a major central component of the catalytic core in the spliceosome, and the spliceosome is responsible for splicing of precursor mRNA that contains introns and exons. Unexpressed introns are removed by the spliceosome complex in order to create a more concise mRNA transcript. Splicing is just one of many different post-transcriptional modifications that mRNA must undergo before translation. Prp8 has also been hypothesized to be a cofactor in RNA catalysis.
References
- ↑ Zhu, Jun; Mayeda, Akila; Krainer, Adrian R. (December 2001). "Exon Identity Established through Differential Antagonism between Exonic Splicing Silencer-Bound hnRNP A1 and Enhancer-Bound SR Proteins". Molecular Cell. 8 (6): 1351–1361. doi: 10.1016/S1097-2765(01)00409-9 . PMID 11779509.
- ↑ Cartegni, Luca; Chew, Shern L.; Krainer, Adrian R. (1 April 2002). "Listening to silence and understanding nonsense: exonic mutations that affect splicing". Nature Reviews Genetics. 3 (4): 285–298. doi:10.1038/nrg775. PMID 11967553. S2CID 15307589.
- ↑ Fairbrother, William G.; Yeo, Gene W.; Yeh, Rufang; Goldstein, Paul; Mawson, Matthew; Sharp, Phillip A.; Burge, Christopher B. (2004-07-01). "RESCUE-ESE identifies candidate exonic splicing enhancers in vertebrate exons". Nucleic Acids Research. 32 (Web Server issue): W187–W190. doi:10.1093/nar/gkh393. ISSN 0305-1048. PMC 441531 . PMID 15215377.
- ↑ Fairbrother WG, Yeh RF, Sharp PA, Burge CB (August 2002). "Predictive identification of exonic splicing enhancers in human genes". Science. 297 (5583): 1007–13. doi: 10.1126/science.1073774 . PMID 12114529.
- ↑ McVety, S; Li, L; Gordon, P H; Chong, G; Foulkes, W D (17 June 2005). "Disruption of an exon splicing enhancer in exon 3 of MLH1 is the cause of HNPCC in a Quebec family". Journal of Medical Genetics. 43 (2): 153–156. doi:10.1136/jmg.2005.031997. PMC 2564635 . PMID 15923275.
- ↑ Carlini, David B.; Genut, Jordan E. (30 November 2005). "Synonymous SNPs Provide Evidence for Selective Constraint on Human Exonic Splicing Enhancers". Journal of Molecular Evolution. 62 (1): 89–98. doi:10.1007/s00239-005-0055-x. PMID 16320116. S2CID 30031983.