
Ubiquitin is a small (8.6 kDa) regulatory protein found in most tissues of eukaryotic organisms, i.e., it is found ubiquitously. It was discovered in 1975 by Gideon Goldstein and further characterized throughout the late 1970s and 1980s. Four genes in the human genome code for ubiquitin: UBB, UBC, UBA52 and RPS27A.

A ubiquitin ligase is a protein that recruits an E2 ubiquitin-conjugating enzyme that has been loaded with ubiquitin, recognizes a protein substrate, and assists or directly catalyzes the transfer of ubiquitin from the E2 to the protein substrate. In simple and more general terms, the ligase enables movement of ubiquitin from a ubiquitin carrier to another protein by some mechanism. The ubiquitin, once it reaches its destination, ends up being attached by an isopeptide bond to a lysine residue, which is part of the target protein. E3 ligases interact with both the target protein and the E2 enzyme, and so impart substrate specificity to the E2. Commonly, E3s polyubiquitinate their substrate with Lys48-linked chains of ubiquitin, targeting the substrate for destruction by the proteasome. However, many other types of linkages are possible and alter a protein's activity, interactions, or localization. Ubiquitination by E3 ligases regulates diverse areas such as cell trafficking, DNA repair, and signaling and is of profound importance in cell biology. E3 ligases are also key players in cell cycle control, mediating the degradation of cyclins, as well as cyclin dependent kinase inhibitor proteins. The human genome encodes over 600 putative E3 ligases, allowing for tremendous diversity in substrates.

Ubiquitin-like modifier activating enzyme 1 (UBA1) is an enzyme which in humans is encoded by the UBA1 gene. UBA1 participates in ubiquitination and the NEDD8 pathway for protein folding and degradation, among many other biological processes. This protein has been linked to X-linked spinal muscular atrophy type 2, neurodegenerative diseases, and cancers.

Ubiquitin-protein ligase E3A (UBE3A) also known as E6AP ubiquitin-protein ligase (E6AP) is an enzyme that in humans is encoded by the UBE3A gene. This enzyme is involved in targeting proteins for degradation within cells.

An isopeptide bond is a type of amide bond formed between a carboxyl group of one amino acid and an amino group of another. An isopeptide bond is the linkage between the side chain amino or carboxyl group of one amino acid to the α-carboxyl, α-amino group, or the side chain of another amino acid. In a typical peptide bond, also known as eupeptide bond, the amide bond always forms between the α-carboxyl group of one amino acid and the α-amino group of the second amino acid. Isopeptide bonds are rarer than regular peptide bonds. Isopeptide bonds lead to branching in the primary sequence of a protein. Proteins formed from normal peptide bonds typically have a linear primary sequence.

ITCH is a HECT domain–containing E3 ubiquitin ligase that is ablated in non-agouti-lethal 18H mice. Itchy mice develop a severe immunological phenotype after birth that includes hyperplasia of lymphoid and hematopoietic cells, and stomach and lung inflammation. In humans ITCH deficiency causes altered physical growth, craniofacial morphology defects, defective muscle development, and aberrant immune system function. The ITCH gene is located on chromosome 20 in humans. ITCH contains a C2 domain, proline-rich region, WW domains, HECT domain, and multiple amino acids that are phosphorylated and ubiquitinated.

E3 ubiquitin-protein ligase NEDD4, also known as neural precursor cell expressed developmentally down-regulated protein 4 is an enzyme that is, in humans, encoded by the NEDD4 gene.

Neural precursor cell expressed developmentally downregulated gene 4-like (NEDD4L) or NEDD4-2 is an enzyme of the NEDD4 family. In human the protein is encoded by the NEDD4L gene. In mouse the protein is commonly known as NEDD4-2 and the gene Nedd4-2.

Ubiquitin-conjugating enzyme E2 L3 (UBE2L3), also called UBCH7, is a protein that in humans is encoded by the UBE2L3 gene. As an E2 enzyme, UBE2L3 participates in ubiquitination to target proteins for degradation. The role of UBE2L3 in the ubiquitination of the NF-κB precursor implicated it in various major autoimmune diseases, including rheumatoid arthritis (RA), celiac disease, Crohn's disease (CD), and systemic lupus erythematosus.

E3 ubiquitin-protein ligase SMURF1 is an enzyme that in humans is encoded by the SMURF1 gene. The SMURF1 Gene encodes a protein with a size of 757 amino acids and the molecular mass of this protein is 86114 Da.

Ubiquitin D is a protein that in humans is encoded by the UBD gene, also known as FAT10. UBD acts like ubiquitin, by covalently modifying proteins and tagging them for destruction in the proteasome.

Ubiquitin-conjugating enzyme E2 D1 is a protein that in humans is encoded by the UBE2D1 gene.

Ubiquitin-conjugating enzyme E2 D2 is a protein that in humans is encoded by the UBE2D2 gene.
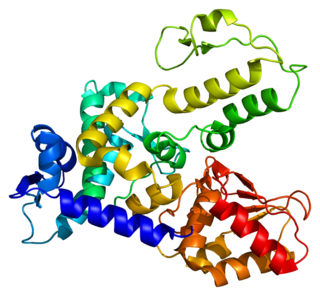
NEDD4-like E3 ubiquitin-protein ligase WWP1 is an enzyme that in humans is encoded by the WWP1 gene.

E3 ubiquitin-protein ligase UBR5 is an enzyme that in humans is encoded by the UBR5 gene.

E3 ubiquitin-protein ligase SMURF2 is an enzyme that in humans is encoded by the SMURF2 gene which is located at chromosome 17q23.3-q24.1.

Ubiquitin/ISG15-conjugating enzyme E2 L6 is a protein that in humans is encoded by the UBE2L6 gene.

WW domain-binding protein 2 is a protein that in humans is encoded by the WBP2 gene.

E3 ISG15–protein ligase HERC5 is an enzyme that in humans is encoded by the HERC5 gene.
In molecular biology, the Ubiquitin-Interacting Motif (UIM), or 'LALAL-motif', is a sequence motif of about 20 amino acid residues, which was first described in the 26S proteasome subunit PSD4/RPN-10 that is known to recognise ubiquitin. In addition, the UIM is found, often in tandem or triplet arrays, in a variety of proteins either involved in ubiquitination and ubiquitin metabolism, or known to interact with ubiquitin-like modifiers. Among the UIM proteins are two different subgroups of the UBP family of deubiquitinating enzymes, one F-box protein, one family of HECT-containing ubiquitin-ligases (E3s) from plants, and several proteins containing ubiquitin-associated UBA and/or UBX domains. In most of these proteins, the UIM occurs in multiple copies and in association with other domains such as UBA, UBX, ENTH domain, EH, VHS, SH3 domain, HECT, VWFA, EF-hand calcium-binding, WD-40, F-box, LIM, protein kinase, ankyrin, PX, phosphatidylinositol 3- and 4-kinase, C2 domain, OTU, DnaJ domain, RING-finger or FYVE-finger. UIMs have been shown to bind ubiquitin and to serve as a specific targeting signal important for monoubiquitination. Thus, UIMs may have several functions in ubiquitin metabolism each of which may require different numbers of UIMs.




















