Related Research Articles

An alpha helix is a sequence of amino acids in a protein that are twisted into a coil.

The beta sheet, (β-sheet) is a common motif of the regular protein secondary structure. Beta sheets consist of beta strands (β-strands) connected laterally by at least two or three backbone hydrogen bonds, forming a generally twisted, pleated sheet. A β-strand is a stretch of polypeptide chain typically 3 to 10 amino acids long with backbone in an extended conformation. The supramolecular association of β-sheets has been implicated in the formation of the fibrils and protein aggregates observed in amyloidosis, Alzheimer's disease and other proteinopathies.

Protein secondary structure is the local spatial conformation of the polypeptide backbone excluding the side chains. The two most common secondary structural elements are alpha helices and beta sheets, though beta turns and omega loops occur as well. Secondary structure elements typically spontaneously form as an intermediate before the protein folds into its three dimensional tertiary structure.
A coiled coil is a structural motif in proteins in which 2–7 alpha-helices are coiled together like the strands of a rope. Many coiled coil-type proteins are involved in important biological functions, such as the regulation of gene expression — e.g., transcription factors. Notable examples are the oncoproteins c-Fos and c-Jun, as well as the muscle protein tropomyosin.

Helix-turn-helix is a DNA-binding protein (DBP). The helix-turn-helix (HTH) is a major structural motif capable of binding DNA. Each monomer incorporates two α helices, joined by a short strand of amino acids, that bind to the major groove of DNA. The HTH motif occurs in many proteins that regulate gene expression. It should not be confused with the helix–loop–helix motif.

A leucine zipper is a common three-dimensional structural motif in proteins. They were first described by Landschulz and collaborators in 1988 when they found that an enhancer binding protein had a very characteristic 30-amino acid segment and the display of these amino acid sequences on an idealized alpha helix revealed a periodic repetition of leucine residues at every seventh position over a distance covering eight helical turns. The polypeptide segments containing these periodic arrays of leucine residues were proposed to exist in an alpha-helical conformation and the leucine side chains from one alpha helix interdigitate with those from the alpha helix of a second polypeptide, facilitating dimerization.
A DNA-binding domain (DBD) is an independently folded protein domain that contains at least one structural motif that recognizes double- or single-stranded DNA. A DBD can recognize a specific DNA sequence or have a general affinity to DNA. Some DNA-binding domains may also include nucleic acids in their folded structure.

A pi helix is a type of secondary structure found in proteins. Discovered by crystallographer Barbara Low in 1952 and once thought to be rare, short π-helices are found in 15% of known protein structures and are believed to be an evolutionary adaptation derived by the insertion of a single amino acid into an α-helix. Because such insertions are highly destabilizing, the formation of π-helices would tend to be selected against unless it provided some functional advantage to the protein. π-helices therefore are typically found near functional sites of proteins.

A 310 helix is a type of secondary structure found in proteins and polypeptides. Of the numerous protein secondary structures present, the 310-helix is the fourth most common type observed; following α-helices, β-sheets and reverse turns. 310-helices constitute nearly 10–15% of all helices in protein secondary structures, and are typically observed as extensions of α-helices found at either their N- or C- termini. Because of the α-helices tendency to consistently fold and unfold, it has been proposed that the 310-helix serves as an intermediary conformation of sorts, and provides insight into the initiation of α-helix folding.
A helix bundle is a small protein fold composed of several alpha helices that are usually nearly parallel or antiparallel to each other.
In polymer science, the Lifson–Roig model is a helix-coil transition model applied to the alpha helix-random coil transition of polypeptides; it is a refinement of the Zimm–Bragg model that recognizes that a polypeptide alpha helix is only stabilized by a hydrogen bond only once three consecutive residues have adopted the helical conformation. To consider three consecutive residues each with two states, the Lifson–Roig model uses a 4x4 transfer matrix instead of the 2x2 transfer matrix of the Zimm–Bragg model, which considers only two consecutive residues. However, the simple nature of the coil state allows this to be reduced to a 3x3 matrix for most applications.
β turns are the most common form of turns—a type of non-regular secondary structure in proteins that cause a change in direction of the polypeptide chain. They are very common motifs in proteins and polypeptides. Each consists of four amino acid residues. They can be defined in two ways:
- By the possession of an intra-main-chain hydrogen bond between the CO of residue i and the NH of residue i+3;
- By having a distance of less than 7Å between the Cα atoms of residues i and i+3.

Schellman loops are commonly occurring structural features of proteins and polypeptides. Each has six amino acid residues with two specific inter-mainchain hydrogen bonds and a characteristic main chain dihedral angle conformation. The CO group of residue i is hydrogen-bonded to the NH of residue i+5, and the CO group of residue i+1 is hydrogen-bonded to the NH of residue i+4. Residues i+1, i+2, and i+3 have negative φ (phi) angle values and the phi value of residue i+4 is positive. Schellman loops incorporate a three amino acid residue RL nest, in which three mainchain NH groups form a concavity for hydrogen bonding to carbonyl oxygens. About 2.5% of amino acids in proteins belong to Schellman loops. Two websites are available for examining small motifs in proteins, Motivated Proteins: ; or PDBeMotif:.
The Asx turn is a structural feature in proteins and polypeptides. It consists of three amino acid residues in which residue i is an aspartate (Asp) or asparagine (Asn) that forms a hydrogen bond from its sidechain CO group to the mainchain NH group of residue i+2. About 14% of Asx residues present in proteins belong to Asx turns.
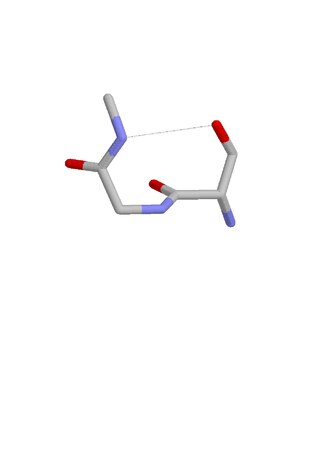
The ST turn is a structural feature in proteins and polypeptides. Each consists of three amino acid residues in which residue i is a serine (S) or threonine (T) that forms a hydrogen bond from its sidechain oxygen group to the mainchain NH group of residue i + 2.
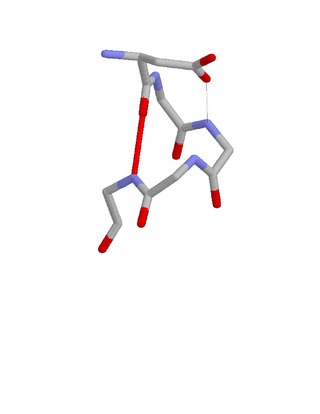
The Asx motif is a commonly occurring feature in proteins and polypeptides. It consists of four or five amino acid residues with either aspartate or asparagine as the first residue. It is defined by two internal hydrogen bonds. One is between the side chain oxygen of residue i and the main chain NH of residue i+2 or i+3; the other is between the main chain oxygen of residue i and the main chain NH of residue i+3 or i+4. Asx motifs occur commonly in proteins and polypeptides.

The ST motif is a commonly occurring feature in proteins and polypeptides. It consists of four or five amino acid residues with either serine or threonine as the first residue. It is defined by two internal hydrogen bonds. One is between the side chain oxygen of residue i and the main chain NH of residue i + 2 or i + 3; the other is between the main chain oxygen of residue i and the main chain NH of residue i + 3 or i + 4. Two websites are available for finding and examining ST motifs in proteins, Motivated Proteins: and PDBeMotif.
The term C cap describes an amino acid in a particular position within a protein or polypeptide. The C cap residue of an alpha helix is the last amino acid residue at the C terminus of the helix. More precisely, it is defined as the last residue (i) whose NH group is hydrogen-bonded to the CO group of residue i-4. Because of this it is sometimes also described as the residue following the helix.
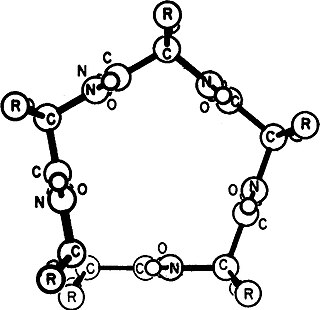
Gamma helix (or γ-helix) is a type of secondary structure in proteins that has been predicted by Pauling, Corey, and Branson, but has never been observed in natural proteins. The hydrogen bond in this type of helix was predicted to be between N-H group of one amino acid and the C=O group of the amino acid six residues earlier (or, as described by Pauling, Corey, Branson, "to the fifth amide group beyond it"). This can also be described as i + 6 → i bond and would be a continuation of the series (310 helix, alpha helix, pi helix and gamma helix). This theoretical helix contains 5.1 residues per turn.However, a fully developed gamma helix has characteristics of a structure that has 2.2 amino acid residues per turn, a rise of 2.75Å per residue, and a pseudo-cyclic (C7) structure closed by intramolecular H-bond. Depending on the amino acid's side chain (R) involved in this main-chain reversal motif, two stereoisomers can occur with their Cα-substituent located either in the axial or in the equatorial position relative to the H-bonded pseudo-cycle.

Transmembrane epididymal protein 1 is a transmembrane protein encoded by the TEDDM1 gene. TEDDM1 is also commonly known as TMEM45C and encodes 273 amino acids that contains six alpha-helix transmembrane regions. The protein contains a 118 amino acid length family of unknown function. While the exact function of TEDDM1 is not understood, it is predicted to be an integral component of the plasma membrane.
References
- ↑ Richardson, JM; Richardson DC (1988). "Amino acid preferences for specific locations at the ends of alpha-helices". Science. 240 (4859): 1648–1652. Bibcode:1988Sci...240.1648R. doi:10.1126/science.3381086. PMID 3381086. S2CID 38467101.
- ↑ Presta, LG; Rose GD (1988). "Helix Caps". Science. 240 (4859): 1632–1641. Bibcode:1988Sci...240.1632P. doi:10.1126/science.2837824. PMID 2837824.
- ↑ Doig, AJ; MacArthur MW (1997). "Structures of N-termini of helices in proteins". Protein Science. 6 (1): 147–155. doi:10.1002/pro.5560060117. PMC 2143508 . PMID 9007987.
- ↑ Aurora, R; Rose GD (1998). "Helix Capping". Protein Science. 7 (1): 21–38. doi:10.1002/pro.5560070103. PMC 2143812 . PMID 9514257.
- ↑ Gunasekaran, K; Nagarajam HA (1998). "Stereochemical punctuation marks in protein structure" (PDF). Journal of Molecular Biology. 275 (5): 917–932. doi:10.1006/jmbi.1997.1505. PMID 9480777.
- ↑ Leader, DP; Milner-White EJ (2011). "The structure of the ends of helices in globular proteins". Proteins. 79 (3): 1010–1019. doi:10.1002/prot.22942. PMID 21287629. S2CID 22240314.