
Oxidative phosphorylation or electron transport-linked phosphorylation or terminal oxidation is the metabolic pathway in which cells use enzymes to oxidize nutrients, thereby releasing chemical energy in order to produce adenosine triphosphate (ATP). In eukaryotes, this takes place inside mitochondria. Almost all aerobic organisms carry out oxidative phosphorylation. This pathway is so pervasive because it releases more energy than alternative fermentation processes such as anaerobic glycolysis.
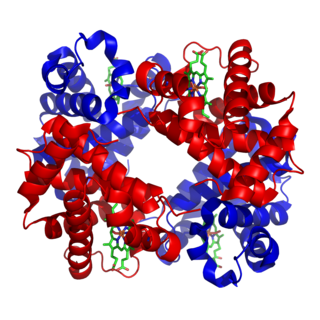
Metalloprotein is a generic term for a protein that contains a metal ion cofactor. A large proportion of all proteins are part of this category. For instance, at least 1000 human proteins contain zinc-binding protein domains although there may be up to 3000 human zinc metalloproteins.

In biology and biochemistry, the active site is the region of an enzyme where substrate molecules bind and undergo a chemical reaction. The active site consists of amino acid residues that form temporary bonds with the substrate, the binding site, and residues that catalyse a reaction of that substrate, the catalytic site. Although the active site occupies only ~10–20% of the volume of an enzyme, it is the most important part as it directly catalyzes the chemical reaction. It usually consists of three to four amino acids, while other amino acids within the protein are required to maintain the tertiary structure of the enzymes.

Photosystem I is one of two photosystems in the photosynthetic light reactions of algae, plants, and cyanobacteria. Photosystem I is an integral membrane protein complex that uses light energy to catalyze the transfer of electrons across the thylakoid membrane from plastocyanin to ferredoxin. Ultimately, the electrons that are transferred by Photosystem I are used to produce the moderate-energy hydrogen carrier NADPH. The photon energy absorbed by Photosystem I also produces a proton-motive force that is used to generate ATP. PSI is composed of more than 110 cofactors, significantly more than Photosystem II.
Iron–sulfur proteins are proteins characterized by the presence of iron–sulfur clusters containing sulfide-linked di-, tri-, and tetrairon centers in variable oxidation states. Iron–sulfur clusters are found in a variety of metalloproteins, such as the ferredoxins, as well as NADH dehydrogenase, hydrogenases, coenzyme Q – cytochrome c reductase, succinate – coenzyme Q reductase and nitrogenase. Iron–sulfur clusters are best known for their role in the oxidation-reduction reactions of electron transport in mitochondria and chloroplasts. Both Complex I and Complex II of oxidative phosphorylation have multiple Fe–S clusters. They have many other functions including catalysis as illustrated by aconitase, generation of radicals as illustrated by SAM-dependent enzymes, and as sulfur donors in the biosynthesis of lipoic acid and biotin. Additionally, some Fe–S proteins regulate gene expression. Fe–S proteins are vulnerable to attack by biogenic nitric oxide, forming dinitrosyl iron complexes. In most Fe–S proteins, the terminal ligands on Fe are thiolate, but exceptions exist.

The cytochrome b6f complex (plastoquinol/plastocyanin reductase or plastoquinol/plastocyanin oxidoreductase; EC 7.1.1.6) is an enzyme found in the thylakoid membrane in chloroplasts of plants, cyanobacteria, and green algae, that catalyzes the transfer of electrons from plastoquinol to plastocyanin:

A photosynthetic reaction center is a complex of several proteins, pigments, and other co-factors that together execute the primary energy conversion reactions of photosynthesis. Molecular excitations, either originating directly from sunlight or transferred as excitation energy via light-harvesting antenna systems, give rise to electron transfer reactions along the path of a series of protein-bound co-factors. These co-factors are light-absorbing molecules (also named chromophores or pigments) such as chlorophyll and pheophytin, as well as quinones. The energy of the photon is used to excite an electron of a pigment. The free energy created is then used, via a chain of nearby electron acceptors, for a transfer of hydrogen atoms (as protons and electrons) from H2O or hydrogen sulfide towards carbon dioxide, eventually producing glucose. These electron transfer steps ultimately result in the conversion of the energy of photons to chemical energy.
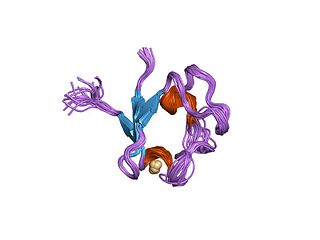
Rubredoxins are a class of low-molecular-weight iron-containing proteins found in sulfur-metabolizing bacteria and archaea. Sometimes rubredoxins are classified as iron-sulfur proteins; however, in contrast to iron-sulfur proteins, rubredoxins do not contain inorganic sulfide. Like cytochromes, ferredoxins and Rieske proteins, rubredoxins are thought to participate in electron transfer in biological systems. Recent work in bacteria and algae have led to the hypothesis that some rubredoxins may instead have a role in delivering iron to metalloproteins.
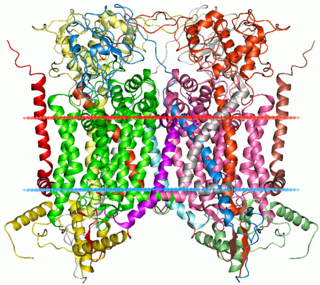
Cytochrome b within both molecular and cell biology, is a protein found in the membranes of aerobic cells. In eukaryotic mitochondria and in aerobic prokaryotes, cytochrome b is a component of respiratory chain complex III — also known as the bc1 complex or ubiquinol-cytochrome c reductase. In plant chloroplasts and cyanobacteria, there is an homologous protein, cytochrome b6, a component of the plastoquinone-plastocyanin reductase, also known as the b6f complex. These complexes are involved in electron transport, the pumping of protons to create a proton-motive force (PMF). This proton gradient is used for the generation of ATP. These complexes play a vital role in cells.
Copper proteins are proteins that contain one or more copper ions as prosthetic groups. Copper proteins are found in all forms of air-breathing life. These proteins are usually associated with electron-transfer with or without the involvement of oxygen (O2). Some organisms even use copper proteins to carry oxygen instead of iron proteins. A prominent copper protein in humans is in cytochrome c oxidase (cco). This enzyme cco mediates the controlled combustion that produces ATP. Other copper proteins include some superoxide dismutases used in defense against free radicals, peptidyl-α-monooxygenase for the production of hormones, and tyrosinase, which affects skin pigmentation.
Nitrite reductase refers to any of several classes of enzymes that catalyze the reduction of nitrite. There are two classes of NIR's. A multi haem enzyme reduces NO2− to a variety of products. Copper containing enzymes carry out a single electron transfer to produce nitric oxide.
P700, or photosystem I primary donor, is the reaction-center chlorophyll a molecular dimer associated with photosystem I in plants, algae, and cyanobacteria.

Stellacyanin is a member of the blue or type I copper protein family. This family of copper proteins is generally involved in electron transfer reactions with the Cu center transitioning between the oxidized Cu(II) form and the reduced Cu(I) form. Stellacyanin is ubiquitous among vascular seed plants.
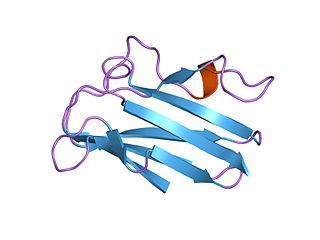
Plastocyanin/azurin family of copper-binding proteins is a family of small proteins that bind a single copper atom and that are characterised by an intense electronic absorption band near 600 nm. The most well-known members of this class of proteins are the plant chloroplastic plastocyanins, which exchange electrons with cytochrome c6, and the distantly related bacterial azurins, which exchange electrons with cytochrome c551. This family of proteins also includes amicyanin from bacteria such as Methylobacterium extorquens or Paracoccus versutus that can grow on methylamine; auracyanins A and B from Chloroflexus aurantiacus; blue copper protein from Alcaligenes faecalis; cupredoxin (CPC) from Cucumis sativus (Cucumber) peelings; cusacyanin from cucumber; halocyanin from Natronomonas pharaonis, a membrane-associated copper-binding protein; pseudoazurin from Pseudomonas; rusticyanin from Thiobacillus ferrooxidans; stellacyanin from Rhus vernicifera ; umecyanin from the roots of Armoracia rusticana (Horseradish); and allergen Ra3 from ragweed. This pollen protein has evolutary relation to the above proteins, but seems to have lost the ability to bind copper. Although there is an appreciable amount of divergence in the sequences of all these proteins, the copper ligand sites are conserved.

Azurin is a small, periplasmic, bacterial blue copper protein found in Pseudomonas, Bordetella, or Alcaligenes bacteria. Azurin moderates single-electron transfer between enzymes associated with the cytochrome chain by undergoing oxidation-reduction between Cu(I) and Cu(II). Each monomer of an azurin tetramer has a molecular weight of approximately 14kDa, contains a single copper atom, is intensively blue, and has a fluorescence emission band centered at 308 nm.
In bioinorganic chemistry, an entatic state is "a state of an atom or group which, due to its binding in a protein, has its geometric or electronic condition adapted for function." The term was coined by Bert Vallee and R. J. P. Williams, following work on the catalytic activity of carbonic anhydrase. These states are thought to enhance the chemistry of metal ions in biological catalysis.

Light-dependent reactions are certain photochemical reactions involved in photosynthesis, the main process by which plants acquire energy. There are two light dependent reactions: the first occurs at photosystem II (PSII) and the second occurs at photosystem I (PSI).
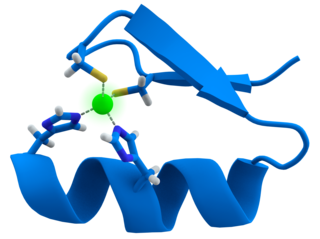
Transition metal thiolate complexes are metal complexes containing thiolate ligands. Thiolates are ligands that can be classified as soft Lewis bases. Therefore, thiolate ligands coordinate most strongly to metals that behave as soft Lewis acids as opposed to those that behave as hard Lewis acids. Most complexes contain other ligands in addition to thiolate, but many homoleptic complexes are known with only thiolate ligands. The amino acid cysteine has a thiol functional group, consequently many cofactors in proteins and enzymes feature cysteinate-metal cofactors.

Galactose oxidase is an enzyme that catalyzes the oxidation of D-galactose in some species of fungi.

Nickel superoxide dismutase (Ni-SOD) is a metalloenzyme that, like the other superoxide dismutases, protects cells from oxidative damage by catalyzing the disproportionation of the cytotoxic superoxide radical to hydrogen peroxide and molecular oxygen. Superoxide is a reactive oxygen species that is produced in large amounts during photosynthesis and aerobic cellular respiration. The equation for the disproportionation of superoxide is shown below:
















