
The SAM-II riboswitch is a RNA element found predominantly in alpha-proteobacteria that binds S-adenosyl methionine (SAM). Its structure and sequence appear to be unrelated to the SAM riboswitch found in Gram-positive bacteria. This SAM riboswitch is located upstream of the metA and metC genes in Agrobacterium tumefaciens, and other methionine and SAM biosynthesis genes in other alpha-proteobacteria. Like the other SAM riboswitch, it probably functions to turn off expression of these genes in response to elevated SAM levels. A significant variant of SAM-II riboswitches was found in Pelagibacter ubique and related marine bacteria and called SAM-V. Also, like many structured RNAs, SAM-II riboswitches can tolerate long loops between their stems.
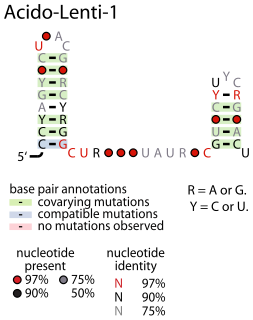
The Acido-Lenti-1 RNA motif describes a predicted non-coding RNA that is found in bacteria within the phyla acidobacteria and lentisphaerae. It is sometimes found nearby to group II introns, but the reason for this apparent association is unknown.

The Bacillaceae-1 RNA motif is a conserved RNA structure identified by bioinformatics within bacteria in the family bacillaceae. The RNA is presumed to operate as a non-coding RNA, and is sometimes adjacent to operons containing ribosomal RNAs. The most characteristic feature is two terminal loops that have the nucleotide consensus RUCCU, where R is either A or G. The motif might be related to the Desulfotalea-1 RNA motif, as the motifs share some similarity in conserved features, and the Desulfotalea-1 RNA motif is also sometimes adjacent to ribosomal RNA operons.
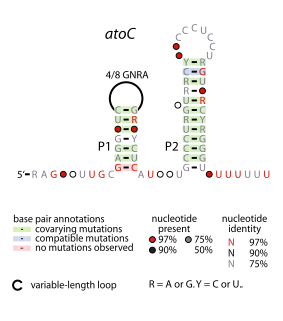
The atoC RNA motif is a conserved RNA-like structure identified by bioinformatics. It consistently appears upstream of protein-coding gene that are predicted to encode oxidoreductase activity, dihydropteroate synthase or DNA-binding response regulators.

The Chlorobi-1 RNA motif is a conserved RNA secondary structure identified by bioinformatics. It is predicted to be used only by Chlorobi, a phylum of bacteria. The motif consists of two stem-loops that are followed by an apparent rho-independent transcription terminator. The motif is presumed to function as an independently transcribed non-coding RNA.

The Chloroflexi-1 RNA motif is a conserved RNA structure detected by bioinformatics within the species Chloroflexus aggregans. C. aggregans has three predicted Chloroflexi-1 RNAs, which are located nearby to one another. This arrangement might suggest a repetitive element. C. aggregans is classified as belonging to the bacterial phylum Chloroflexi.

The Collinsella-1 RNA motif denotes a particular conserved RNA structure discovered by bioinformatics. Of the six sequences belonging to this motif that were originally identified, five are from uncultivated bacteria residing in the human gut, while only the sixth is in a cultivated species, Collinsella aerofaciens. The evidence supporting the stem-loops designated as "P1" and "P2" is ambiguous.

The Flavo-1 RNA motif is a conserved RNA structure that was identified by bioinformatics. The vast majority of Flavo-1 RNAs are found in Flavobacteria, but some were detected in the phylum Bacteroidetes, which contains Flavobacteria, or the phylum Spirochaetes, which is evolutionarily related to Bacteroidetes. It was presumed that Flavo-1 RNAs function as non-coding RNAs.

The Lacto-usp RNA motif is a conserved RNA structure identified in bacteria by bioinformatics. Lacto-usp RNAs are found exclusively in lactic acid bacteria, and exclusively in the possible 5′ untranslated regions of operons that contain a hypothetical gene and a usp gene. The usp gene encodes the universal stress protein. It was proposed that the Lacto-usp might correspond to the 6S RNA of the relevant species, because four of five of these species lack a predicted 6S RNA, and 6S RNAs commonly occur in 5′ UTRs of usp genes. However, given that the Lacto-usp RNA motif is much shorter than the standard 6S RNA structure, the function of Lacto-usp RNAs remains unclear.

The mraW RNA motif is a conserved, structured RNA found in certain bacteria. Specifically, it is predicted in many, though not all, species of actinobacteria, and especially within the genus Mycobacterium. Structurally, the motif consists of a hairpin with a highly conserved terminal loop sequence. mraW RNAs are consistently in the presumed 5' untranslated regions of mraW genes. These mraW genes likely form operons with immediately downstream ftsI genes, and multiple types of mur genes. These genes are associated with peptidoglycan synthesis, and it was hypothesized that the mraW RNA motif might regulate these genes.

The pan RNA motif defines a conserved RNA structure that was identified using bioinformatics. pan motif RNAs are present in three phyla: Chloroflexi, Firmicutes and Proteobacteria, although within the latter phylum they are only known in deltaproteobacteria. A pan RNA is present in the Firmicute Bacillus subtilis, which is one of the most extensively studied bacteria.

PhotoRC RNA motifs refer to conserved RNA structures that are associated with genes acting in the photosynthetic reaction centre of photosynthetic bacteria. Two such RNA classes were identified and called the PhotoRC-I and PhotoRC-II motifs. PhotoRC-I RNAs were detected in the genomes of some cyanobacteria. Although no PhotoRC-II RNA has been detected in cyanobacteria, one is found in the genome of a purified phage that infects cyanobacteria. Both PhotoRC-I and PhotoRC-II RNAs are present in sequences derived from DNA that was extracted from uncultivated marine bacteria.

The Polynucleobacter-1 RNA motif is a conserved RNA structure that was identified by bioinformatics. The RNA structure is predominantly located in genome sequences derived from DNA extracted from uncultivated marine samples. However it was also predicted in the genome of Polynucleobacter species QLW-P1DMWA-1, a kind of betaproteobacteria. The RNAs are often located near to a conserved gene that might be homologous to a gene found in a phage that infects cyanobacteria. However, it is unknown if the RNA is used by phages.

The radC RNA motif is a conserved RNA structure identified by bioinformatics. The radC RNA motif is found in certain bacteria where it is consistent located in the presumed 5' untranslated regions of genes whose encoded proteins bind DNA are interact with other proteins that bind DNA. These proteins include integrases, methyltransferases that might methylate DNA, proteins that inhibit restriction enzymes and radC genes. Although radC genes were thought to encode DNA repair proteins, this conclusion was based on mutation data that was later shown to affect a different gene. However, it is still possible that radC genes play some DNA-related role. No radC RNAs have been detected in any purified phage whose sequence was available as of 2010, although integrases are often used by phages.

The Rhizobiales-2 RNA motif is a set of RNAs found in certain bacteria that are presumed to be homologous because they conserve a common primary and secondary structure. The motif was discovered using bioinformatics, and is found only within bacteria that belong to the order Hyphomicrobiales, in turn a kind of alphaproteobacteria. Because Rhizobiales-2 RNAs are not consistently located in proximity to genes of a consistent class or function, these RNAs are presumed to function as non-coding RNAs.

The Ssbp, Topoisomerase, Antirestriction, XerDC Integrase RNA motif is a conserved RNA-like structure identified using bioinformatics. STAXI RNAs are located near to genes encoding proteins that interact with DNA or are associated with such proteins. This observation raised the possibility that instances of the STAXI motif function as single-stranded DNA molecules, perhaps during DNA replication or DNA repair. On the other hand, STAXI motifs often contain terminal loops conforming to the stable UNCG tetraloop, but the DNA version of this tetraloop (TNCG) is not especially stable. The STAXI motif consists of a simple pseudoknot structure that is repeated two or more times.

The sucA-II RNA motif is a conserved RNA structure identified by bioinformatics. It is consistently found in the presumed 5' untranslated regions of sucA genes, which encode Oxoglutarate dehydrogenase enzymes that participate in the citric acid cycle. Given this arrangement, sucA-II RNAs might regulate the downstream sucA gene. This genetic arrangement is similar to the previously reported sucA RNA motif. However, sucA-II RNAs are found only in bacteria classified within the genus Pseudomonas, whereas the previously reported motif is found only in betaproteobacteria.

The sucC RNA motif is a conserved RNA structure discovered using bioinformatics. sucC RNAs are found in the genus Pseudomonas, and are consistently found in possible 5' untranslated regions of sucC genes. These genes encode Succinyl coenzyme A synthetase, and are hypothesised to be regulated by the sucC RNAs. sucC genes participate in the citric acid cycle, and another gene involved in the citric acid cycle, sucA, is also predicted to be regulated by a conserved RNA structure.

The TwoAYGGAY RNA motif is a conserved RNA structure identified by bioinformatics. Its name refers to the conserved AYGGAY nucleotide sequence found in the motif's two terminal loops. The RNAs are found in sequences derived from DNA extracted from uncultivated bacteria present in the human gut, as well as some bacteria in the classes Clostridia and Gammaproteobacteria.

Yfr2 is a family of non-coding RNAs. Members of the Yrf2 family have been identified in almost all studied species of cyanobacteria. The family was identified through a bioinformatics screen of published cyanobacterial genomes, having previously been grouped in a family of Yfr2–5.




















