DNA repair protein XRCC3 is a protein that in humans is encoded by the XRCC3 gene. [5]
DNA repair protein XRCC3 is a protein that in humans is encoded by the XRCC3 gene. [5]
This gene encodes a member of the RecA/Rad51-related protein family that participates in homologous recombination to maintain chromosome stability and repair DNA damage. This gene functionally complements Chinese hamster irs1SF, a repair-deficient mutant that exhibits hypersensitivity to a number of different DNA-damaging agents and is chromosomally unstable. A rare microsatellite polymorphism in this gene is associated with cancer in patients of varying radiosensitivity. [6]
The XRCC3 protein is one of five paralogs of RAD51, including RAD51B (RAD51L1), RAD51C (RAD51L2), RAD51D (RAD51L3), XRCC2 and XRCC3. They each share about 25% amino acid sequence identity with RAD51 and each other. [7]
The RAD51 paralogs are all required for efficient DNA double-strand break repair by homologous recombination and depletion of any paralog results in significant decreases in homologous recombination frequency. [8]
Two paralogs form a complex designated CX3 (RAD51C-XRCC3). Four paralogs form a second complex designated BCDX2 (RAD51B-RAD51C-RAD51D-XRCC2). These two complexes act at two different stages of homologous recombinational DNA repair.
The CX3 complex acts downstream of RAD51, after its recruitment to damage sites. [8] The CX3 complex associates with Holliday junction resolvase activity, probably in a role of stabilizing gene conversion tracts. [8]
The BCDX2 complex is responsible for RAD51 recruitment or stabilization at damage sites. [8] The BCDX2 complex appears to act by facilitating the assembly or stability of the RAD51 nucleoprotein filament.
XRCC3 has been shown to interact with RAD51C. [9] [10] [11] [12]
There is an epigenetic cause of XRCC3 deficiency that appears to increase cancer risk. This is the repression of XRCC3 by over-expression of EZH2 protein.
Increased expression of EZH2 leads to epigenetic repression of RAD51 paralogs, including XRCC3, and thus reduces homologous recombinational repair. [13] This reduction was proposed to be a cause of breast cancer. [13] EZH2 is the catalytic subunit of Polycomb Repressor Complex 2 (PRC2) which catalyzes methylation of histone H3 at lysine 27 (H3K27me) and mediates gene silencing of target genes via local chromatin reorganization. [14] EZH2 protein is up-regulated in numerous cancers. [14] [15] EZH2 mRNA is up-regulated, on average, 7.5-fold in breast cancer, and between 40% to 75% of breast cancers have over-expressed EZH2 protein. [16]
Click on genes, proteins and metabolites below to link to respective articles. [§ 1]
RecQ helicase is a family of helicase enzymes initially found in Escherichia coli that has been shown to be important in genome maintenance. They function through catalyzing the reaction ATP + H2O → ADP + P and thus driving the unwinding of paired DNA and translocating in the 3' to 5' direction. These enzymes can also drive the reaction NTP + H2O → NDP + P to drive the unwinding of either DNA or RNA.

Non-homologous end joining (NHEJ) is a pathway that repairs double-strand breaks in DNA. NHEJ is referred to as "non-homologous" because the break ends are directly ligated without the need for a homologous template, in contrast to homology directed repair(HDR), which requires a homologous sequence to guide repair. NHEJ is active in both non-dividing and proliferating cells, while HDR is not readily accessible in non-dividing cells. The term "non-homologous end joining" was coined in 1996 by Moore and Haber.

Homologous recombination is a type of genetic recombination in which genetic information is exchanged between two similar or identical molecules of double-stranded or single-stranded nucleic acids.
DNA repair protein RAD51 homolog 1 is a protein encoded by the gene RAD51. The enzyme encoded by this gene is a member of the RAD51 protein family which assists in repair of DNA double strand breaks. RAD51 family members are homologous to the bacterial RecA, Archaeal RadA and yeast Rad51. The protein is highly conserved in most eukaryotes, from yeast to humans.

DNA repair protein XRCC4 also known as X-ray repair cross-complementing protein 4 or XRCC4 is a protein that in humans is encoded by the XRCC4 gene. In addition to humans, the XRCC4 protein is also expressed in many other metazoans, fungi and in plants. The X-ray repair cross-complementing protein 4 is one of several core proteins involved in the non-homologous end joining (NHEJ) pathway to repair DNA double strand breaks (DSBs).

Double-strand break repair protein MRE11 is an enzyme that in humans is encoded by the MRE11 gene. The gene has been designated MRE11A to distinguish it from the pseudogene MRE11B that is nowadays named MRE11P1.

Fanconi anemia group G protein is a protein that in humans is encoded by the FANCG gene.

RAD52 homolog , also known as RAD52, is a protein which in humans is encoded by the RAD52 gene.

DNA repair protein RAD51 homolog 2 is a protein that in humans is encoded by the RAD51L1 gene.

RAD51 homolog C , also known as RAD51C, is a protein which in humans is encoded by the RAD51C gene.

DNA repair protein RAD51 homolog 4 is a protein that in humans is encoded by the RAD51L3 gene.

DNA repair protein XRCC2 is a protein that in humans is encoded by the XRCC2 gene.

DNA repair and recombination protein RAD54-like is a protein that in humans is encoded by the RAD54L gene.

E3 ubiquitin-protein ligase FANCL is an enzyme that in humans is encoded by the FANCL gene.

DNA repair and recombination protein RAD54B is a protein that in humans is encoded by the RAD54B gene.

Partner and localizer of BRCA2, also known as PALB2 or FANCN, is a protein which in humans is encoded by the PALB2 gene.
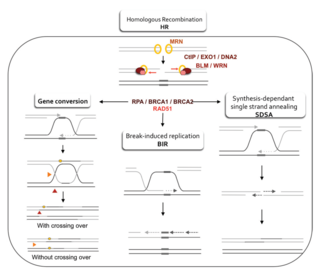
Homology-directed repair (HDR) is a mechanism in cells to repair double-strand DNA lesions. The most common form of HDR is homologous recombination. The HDR mechanism can only be used by the cell when there is a homologous piece of DNA present in the nucleus, mostly in G2 and S phase of the cell cycle. Other examples of homology-directed repair include single-strand annealing and breakage-induced replication. When the homologous DNA is absent, another process called non-homologous end joining (NHEJ) takes place instead.

DNA ligase 3 is an enzyme that, in humans, is encoded by the LIG3 gene. The human LIG3 gene encodes ATP-dependent DNA ligases that seal interruptions in the phosphodiester backbone of duplex DNA.

RI-1 is a selective inhibitor of RAD51, which is a central gene molecule of homologous recombination, with IC50 ranging from 5 to 30 μM.

A double-strand break repair model refers to the various models of pathways that cells undertake to repair double strand-breaks (DSB). DSB repair is an important cellular process, as the accumulation of unrepaired DSB could lead to chromosomal rearrangements, tumorigenesis or even cell death. In human cells, there are two main DSB repair mechanisms: Homologous recombination (HR) and non-homologous end joining (NHEJ). HR relies on undamaged template DNA as reference to repair the DSB, resulting in the restoration of the original sequence. NHEJ modifies and ligates the damaged ends regardless of homology. In terms of DSB repair pathway choice, most mammalian cells appear to favor NHEJ rather than HR. This is because the employment of HR may lead to gene deletion or amplification in cells which contains repetitive sequences. In terms of repair models in the cell cycle, HR is only possible during the S and G2 phases, while NHEJ can occur throughout whole process. These repair pathways are all regulated by the overarching DNA damage response mechanism. Besides HR and NHEJ, there are also other repair models which exists in cells. Some are categorized under HR, such as synthesis-dependent strain annealing, break-induced replication, and single-strand annealing; while others are an entirely alternate repair model, namely, the pathway microhomology-mediated end joining (MMEJ).