DNA polymerase alpha catalytic subunit is an enzyme that in humans is encoded by the POLA1 gene. [5]
Contents

DNA polymerase alpha catalytic subunit is an enzyme that in humans is encoded by the POLA1 gene. [5]

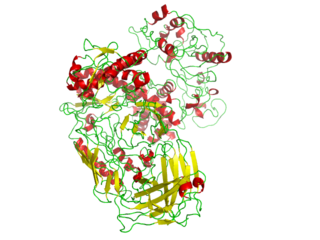
This gene encodes the p180 catalytic subunit of DNA polymerase α-primase. Pol α has limited processivity and lacks 3′ exonuclease activity for proofreading errors. Thus it is not well suited to efficiently and accurately copy long templates (unlike Pol Delta and Epsilon). Instead it plays a more limited role in replication. Pol α is responsible for the initiation of DNA replication at origins of replication (on both the leading and lagging strands) and during synthesis of Okazaki fragments on the lagging strand. The Pol α complex (pol α-DNA primase complex) consists of four subunits: the catalytic subunit POLA1, the regulatory subunit POLA2, and the small and the large primase subunits PRIM1 and PRIM2 respectively. Once primase has created the RNA primer, Pol α starts replication elongating the primer with ~20 nucleotides.
In addition to its role during DNA replication, POLA1 plays a role in type I interferon activation. The POLA1 gene was found to be the site of a mutation resulting in X-linked reticulate pigmentary disorder (XLPDR), OMIM 301220). This leads to altered mRNA splicing and decreased expression of POLA1 protein to a level that does not impair DNA replication. The reduction in POLA1 expression is accompanied by marked reduction in cytosolic RNA:DNA hybrid molecules and a concomitant hyperactivation of the IRF3 pathway, with consequent overproduction of type I interferons. [7]
Moreover, POLA1 deficiency, typical for XLPDR, also impair direct cytotoxicity of NK cells. POLA1 inhibition or a natural deficiency (XLPDR) affects the way the lytic granules secreted toward target cells. As a result, NK cells in XLPDR patients display functional deficiency. Interestingly, the POLA1 deficiency typical for XLPDR is not associated with any genomic damages or cell cycle arrest. [8] [9]
While the XLPDR mutation is resided in intron 13th, other somatic mutations in POLA1 were also described. Somatic mutation are associated with more profound deficiency of POLA1, with develops into X-linked intellectual disability (XLID). In a case of non-XLPDR mutations, beside of type I interferon signature patients also display mild to medium signs of intellectual disability, cell cycle arrest, proportionate short stature, microcephaly and hypogonadism. [10]
DNA dependent polymerase alpha (Pol α) has been shown to interact with MCM4 and GINS1, [8] Retinoblastoma protein, [11] PARP1 [12] [13] and RBMS1. [14]

In molecular biology, DNA replication is the biological process of producing two identical replicas of DNA from one original DNA molecule. DNA replication occurs in all living organisms acting as the most essential part of biological inheritance. This is essential for cell division during growth and repair of damaged tissues, while it also ensures that each of the new cells receives its own copy of the DNA. The cell possesses the distinctive property of division, which makes replication of DNA essential.
A polymerase is an enzyme that synthesizes long chains of polymers or nucleic acids. DNA polymerase and RNA polymerase are used to assemble DNA and RNA molecules, respectively, by copying a DNA template strand using base-pairing interactions or RNA by half ladder replication.

In molecular biology, RNA polymerase, or more specifically DNA-directed/dependent RNA polymerase (DdRP), is an enzyme that catalyzes the chemical reactions that synthesize RNA from a DNA template.

A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create two identical DNA duplexes from a single original DNA duplex. During this process, DNA polymerase "reads" the existing DNA strands to create two new strands that match the existing ones. These enzymes catalyze the chemical reaction
DNA primase is an enzyme involved in the replication of DNA and is a type of RNA polymerase. Primase catalyzes the synthesis of a short RNA segment called a primer complementary to a ssDNA template. After this elongation, the RNA piece is removed by a 5' to 3' exonuclease and refilled with DNA.

DNA polymerase I is an enzyme that participates in the process of prokaryotic DNA replication. Discovered by Arthur Kornberg in 1956, it was the first known DNA polymerase. It was initially characterized in E. coli and is ubiquitous in prokaryotes. In E. coli and many other bacteria, the gene that encodes Pol I is known as polA. The E. coli Pol I enzyme is composed of 928 amino acids, and is an example of a processive enzyme — it can sequentially catalyze multiple polymerisation steps without releasing the single-stranded template. The physiological function of Pol I is mainly to support repair of damaged DNA, but it also contributes to connecting Okazaki fragments by deleting RNA primers and replacing the ribonucleotides with DNA.
dnaQ is the gene encoding the ε subunit of DNA polymerase III in Escherichia coli. The ε subunit is one of three core proteins in the DNA polymerase complex. It functions as a 3’→5’ DNA directed proofreading exonuclease that removes incorrectly incorporated bases during replication. dnaQ may also be referred to as mutD.

DNA polymerase II is a prokaryotic DNA-dependent DNA polymerase encoded by the PolB gene.

Proliferating cell nuclear antigen (PCNA) is a DNA clamp that acts as a processivity factor for DNA polymerase δ in eukaryotic cells and is essential for replication. PCNA is a homotrimer and achieves its processivity by encircling the DNA, where it acts as a scaffold to recruit proteins involved in DNA replication, DNA repair, chromatin remodeling and epigenetics.

A DNA clamp, also known as a sliding clamp, is a protein complex that serves as a processivity-promoting factor in DNA replication. As a critical component of the DNA polymerase III holoenzyme, the clamp protein binds DNA polymerase and prevents this enzyme from dissociating from the template DNA strand. The clamp-polymerase protein–protein interactions are stronger and more specific than the direct interactions between the polymerase and the template DNA strand; because one of the rate-limiting steps in the DNA synthesis reaction is the association of the polymerase with the DNA template, the presence of the sliding clamp dramatically increases the number of nucleotides that the polymerase can add to the growing strand per association event. The presence of the DNA clamp can increase the rate of DNA synthesis up to 1,000-fold compared with a nonprocessive polymerase.

Eukaryotic DNA replication is a conserved mechanism that restricts DNA replication to once per cell cycle. Eukaryotic DNA replication of chromosomal DNA is central for the duplication of a cell and is necessary for the maintenance of the eukaryotic genome.

The gene polymerase delta 1 (POLD1) encodes the large, POLD1/p125, catalytic subunit of the DNA polymerase delta (Polδ) complex. The Polδ enzyme is responsible for synthesizing the lagging strand of DNA, and has also been implicated in some activities at the leading strand. The POLD1/p125 subunit encodes both DNA polymerizing and exonuclease domains, which provide the protein an important second function in proofreading to ensure replication accuracy during DNA synthesis, and in a number of types of replication-linked DNA repair following DNA damage.

DNA primase large subunit is an enzyme that in humans is encoded by the PRIM2 gene.
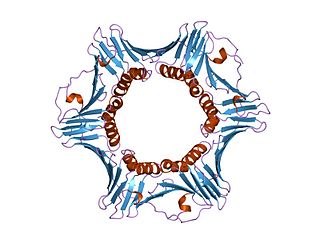
DNA polymerase delta subunit 3 is an enzyme that in humans is encoded by the POLD3 gene. It is a component of the DNA polymerase delta complex.

T7 DNA polymerase is an enzyme used during the DNA replication of the T7 bacteriophage. During this process, the DNA polymerase “reads” existing DNA strands and creates two new strands that match the existing ones. The T7 DNA polymerase requires a host factor, E. coli thioredoxin, in order to carry out its function. This helps stabilize the binding of the necessary protein to the primer-template to improve processivity by more than 100-fold, which is a feature unique to this enzyme. It is a member of the Family A DNA polymerases, which include E. coli DNA polymerase I and Taq DNA polymerase.

DNA polymerase epsilon catalytic subunit is an enzyme that in humans is encoded by the POLE gene. It is the central catalytic subunit of DNA polymerase epsilon.

PrimPol is a protein encoded by the PRIMPOL gene in humans. PrimPol is a eukaryotic protein with both DNA polymerase and DNA Primase activities involved in translesion DNA synthesis. It is the first eukaryotic protein to be identified with priming activity using deoxyribonucleotides. It is also the first protein identified in the mitochondria to have translesion DNA synthesis activities.

DNA polymerase alpha subunit 2 is an enzyme that in humans is encoded by the POLA2 gene.

DNA primase small subunit is an enzyme that in humans is encoded by the PRIM1 gene.

DNA polymerase alpha also known as Pol α is an enzyme complex found in eukaryotes that is involved in initiation of DNA replication. The DNA polymerase alpha complex consists of 4 subunits: POLA1, POLA2, PRIM1, and PRIM2.