Oxidative phosphorylation or electron transport-linked phosphorylation or terminal oxidation is the metabolic pathway in which cells use enzymes to oxidize nutrients, thereby releasing chemical energy in order to produce adenosine triphosphate (ATP). In eukaryotes, this takes place inside mitochondria. Almost all aerobic organisms carry out oxidative phosphorylation. This pathway is so pervasive because it releases more energy than alternative fermentation processes such as anaerobic glycolysis.
An electron transport chain (ETC) is a series of protein complexes and other molecules that transfer electrons from electron donors to electron acceptors via redox reactions (both reduction and oxidation occurring simultaneously) and couples this electron transfer with the transfer of protons (H+ ions) across a membrane. Many of the enzymes in the electron transport chain are embedded within the membrane.

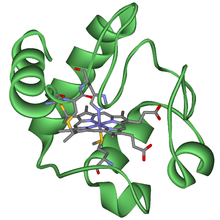
The cytochrome complex, or cyt c, is a small hemeprotein found loosely associated with the inner membrane of the mitochondrion where it plays a critical role in cellular respiration. It transfers electrons between Complexes III and IV. Cytochrome c is highly water-soluble, unlike other cytochromes. It is capable of undergoing oxidation and reduction as its iron atom converts between the ferrous and ferric forms, but does not bind oxygen. It also plays a major role in cell apoptosis. In humans, cytochrome c is encoded by the CYCS gene.

A hemeprotein, or heme protein, is a protein that contains a heme prosthetic group. They are a very large class of metalloproteins. The heme group confers functionality, which can include oxygen carrying, oxygen reduction, electron transfer, and other processes. Heme is bound to the protein either covalently or noncovalently or both.

The coenzyme Q : cytochrome c – oxidoreductase, sometimes called the cytochrome bc1 complex, and at other times complex III, is the third complex in the electron transport chain, playing a critical role in biochemical generation of ATP. Complex III is a multisubunit transmembrane protein encoded by both the mitochondrial and the nuclear genomes. Complex III is present in the mitochondria of all animals and all aerobic eukaryotes and the inner membranes of most eubacteria. Mutations in Complex III cause exercise intolerance as well as multisystem disorders. The bc1 complex contains 11 subunits, 3 respiratory subunits, 2 core proteins and 6 low-molecular weight proteins.

Heme, or haem, is a precursor to hemoglobin, which is necessary to bind oxygen in the bloodstream. Heme is biosynthesized in both the bone marrow and the liver.
Cytochrome C1 is a protein encoded by the CYC1 gene. Cytochrome is a heme-containing subunit of the cytochrome b-c1 complex, which accepts electrons from Rieske protein and transfers electrons to cytochrome c in the mitochondrial respiratory chain. It is formed in the cytosol and targeted to the mitochondrial intermembrane space. Cytochrome c1 belongs to the cytochrome c family of proteins.

In biochemistry, flavin adenine dinucleotide (FAD) is a redox-active coenzyme associated with various proteins, which is involved with several enzymatic reactions in metabolism. A flavoprotein is a protein that contains a flavin group, which may be in the form of FAD or flavin mononucleotide (FMN). Many flavoproteins are known: components of the succinate dehydrogenase complex, α-ketoglutarate dehydrogenase, and a component of the pyruvate dehydrogenase complex.

The cytochrome b6f complex (plastoquinol/plastocyanin reductase or plastoquinol/plastocyanin oxidoreductase; EC 7.1.1.6) is an enzyme found in the thylakoid membrane in chloroplasts of plants, cyanobacteria, and green algae, that catalyzes the transfer of electrons from plastoquinol to plastocyanin:

Rieske proteins are iron–sulfur protein (ISP) components of cytochrome bc1 complexes and cytochrome b6f complexes and are responsible for electron transfer in some biological systems. John S. Rieske and co-workers first discovered the protein and in 1964 isolated an acetylated form of the bovine mitochondrial protein. In 1979 Trumpower's lab isolated the "oxidation factor" from bovine mitochondria and showed it was a reconstitutively-active form of the Rieske iron-sulfur protein
It is a unique [2Fe-2S] cluster in that one of the two Fe atoms is coordinated by two histidine residues rather than two cysteine residues. They have since been found in plants, animals, and bacteria with widely ranging electron reduction potentials from -150 to +400 mV.
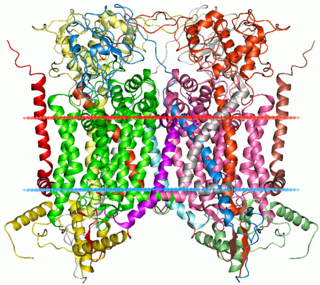
Cytochrome b within both molecular and cell biology, is a protein found in the membranes of aerobic cells. In eukaryotic mitochondria and in aerobic prokaryotes, cytochrome b is a component of respiratory chain complex III — also known as the bc1 complex or ubiquinol-cytochrome c reductase. In plant chloroplasts and cyanobacteria, there is an homologous protein, cytochrome b6, a component of the plastoquinone-plastocyanin reductase, also known as the b6f complex. These complexes are involved in electron transport, the pumping of protons to create a proton-motive force (PMF). This proton gradient is used for the generation of ATP. These complexes play a vital role in cells.
Iron-binding proteins are carrier proteins and metalloproteins that are important in iron metabolism and the immune response. Iron is required for life.

Heme C is an important kind of heme.
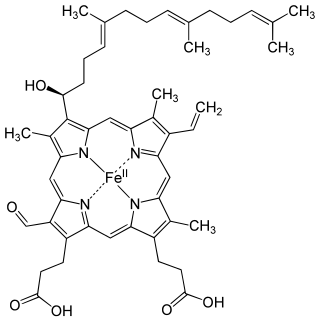
Heme A is a heme, a coordination complex consisting of a macrocyclic ligand called a porphyrin, chelating an iron atom. Heme A is a biomolecule and is produced naturally by many organisms. Heme A, often appears a dichroic green/red when in solution, is a structural relative of heme B, a component of hemoglobin, the red pigment in blood.
In enzymology, a cellobiose dehydrogenase (acceptor) (EC 1.1.99.18) is an enzyme that catalyzes the chemical reaction
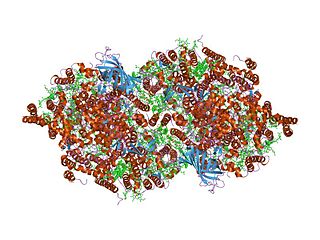
Cytochrome b559 is an important component of Photosystem II (PSII) is a multisubunit protein-pigment complex containing polypeptides both intrinsic and extrinsic to the photosynthetic membrane. Within the core of the complex, the chlorophyll and beta-carotene pigments are mainly bound to the antenna proteins CP43 (PsbC) and CP47 (PsbB), which pass the excitation energy on to chlorophylls in the reaction centre proteins D1 and D2 that bind all the redox-active cofactors involved in the energy conversion process. The PSII oxygen-evolving complex (OEC) provides electrons to re-reduce the PSII reaction center, and oxidizes 2 water molecules to recover its reduced initial state. It consists of OEE1 (PsbO), OEE2 (PsbP) and OEE3 (PsbQ). The remaining subunits in PSII are of low molecular weight, and are involved in PSII assembly, stabilisation, dimerization, and photoprotection.

Cytochromes c cytochromes, or heme-containing proteins, that have heme C covalently attached to the peptide backbone via one or two thioether bonds. These bonds are in most cases part of a specific Cys-X-X-Cys-His (CXXCH) binding motif, where X denotes a miscellaneous amino acid. Two thioether bonds of cysteine residues bind to the vinyl sidechains of heme, and the histidine residue coordinates one axial binding site of the heme iron. Less common binding motifs can include a single thioether linkage, a lysine or a methionine instead of the axial histidine or a CXnCH binding motif with n>2. The second axial site of the iron can be coordinated by amino acids of the protein, substrate molecules or water. Cytochromes c possess a wide range of properties and function as electron transfer proteins or catalyse chemical reactions involving redox processes. A prominent member of this family is mitochondrial cytochrome c.

Light-dependent reactions refers to certain photochemical reactions that are involved in photosynthesis, the main process by which plants acquire energy. There are two light dependent reactions, the first occurs at photosystem II (PSII) and the second occurs at photosystem I (PSI).

Cytochrome d, previously known as cytochrome a2, is a name for all cytochromes that contain heme D as a cofactor. Two unrelated classes of cytochrome d are known: Cytochrome bd, an enzyme that generates a charge across the membrane so that protons will move, and cytochrome cd1, a nitrite reductase.
In the field of enzymology, murburn is a term coined by Kelath Murali Manoj that explains the catalytic mechanism of certain redox-active proteins. The term describes the equilibrium among molecules, unbound ions and radicals, signifying a process of "mild unrestricted redox catalysis".