Related Research Articles

Heredity, also called inheritance or biological inheritance, is the passing on of traits from parents to their offspring; either through asexual reproduction or sexual reproduction, the offspring cells or organisms acquire the genetic information of their parents. Through heredity, variations between individuals can accumulate and cause species to evolve by natural selection. The study of heredity in biology is genetics.
Genomic imprinting is an epigenetic phenomenon that causes genes to be expressed or not, depending on whether they are inherited from the female or male parent. Genes can also be partially imprinted. Partial imprinting occurs when alleles from both parents are differently expressed rather than complete expression and complete suppression of one parent's allele. Forms of genomic imprinting have been demonstrated in fungi, plants and animals. In 2014, there were about 150 imprinted genes known in mice and about half that in humans. As of 2019, 260 imprinted genes have been reported in mice and 228 in humans.

In biology, epigenetics is the study of heritable traits, or a stable change of cell function, that happen without changes to the DNA sequence. The Greek prefix epi- in epigenetics implies features that are "on top of" or "in addition to" the traditional genetic mechanism of inheritance. Epigenetics usually involves a change that is not erased by cell division, and affects the regulation of gene expression. Such effects on cellular and physiological phenotypic traits may result from environmental factors, or be part of normal development. Epigenetic factors can also lead to cancer.
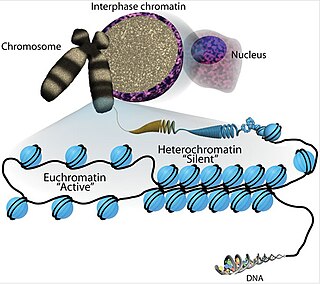
Euchromatin is a lightly packed form of chromatin that is enriched in genes, and is often under active transcription. Euchromatin stands in contrast to heterochromatin, which is tightly packed and less accessible for transcription. 92% of the human genome is euchromatic.
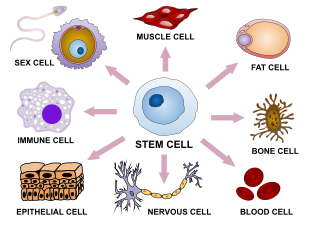
Cellular differentiation is the process in which a stem cell changes from one type to a differentiated one. Usually, the cell changes to a more specialized type. Differentiation happens multiple times during the development of a multicellular organism as it changes from a simple zygote to a complex system of tissues and cell types. Differentiation continues in adulthood as adult stem cells divide and create fully differentiated daughter cells during tissue repair and during normal cell turnover. Some differentiation occurs in response to antigen exposure. Differentiation dramatically changes a cell's size, shape, membrane potential, metabolic activity, and responsiveness to signals. These changes are largely due to highly controlled modifications in gene expression and are the study of epigenetics. With a few exceptions, cellular differentiation almost never involves a change in the DNA sequence itself. Metabolic composition, however, gets dramatically altered where stem cells are characterized by abundant metabolites with highly unsaturated structures whose levels decrease upon differentiation. Thus, different cells can have very different physical characteristics despite having the same genome.
A maternal effect is a situation where the phenotype of an organism is determined not only by the environment it experiences and its genotype, but also by the environment and genotype of its mother. In genetics, maternal effects occur when an organism shows the phenotype expected from the genotype of the mother, irrespective of its own genotype, often due to the mother supplying messenger RNA or proteins to the egg. Maternal effects can also be caused by the maternal environment independent of genotype, sometimes controlling the size, sex, or behaviour of the offspring. These adaptive maternal effects lead to phenotypes of offspring that increase their fitness. Further, it introduces the concept of phenotypic plasticity, an important evolutionary concept. It has been proposed that maternal effects are important for the evolution of adaptive responses to environmental heterogeneity.

In biology, the epigenome of an organism is the collection of chemical changes to its DNA and histone proteins that affects when, where, and how the DNA is expressed; these changes can be passed down to an organism's offspring via transgenerational epigenetic inheritance. Changes to the epigenome can result in changes to the structure of chromatin and changes to the function of the genome. The human epigenome, including DNA methylation and histone modification, is maintained through cell division. The epigenome is essential for normal development and cellular differentiation, enabling cells with the same genetic code to perform different functions. The human epigenome is dynamic and can be influenced by environmental factors such as diet, stress, and toxins.

In epigenetics, a paramutation is an interaction between two alleles at a single locus, whereby one allele induces a heritable change in the other allele. The change may be in the pattern of DNA methylation or histone modifications. The allele inducing the change is said to be paramutagenic, while the allele that has been epigenetically altered is termed paramutable. A paramutable allele may have altered levels of gene expression, which may continue in offspring which inherit that allele, even though the paramutagenic allele may no longer be present. Through proper breeding, paramutation can result in siblings that have the same genetic sequence, but with drastically different phenotypes.
Histone methylation is a process by which methyl groups are transferred to amino acids of histone proteins that make up nucleosomes, which the DNA double helix wraps around to form chromosomes. Methylation of histones can either increase or decrease transcription of genes, depending on which amino acids in the histones are methylated, and how many methyl groups are attached. Methylation events that weaken chemical attractions between histone tails and DNA increase transcription because they enable the DNA to uncoil from nucleosomes so that transcription factor proteins and RNA polymerase can access the DNA. This process is critical for the regulation of gene expression that allows different cells to express different genes.
Epigenomics is the study of the complete set of epigenetic modifications on the genetic material of a cell, known as the epigenome. The field is analogous to genomics and proteomics, which are the study of the genome and proteome of a cell. Epigenetic modifications are reversible modifications on a cell's DNA or histones that affect gene expression without altering the DNA sequence. Epigenomic maintenance is a continuous process and plays an important role in stability of eukaryotic genomes by taking part in crucial biological mechanisms like DNA repair. Plant flavones are said to be inhibiting epigenomic marks that cause cancers. Two of the most characterized epigenetic modifications are DNA methylation and histone modification. Epigenetic modifications play an important role in gene expression and regulation, and are involved in numerous cellular processes such as in differentiation/development and tumorigenesis. The study of epigenetics on a global level has been made possible only recently through the adaptation of genomic high-throughput assays.

Transgenerational epigenetic inheritance is the proposed transmission of epigenetic markers and modifications from one generation to multiple subsequent generations without altering the primary structure of DNA. Thus, the regulation of genes via epigenetic mechanisms can be heritable; the amount of transcripts and proteins produced can be altered by inherited epigenetic changes. In order for epigenetic marks to be heritable, however, they must occur in the gametes in animals, but since plants lack a definitive germline and can propagate, epigenetic marks in any tissue can be heritable.
Behavioral epigenetics is the field of study examining the role of epigenetics in shaping animal and human behavior. It seeks to explain how nurture shapes nature, where nature refers to biological heredity and nurture refers to virtually everything that occurs during the life-span. Behavioral epigenetics attempts to provide a framework for understanding how the expression of genes is influenced by experiences and the environment to produce individual differences in behaviour, cognition, personality, and mental health.
The epigenetics of schizophrenia is the study of how inherited epigenetic changes are regulated and modified by the environment and external factors and how these changes influence the onset and development of, and vulnerability to, schizophrenia. Epigenetics concerns the heritability of those changes, too. Schizophrenia is a debilitating and often misunderstood disorder that affects up to 1% of the world's population. Although schizophrenia is a heavily studied disorder, it has remained largely impervious to scientific understanding; epigenetics offers a new avenue for research, understanding, and treatment.
Epigenetics of human development is the study of how epigenetics effects human development.
Neuroepigenetics is the study of how epigenetic changes to genes affect the nervous system. These changes may effect underlying conditions such as addiction, cognition, and neurological development.

An epigenome-wide association study (EWAS) is an examination of a genome-wide set of quantifiable epigenetic marks, such as DNA methylation, in different individuals to derive associations between epigenetic variation and a particular identifiable phenotype/trait. When patterns change such as DNA methylation at specific loci, discriminating the phenotypically affected cases from control individuals, this is considered an indication that epigenetic perturbation has taken place that is associated, causally or consequentially, with the phenotype.
Epigenetics of anxiety and stress–related disorders is the field studying the relationship between epigenetic modifications of genes and anxiety and stress-related disorders, including mental health disorders such as generalized anxiety disorder (GAD), post-traumatic stress disorder, obsessive-compulsive disorder (OCD), and more. These changes can lead to transgenerational stress inheritance.
Transgenerational epigenetic inheritance in plants involves mechanisms for the passing of epigenetic marks from parent to offspring that differ from those reported in animals. There are several kinds of epigenetic markers, but they all provide a mechanism to facilitate greater phenotypic plasticity by influencing the expression of genes without altering the DNA code. These modifications represent responses to environmental input and are reversible changes to gene expression patterns that can be passed down through generations. In plants, transgenerational epigenetic inheritance could potentially represent an evolutionary adaptation for sessile organisms to quickly adapt to their changing environment.
Sleep epigenetics is the field of how epigenetics affects sleep.
Environmental epigenetics is a branch of epigenetics that studies the influence of external environmental factors on the gene expression of a developing embryo. The way that genes are expressed may be passed down from parent to offspring through epigenetic modifications, although environmental influences do not alter the genome itself.
References
- ↑ Suter CM, Boffelli D, Martin DI (November 2013). "A role for epigenetic inheritance in modern evolutionary theory? A comment in response to Dickins and Rahman". Proceedings. Biological Sciences. 280 (1771): 20130903, discussion 20131820. doi:10.1098/rspb.2013.0903. PMC 3790474 . PMID 24089330.
- ↑ Sapozhnikov, Daniel M.; Szyf, Moshe (2021-09-29). "Unraveling the functional role of DNA demethylation at specific promoters by targeted steric blockage of DNA methyltransferase with CRISPR/dCas9". Nature Communications. 12 (1): 5711. Bibcode:2021NatCo..12.5711S. doi:10.1038/s41467-021-25991-9. ISSN 2041-1723. PMC 8481236 . PMID 34588447.
- 1 2 Turck F, Coupland G (March 2014). "Natural variation in epigenetic gene regulation and its effects on plant developmental traits". Evolution; International Journal of Organic Evolution. 68 (3): 620–631. doi:10.1111/evo.12286. hdl: 11858/00-001M-0000-0024-04A8-4 . PMID 24117443. S2CID 10225862.
- ↑ Zhang YY, Fischer M, Colot V, Bossdorf O (January 2013). "Epigenetic variation creates potential for evolution of plant phenotypic plasticity". The New Phytologist. 197 (1): 314–322. Bibcode:2013NewPh.197..314Z. doi: 10.1111/nph.12010 . PMID 23121242.
- ↑ Verhoeven KJ, Jansen JJ, van Dijk PJ, Biere A (March 2010). "Stress-induced DNA methylation changes and their heritability in asexual dandelions". The New Phytologist. 185 (4): 1108–18. Bibcode:2010NewPh.185.1108V. doi: 10.1111/j.1469-8137.2009.03121.x . PMID 20003072.
- ↑ Hernando-Herraez I, Prado-Martinez J, Garg P, Fernandez-Callejo M, Heyn H, et al. (2013) Dynamics of DNA Methylation in Recent Human and Great Ape Evolution. PLoS Genet 9(9): e1003763. doi: 10.1371/journal.pgen.1003763
- ↑ Shulha HP, Crisci JL, Reshetov D, Tushir JS, Cheung I, Bharadwaj R, Chou HJ, Houston IB, Peter CJ, Mitchell AC, Yao WD, Myers RH, Chen JF, Preuss TM, Rogaev EI, Jensen JD, Weng Z, Akbarian S (2012). "Human-specific histone methylation signatures at transcription start sites in prefrontal neurons". PLOS Biology. 10 (11): e1001427. doi: 10.1371/journal.pbio.1001427 . PMC 3502543 . PMID 23185133.
- ↑ Molaro A, Hodges E, Fang F, Song Q, McCombie WR, Hannon GJ, Smith AD (September 2011). "Sperm methylation profiles reveal features of epigenetic inheritance and evolution in primates". Cell. 146 (6): 1029–41. doi:10.1016/j.cell.2011.08.016. PMC 3205962 . PMID 21925323.
- ↑ Blake LE, Roux J, Hernando-Herraez I, Banovich NE, Perez RG, Hsiao CJ, et al. (February 2020). "A comparison of gene expression and DNA methylation patterns across tissues and species". Genome Research. 30 (2): 250–262. doi:10.1101/gr.254904.119. PMC 7050529 . PMID 31953346.
- ↑ Nätt D, Rubin CJ, Wright D, Johnsson M, Beltéky J, Andersson L, Jensen P (February 2012). "Heritable genome-wide variation of gene expression and promoter methylation between wild and domesticated chickens". BMC Genomics. 13: 59. doi: 10.1186/1471-2164-13-59 . PMC 3297523 . PMID 22305654.
- ↑ Dickins TE, Rahman Q (August 2012). "The extended evolutionary synthesis and the role of soft inheritance in evolution". Proceedings. Biological Sciences. 279 (1740): 2913–21. doi:10.1098/rspb.2012.0273. PMC 3385474 . PMID 22593110.
- ↑ Rando OJ, Verstrepen KJ (February 2007). "Timescales of genetic and epigenetic inheritance" (PDF). Cell. 128 (4): 655–68. doi: 10.1016/j.cell.2007.01.023 . PMID 17320504. S2CID 17964015.
- ↑ Lancaster AK, Masel J (September 2009). "The evolution of reversible switches in the presence of irreversible mimics". Evolution; International Journal of Organic Evolution. 63 (9): 2350–62. doi:10.1111/j.1558-5646.2009.00729.x. PMC 2770902 . PMID 19486147.
- ↑ Pennisi E (September 2013). "Plant biology. Evolution heresy? Epigenetics underlies heritable plant traits". Science. 341 (6150): 1055. doi:10.1126/science.341.6150.1055. PMID 24009370.