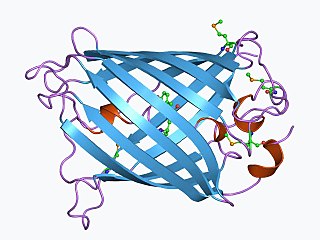
The green fluorescent protein (GFP) is a protein that exhibits bright green fluorescence when exposed to light in the blue to ultraviolet range. The label GFP traditionally refers to the protein first isolated from the jellyfish Aequorea victoria and is sometimes called avGFP. However, GFPs have been found in other organisms including corals, sea anemones, zoanithids, copepods and lancelets.
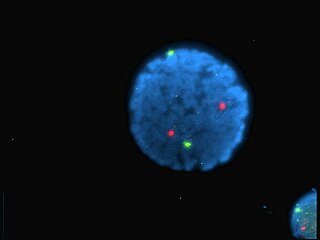
A fluorophore is a fluorescent chemical compound that can re-emit light upon light excitation. Fluorophores typically contain several combined aromatic groups, or planar or cyclic molecules with several π bonds.

Aequorin is a calcium-activated photoprotein isolated from the hydrozoan Aequorea victoria. Its bioluminescence was studied decades before the protein was isolated from the animal by Osamu Shimomura in 1962. In the animal, the protein occurs together with the green fluorescent protein to produce green light by resonant energy transfer, while aequorin by itself generates blue light.

Fura-2-acetoxymethyl ester, often abbreviated Fura-2AM, is a membrane-permeant derivative of the ratiometric calcium indicator Fura-2 used in biochemistry to measure cellular calcium concentrations by fluorescence. When added to cells, Fura-2AM crosses cell membranes and once inside the cell, the acetoxymethyl groups are removed by cellular esterases. Removal of the acetoxymethyl esters regenerates "Fura-2", the pentacarboxylate calcium indicator. Measurement of Ca2+-induced fluorescence at both 340 nm and 380 nm allows for calculation of calcium concentrations based 340/380 ratios. The use of the ratio automatically cancels out certain variables such as local differences in fura-2 concentration or cell thickness that would otherwise lead to artifacts when attempting to image calcium concentrations in cells.
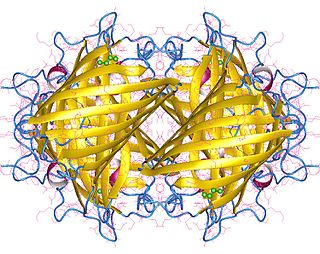
Yellow fluorescent protein (YFP) is a genetic mutant of green fluorescent protein (GFP) originally derived from the jellyfish Aequorea victoria. Its excitation peak is 513 nm and its emission peak is 527 nm. Like the parent GFP, YFP is a useful tool in cell and molecular biology because the excitation and emission peaks of YFP are distinguishable from GFP which allows for the study of multiple processes/proteins within the same experiment.
Cameleon is an engineered protein based on variant of green fluorescent protein used to visualize calcium levels in living cells. It is a genetically encoded calcium sensor created by Roger Y. Tsien and coworkers. The name is a conflation of CaM and chameleon to indicate the fact that the sensor protein undergoes a conformation change and radiates at an altered wavelength upon calcium binding to the calmodulin element of the Cameleon. Cameleon was the first genetically encoded calcium sensor that could be used for ratiometric measurements and the first to be used in a transgenic animal to record activity in neurons and muscle cells. Cameleon and other genetically-encoded calcium indicators (GECIs) have found many applications in neuroscience and other fields of biology. It was created by fusing BFP, calmodulin, calmodulin-binding peptide M13 and EGFP.
Calcein, also known as fluorexon, fluorescein complex, is a fluorescent dye with excitation and emission wavelengths of 495/515 nm, respectively, and has the appearance of orange crystals. Calcein self-quenches at concentrations above 70mM and is commonly used as an indicator of lipid vesicle leakage. It is also used traditionally as a complexometric indicator for titration of calcium ions with EDTA, and for fluorometric determination of calcium.

Roger Yonchien Tsien was an American biochemist. He was a professor of chemistry and biochemistry at the University of California, San Diego and was awarded the Nobel Prize in Chemistry in 2008 for his discovery and development of the green fluorescent protein, in collaboration with organic chemist Osamu Shimomura and neurobiologist Martin Chalfie. Tsien was also a pioneer of calcium imaging.
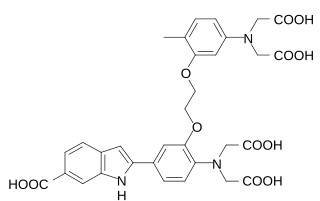
Indo-1 is a popular calcium indicator similar to Fura-2. In contrast to Fura-2, Indo-1 has a dual emissions peak. The main emission peak in calcium-free solution is 475 nm while in the presence of calcium the emission is shifted to 400 nm. It is widely used in flow cytometry. The penta potassium salt is commercially available and preferred to the free acid because of its higher solubility in water. While Indo-1 is not cell permeable the penta acetoxymethyl ester Indo-1 AM enters the cell where it is cleaved by intracellular esterases to Indo-1. The synthesis and properties of Indo-1 were presented in 1985 by the group of Roger Y Tsien.

Fluorescence is used in the life sciences generally as a non-destructive way of tracking or analysing biological molecules. Some proteins or small molecules in cells are naturally fluorescent, which is called intrinsic fluorescence or autofluorescence. Alternatively, specific or general proteins, nucleic acids, lipids or small molecules can be "labelled" with an extrinsic fluorophore, a fluorescent dye which can be a small molecule, protein or quantum dot. Several techniques exist to exploit additional properties of fluorophores, such as fluorescence resonance energy transfer, where the energy is passed non-radiatively to a particular neighbouring dye, allowing proximity or protein activation to be detected; another is the change in properties, such as intensity, of certain dyes depending on their environment allowing their use in structural studies.

GCaMP is a genetically encoded calcium indicator (GECI) initially developed in 2001 by Junichi Nakai. It is a synthetic fusion of green fluorescent protein (GFP), calmodulin (CaM), and M13, a peptide sequence from myosin light-chain kinase. When bound to Ca2+, GCaMP fluoresces green with a peak excitation wavelength of 480 nm and a peak emission wavelength of 510 nm. It is used in biological research to measure intracellular Ca2+ levels both in vitro and in vivo using virally transfected or transgenic cell and animal lines. The genetic sequence encoding GCaMP can be inserted under the control of promoters exclusive to certain cell types, allowing for cell-type specific expression of GCaMP. Since Ca2+ is a second messenger that contributes to many cellular mechanisms and signaling pathways, GCaMP allows researchers to quantify the activity of Ca2+-based mechanisms and study the role of Ca2+ ions in biological processes of interest.
mCherry is a member of the mFruits family of monomeric red fluorescent proteins (mRFPs). As a RFP, mCherry was derived from DsRed of Discosoma sea anemones unlike green fluorescent proteins (GFPs) which are often derived from Aequorea victoria jellyfish. Fluorescent proteins are used to tag components in the cell, so they can be studied using fluorescence spectroscopy and fluorescence microscopy. mCherry absorbs light between 540-590 nm and emits light in the range of 550-650 nm. mCherry belongs to the group of fluorescent protein chromophores used as instruments to visualize genes and analyze their functions in experiments. Genome editing has been improved greatly through the precise insertion of these fluorescent protein tags into the genetic material of many diverse organisms. Most comparisons between the brightness and photostability of different fluorescent proteins have been made in vitro, removed from biological variables that affect protein performance in cells or organisms. It is hard to perfectly simulate cellular environments in vitro, and the difference in environment could have an effect on the brightness and photostability.

Fluo-3 is a fluorescence indicator of intracellular calcium (Ca2+), developed by Roger Y. Tsien. It is used to measure Ca2+ inside living cells in flow cytometry, and confocal laser scanning microscopy using visible light excitation (compatible with argon laser sources operating at 488 nm). Fluo-3 and derivatives (Fluo-4, Fluo-5 etc) have also been widely used with two-photon excitation microscopy. Fluo-3 is an essentially nonfluorescent compound, but upon binding of Ca2+ its fluorescence increases sharply with an emission maximum at 525 nm suitable for conventionally used detectors designed for fluorescein isothiocyanate (FITC) measurements. This large change in fluorescence coupled with a good yield of photons provides very high contrast which allowed the detection of microscopic Ca2+ release events inside cells called "Calcium sparks". Whereas the salts of fluo-3 are unable to penetrate cells, loading can be achieved using its acetoxymethyl (AM) ester derivative. Once inside the cell, unspecific esterases cleave the ester effectively trapping fluo-3.

FlAsH-EDT2 is an organoarsenic compound with molecular formula C24H18As2O5S4. Its structure is based around a fluorescein core with two 1,3,2-dithiarsolane substituents. It is used in bioanalytical research as a fluorescent label for visualising proteins in living cells. FlAsH-EDT2 is an abbreviation for fluorescin arsenical hairpin binder-ethanedithiol, and is a pale yellow or pinkish fluorogenic solid. It has a semi-structural formula (C2H4AsS2)2-(C13H5O3)-C6H4COOH, representing the dithiarsolane substituents bound to the hydroxyxanthone core, attached to an o-substituted molecule of benzoic acid.
Calcium imaging is a microscopy technique to optically measure the calcium (Ca2+) status of an isolated cell, tissue or medium. Calcium imaging takes advantage of calcium indicators, fluorescent molecules that respond to the binding of Ca2+ ions by fluorescence properties. Two main classes of calcium indicators exist: chemical indicators and genetically encoded calcium indicators (GECI). This technique has allowed studies of calcium signalling in a wide variety of cell types. In neurons, electrical activity is always accompanied by an influx of Ca2+ ions. Thus, calcium imaging can be used to monitor the electrical activity in hundreds of neurons in cell culture or in living animals, which has made it possible to dissect the function of neuronal circuits.

Small ultra red fluorescent protein (smURFP) is a class of far-red fluorescent protein evolved from a cyanobacterial phycobiliprotein, α-allophycocyanin. Native α-allophycocyanin requires an exogenous protein, known as a lyase, to attach the chromophore, phycocyanobilin. Phycocyanobilin is not present in mammalian cells. smURFP was evolved to covalently attach phycocyanobilin without a lyase and fluoresce, covalently attach biliverdin and fluoresce, blue-shift fluorescence to match the organic fluorophore, Cy5, and not inhibit E. coli growth. smURFP was found after 12 rounds of random mutagenesis and manually screening 10,000,000 bacterial colonies.
In systems biology, live single-cell imaging is a live cell imaging technique that combines traditional live cell imaging and time-lapse microscopy techniques with automated cell tracking and feature extraction, drawing many techniques from high-content screening. It is used to study signalling dynamics and behaviour in populations of individual living cells. Live single cell studies can reveal key behaviours that would otherwise be masked in population averaging experiments such as western blots.
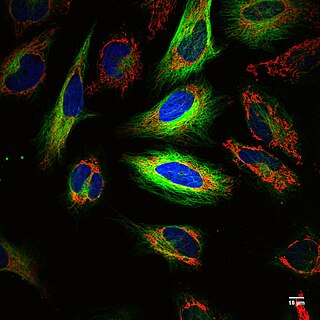
Fluorescence imaging is a type of non-invasive imaging technique that can help visualize biological processes taking place in a living organism. Images can be produced from a variety of methods including: microscopy, imaging probes, and spectroscopy.
Fiber photometry is a calcium imaging technique that captures 'bulk' or population-level calcium (Ca2+) activity from specific cell-types within a brain region or functional network in order to study neural circuits Population-level calcium activity can be correlated with behavioral tasks, such as spatial learning, memory recall and goal-directed behaviors. The technique involves the surgical implantation of fiber optics into the brains of living animals. The benefits to researchers are that optical fibers are simpler to implant, less invasive and less expensive than other calcium methods, and there is less weight and stress on the animal, as compared to miniscopes. It also allows for imaging of multiple interacting brain regions and integration with other neuroscience techniques. The limitations of fiber photometry are low cellular and spatial resolution, and the fact that animals must be securely tethered to a rigid fiber bundle, which may impact the naturalistic behavior of smaller mammals such as mice.
Optogenetics began with methods to alter neuronal activity with light, using e.g. channelrhodopsins. In a broader sense, optogenetic approaches also include the use of genetically encoded biosensors to monitor the activity of neurons or other cell types by measuring fluorescence or bioluminescence. Genetically encoded calcium indicators (GECIs) are used frequently to monitor neuronal activity, but other cellular parameters such as membrane voltage or second messenger activity can also be recorded optically. The use of optogenetic sensors is not restricted to neuroscience, but plays increasingly important roles in immunology, cardiology and cancer research.













