
The three-domain system is a biological classification introduced by Carl Woese et al. in 1990 that divides cellular life forms into archaea, bacteria, and eukaryote domains. The key difference from earlier classifications is the splitting of archaea from bacteria.

In molecular biology and genetics, translation is the process in which ribosomes in the cytoplasm or endoplasmic reticulum synthesize proteins after the process transcription of DNA to RNA in the cell's nucleus. The entire process is called gene expression.

Ribonuclease is a type of nuclease that catalyzes the degradation of RNA into smaller components. Ribonucleases can be divided into endoribonucleases and exoribonucleases, and comprise several sub-classes within the EC 2.7 and 3.1 classes of enzymes.

Ribonuclease H is a family of non-sequence-specific endonuclease enzymes that catalyze the cleavage of RNA in an RNA/DNA substrate via a hydrolytic mechanism. Members of the RNase H family can be found in nearly all organisms, from bacteria to archaea to eukaryotes.

Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosomal DNA (rDNA) and then bound to ribosomal proteins to form small and large ribosome subunits. rRNA is the physical and mechanical actor of the ribosome that forces transfer RNA (tRNA) and messenger RNA (mRNA) to process and translate the latter into proteins. Ribosomal RNA is the predominant form of RNA found in most cells; it makes up about 80% of cellular RNA despite never being translated into proteins itself. Ribosomes are composed of approximately 60% rRNA and 40% ribosomal proteins by mass.
Gene structure is the organisation of specialised sequence elements within a gene. Genes contain the information necessary for living cells to survive and reproduce. In most organisms, genes are made of DNA, where the particular DNA sequence determines the function of the gene. A gene is transcribed (copied) from DNA into RNA, which can either be non-coding (ncRNA) with a direct function, or an intermediate messenger (mRNA) that is then translated into protein. Each of these steps is controlled by specific sequence elements, or regions, within the gene. Every gene, therefore, requires multiple sequence elements to be functional. This includes the sequence that actually encodes the functional protein or ncRNA, as well as multiple regulatory sequence regions. These regions may be as short as a few base pairs, up to many thousands of base pairs long.

EF-Tu is a prokaryotic elongation factor responsible for catalyzing the binding of an aminoacyl-tRNA (aa-tRNA) to the ribosome. It is a G-protein, and facilitates the selection and binding of an aa-tRNA to the A-site of the ribosome. As a reflection of its crucial role in translation, EF-Tu is one of the most abundant and highly conserved proteins in prokaryotes. It is found in eukaryotic mitochrondria as TUFM.
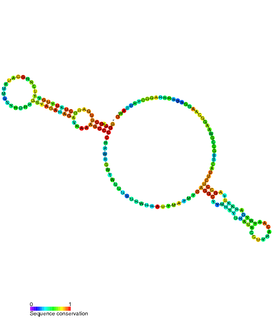
The hok/sok system is a postsegregational killing mechanism employed by the R1 plasmid in Escherichia coli. It was the first type I toxin-antitoxin pair to be identified through characterisation of a plasmid-stabilising locus. It is a type I system because the toxin is neutralised by a complementary RNA, rather than a partnered protein.

Telomerase-binding protein EST1A is an enzyme that in humans is encoded by the SMG6 gene on chromosome 17. It is ubiquitously expressed in many tissues and cell types. The C-terminus of the EST1A protein contains a PilT N-terminus (PIN) domain. This structure for this domain has been determined by X-ray crystallography. SMG6 functions to bind single-stranded DNA in telomere maintenance and single-stranded RNA in nonsense-mediated mRNA decay (NMD). The SMG6 gene also contains one of 27 SNPs associated with increased risk of coronary artery disease.
In a screen of the Bacillus subtilis genome for genes encoding ncRNAs, Saito et al. focused on 123 intergenic regions (IGRs) over 500 base pairs in length, the authors analyzed expression from these regions. Seven IGRs termed bsrC, bsrD, bsrE, bsrF, bsrG, bsrH and bsrI expressed RNAs smaller than 380 nt. All the small RNAs except BsrD RNA were expressed in transformed Escherichia coli cells harboring a plasmid with PCR-amplified IGRs of B. subtilis, indicating that their own promoters independently express small RNAs. Under non-stressed condition, depletion of the genes for the small RNAs did not affect growth. Although their functions are unknown, gene expression profiles at several time points showed that most of the genes except for bsrD were expressed during the vegetative phase, but undetectable during the stationary phase. Mapping the 5' ends of the 6 small RNAs revealed that the genes for BsrE, BsrF, BsrG, BsrH, and BsrI RNAs are preceded by a recognition site for RNA polymerase sigma factor σA.

A ribosome-inactivating protein (RIP) is a protein synthesis inhibitor that acts at the ribosome.
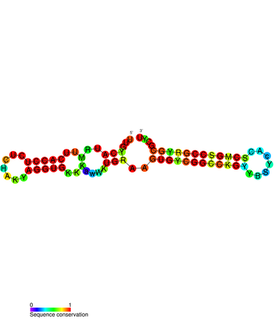
PtaRNA1 is a family of non-coding RNAs. Homologs of PtaRNA1 can be found in the proteobacteria families, Betaproteobacteria and Gammaproteobacteria. In all cases the PtaRNA1 is located anti-sense to a short protein-coding gene. In Xanthomonas campestris pv. vesicatoria, this gene is annotated as XCV2162 and is included in the plasmid toxin family of proteins.

A toxin-antitoxin system is a set of two or more closely linked genes that together encode both a "toxin" protein and a corresponding "antitoxin". Toxin-antitoxin systems are widely distributed in prokaryotes, and organisms often have them in multiple copies. When these systems are contained on plasmids – transferable genetic elements – they ensure that only the daughter cells that inherit the plasmid survive after cell division. If the plasmid is absent in a daughter cell, the unstable antitoxin is degraded and the stable toxic protein kills the new cell; this is known as 'post-segregational killing' (PSK).
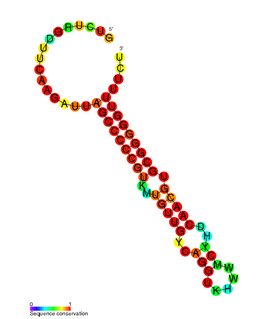
RdlD RNA is a family of small non-coding RNAs which repress the protein LdrD in a type I toxin-antitoxin system. It was discovered in Escherichia coli strain K-12 in a long direct repeat (LDR) named LDR-D. This locus encodes two products: a 35 amino acid peptide toxin (ldrD) and a 60 nucleotide RNA antitoxin. The 374nt toxin mRNA has a half-life of around 30 minutes while rdlD RNA has a half-life of only 2 minutes. This is in keeping with other type I toxin-antitoxin systems.
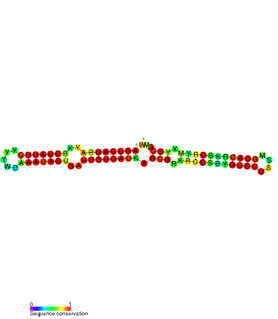
The SymE-SymR toxin-antitoxin system consists of a small symbiotic endonuclease toxin, SymE, and a non-coding RNA symbiotic RNA antitoxin, SymR, which inhibits SymE translation. SymE-SymR is a type I toxin-antitoxin system, and is under regulation by the antitoxin, SymR. The SymE-SymR complex is believed to play an important role in recycling damaged RNA and DNA. The relationship and corresponding structures of SymE and SymR provide insight into the mechanism of toxicity and overall role in prokaryotic systems.
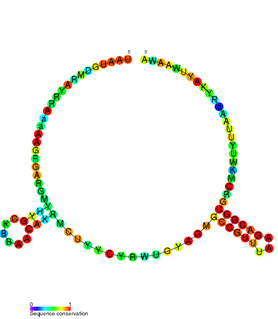
The TxpA/RatA toxin-antitoxin system was first identified in Bacillus subtilis. It consists of a non-coding 222nt sRNA called RatA and a protein toxin named TxpA.
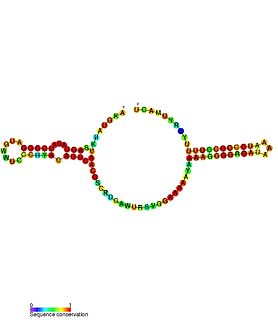
Escherichia coli contains a number of small RNAs located in intergenic regions of its genome. The presence of at least 55 of these has been verified experimentally. 275 potential sRNA-encoding loci were identified computationally using the QRNA program. These loci will include false positives, so the number of sRNA genes in E. coli is likely to be less than 275. A computational screen based on promoter sequences recognised by the sigma factor sigma 70 and on Rho-independent terminators predicted 24 putative sRNA genes, 14 of these were verified experimentally by northern blotting. The experimentally verified sRNAs included the well characterised sRNAs RprA and RyhB. Many of the sRNAs identified in this screen, including RprA, RyhB, SraB and SraL, are only expressed in the stationary phase of bacterial cell growth. A screen for sRNA genes based on homology to Salmonella and Klebsiella identified 59 candidate sRNA genes. From this set of candidate genes, microarray analysis and northern blotting confirmed the existence of 17 previously undescribed sRNAs, many of which bind to the chaperone protein Hfq and regulate the translation of RpoS. UptR sRNA transcribed from the uptR gene is implicated in suppressing extracytoplasmic toxicity by reducing the amount of membrane-bound toxic hybrid protein.
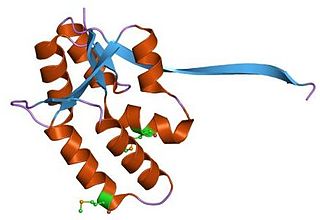
VapBC is the largest family of type II toxin-antitoxin system genetic loci in prokaryotes. VapBC operons consist of two genes: VapC encodes a toxic PilT N-terminus (PIN) domain, and VapB encodes a matching antitoxin. The toxins in this family are thought to perform RNA cleavage, which is inhibited by the co-expression of the antitoxin, in a manner analogous to a poison and antidote.

In molecular biology, the single-domain protein SUI1 is a translation initiation factor often found in the fungus, Saccharomyces cerevisiae but it is also found in other eukaryotes and prokaryotes as well as archaea. It is otherwise known as Eukaryotic translation initiation factor 1 (eIF1) in eukaryotes or YciH in bacteria.

In molecular biology, the CRM domain is an approximately 100-amino acid RNA-binding domain. The name CRM has been suggested to reflect the functions established for four characterised members of the family: Zea mays (Maize) CRS1, CAF1 and CAF2 proteins and the Escherichia coli protein YhbY. Proteins containing the CRM domain are found in eubacteria, archaea, and plants. The CRM domain is represented as a stand-alone protein in archaea and bacteria, and in single- and multi-domain proteins in plants. It has been suggested that prokaryotic CRM proteins existed as ribosome-associated proteins prior to the divergence of archaea and bacteria, and that they were co-opted in the plant lineage as RNA binding modules by incorporation into diverse protein contexts. Plant CRM domains are predicted to reside not only in the chloroplast, but also in the mitochondrion and the nucleo/cytoplasmic compartment. The diversity of the CRM domain family in plants suggests a diverse set of RNA targets.


















