Chemistry at Harvard Macromolecular Mechanics (CHARMM) is the name of a widely used set of force fields for molecular dynamics, and the name for the molecular dynamics simulation and analysis computer software package associated with them. The CHARMM Development Project involves a worldwide network of developers working with Martin Karplus and his group at Harvard to develop and maintain the CHARMM program. Licenses for this software are available, for a fee, to people and groups working in academia.

Molecular mechanics uses classical mechanics to model molecular systems. The Born–Oppenheimer approximation is assumed valid and the potential energy of all systems is calculated as a function of the nuclear coordinates using force fields. Molecular mechanics can be used to study molecule systems ranging in size and complexity from small to large biological systems or material assemblies with many thousands to millions of atoms.
Gaussian is a general purpose computational chemistry software package initially released in 1970 by John Pople and his research group at Carnegie Mellon University as Gaussian 70. It has been continuously updated since then. The name originates from Pople's use of Gaussian orbitals to speed up molecular electronic structure calculations as opposed to using Slater-type orbitals, a choice made to improve performance on the limited computing capacities of then-current computer hardware for Hartree–Fock calculations. The current version of the program is Gaussian 16. Originally available through the Quantum Chemistry Program Exchange, it was later licensed out of Carnegie Mellon University, and since 1987 has been developed and licensed by Gaussian, Inc.
In theoretical and computational chemistry, a basis set is a set of functions that is used to represent the electronic wave function in the Hartree–Fock method or density-functional theory in order to turn the partial differential equations of the model into algebraic equations suitable for efficient implementation on a computer.
MNDO, or Modified Neglect of Diatomic Overlap is a semi-empirical method for the quantum calculation of molecular electronic structure in computational chemistry. It is based on the Neglect of Diatomic Differential Overlap integral approximation. Similarly, this method replaced the earlier MINDO method. It is part of the MOPAC program and was developed in the group of Michael Dewar. It is also part of the AMPAC, GAMESS (US), PC GAMESS, GAMESS (UK), Gaussian, ORCA and CP2K programs.
Austin Model 1, or AM1, is a semi-empirical method for the quantum calculation of molecular electronic structure in computational chemistry. It is based on the Neglect of Differential Diatomic Overlap integral approximation. Specifically, it is a generalization of the modified neglect of differential diatomic overlap approximation. Related methods are PM3 and the older MINDO.

Ghemical is a computational chemistry software package written in C++ and released under the GNU General Public License. The program has graphical user interface based on GTK+2 and supports quantum mechanical and molecular mechanic models, with geometry optimization, molecular dynamics, and a large set of visualization tools. Ghemical relies on external code to provide the quantum-mechanical calculations — MOPAC provides the semi-empirical MNDO, MINDO, AM1, and PM3 methods, and MPQC methods based on Hartree–Fock calculations.

PQS is a general purpose quantum chemistry program. Its roots go back to the first ab initio gradient program developed in Professor Peter Pulay's group but now it is developed and distributed commercially by Parallel Quantum Solutions. There is a reduction in cost for academic users and a site license. Its strong points are geometry optimization, NMR chemical shift calculations, and large MP2 calculations, and high parallel efficiency on computing clusters. It includes many other capabilities including Density functional theory, the semiempirical methods, MINDO/3, MNDO, AM1 and PM3, Molecular mechanics using the SYBYL 5.0 Force Field, the quantum mechanics/molecular mechanics mixed method using the ONIOM method, natural bond orbital (NBO) analysis and COSMO solvation models. Recently, a highly efficient parallel CCSD(T) code for closed shell systems has been developed. This code includes many other post Hartree–Fock methods: MP2, MP3, MP4, CISD, CEPA, QCISD and so on.
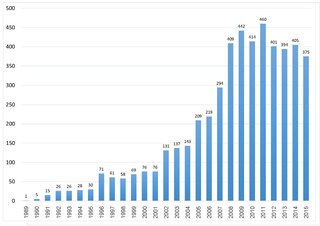
TURBOMOLE is an ab initio computational chemistry program that implements various quantum chemistry methods. It was initially developed by the group of Prof. Reinhart Ahlrichs at the University of Karlsruhe. In 2007, TURBOMOLE GmbH, founded by R. Ahlrichs, F. Furche, C. Hättig, W. Klopper, M. Sierka, and F. Weigend, took over the responsibility for the coordination of the scientific development of TURBOMOLE program, for which the company holds all copy and intellectual property rights. In 2018 David P. Tew joined the TURBOMOLE GmbH. Since 1987, this program is one of the useful tools as it involves in many fields of research including heterogeneous and homogeneous catalysis, organic and inorganic chemistry, spectroscopy as well as biochemistry. This can be illustrated by citation records of Ahlrich's 1989 publication which is more than 6700 times as of 18 July 2020. In the year 2014, the second Turbomole article has been published. The number of citations from both papers indicates that the Turbomole's user base is expanding.
Internal Coordinate Mechanics (ICM) is a software program and algorithm to predict low-energy conformations of molecules by sampling the space of internal coordinates defining molecular geometry. In ICM each molecule is constructed as a tree from an entry atom where each next atom is built iteratively from the preceding three atoms via three internal variables. The rings kept rigid or imposed via additional restraints. ICM is used for modelling peptides and interactions with substrates and coenzymes.

Spartan is a molecular modelling and computational chemistry application from Wavefunction. It contains code for molecular mechanics, semi-empirical methods, ab initio models, density functional models, post-Hartree–Fock models, and thermochemical recipes including G3(MP2) and T1. Quantum chemistry calculations in Spartan are powered by Q-Chem.
Semi-empirical quantum chemistry methods are based on the Hartree–Fock formalism, but make many approximations and obtain some parameters from empirical data. They are very important in computational chemistry for treating large molecules where the full Hartree–Fock method without the approximations is too expensive. The use of empirical parameters appears to allow some inclusion of electron correlation effects into the methods.
Merck molecular force field (MMFF) is a family of chemistry force fields developed by Merck Research Laboratories. They are based on the MM3 force field. MMFF is not optimized for one use, such as simulating proteins or small molecules, but tries to perform well for a wide range of organic chemistry calculations. The parameters in the force field have been derived from computational data consisting of approximately 2800 structures spanning a wide range of chemical classes.

The accessible surface area (ASA) or solvent-accessible surface area (SASA) is the surface area of a biomolecule that is accessible to a solvent. Measurement of ASA is usually described in units of square angstroms. ASA was first described by Lee & Richards in 1971 and is sometimes called the Lee-Richards molecular surface. ASA is typically calculated using the 'rolling ball' algorithm developed by Shrake & Rupley in 1973. This algorithm uses a sphere of a particular radius to 'probe' the surface of the molecule.
Quantum chemistry composite methods are computational chemistry methods that aim for high accuracy by combining the results of several calculations. They combine methods with a high level of theory and a small basis set with methods that employ lower levels of theory with larger basis sets. They are commonly used to calculate thermodynamic quantities such as enthalpies of formation, atomization energies, ionization energies and electron affinities. They aim for chemical accuracy which is usually defined as within 1 kcal/mol of the experimental value. The first systematic model chemistry of this type with broad applicability was called Gaussian-1 (G1) introduced by John Pople. This was quickly replaced by the Gaussian-2 (G2) which has been used extensively. The Gaussian-3 (G3) was introduced later.
AM1* is a semiempirical molecular orbital technique in computational chemistry. The method was developed by Timothy Clark and co-workers and published first in 2003.
Molecular modeling on GPU is the technique of using a graphics processing unit (GPU) for molecular simulations.
Biochemical and Organic Simulation System (BOSS) is a general-purpose molecular modeling program that performs molecular mechanics calculations, Metropolis Monte Carlo statistical mechanics simulations, and semiempirical Austin Model 1 (AM1), PM3, and PDDG/PM3 quantum mechanics calculations. The molecular mechanics calculations cover energy minimizations, normal mode analysis and conformational searching with the Optimized Potentials for Liquid Simulations (OPLS) force fields. BOSS is developed by Prof. William L. Jorgensen at Yale University, and distributed commercially by Cemcomco, LLC and Schrödinger, Inc.

Ascalaph Designer is a computer program for general purpose molecular modelling for molecular design and simulations. It provides a graphical environment for the common programs of quantum and classical molecular modelling ORCA, NWChem, Firefly, CP2K and MDynaMix . The molecular mechanics calculations cover model building, energy optimizations and molecular dynamics. Firefly covers a wide range of quantum chemistry methods. Ascalaph Designer is free and open-source software, released under the GNU General Public License, version 2 (GPLv2).
In computational chemistry, Natural Resonance Theory (NRT) is an iterative, variational functional embedded into the Natural Bond Orbital (NBO) program, commonly run in Gaussian, GAMESS, ORCA, Ampac and other software packages. NRT was developed in 1997 by Frank A. Weinhold and Eric D. Glendening, chemistry professors at University of Wisconsin-Madison and Indiana State University, respectively. Given a list of NBOs for an idealized natural Lewis structure, the NRT functional creates a list of Lewis resonance structures and calculates the resonance weights of each contributing resonance structure. Structural and chemical properties, such as bond order, valency, and bond polarity, may be calculated from resonance weights. Specifically, bond orders may be divided into their covalent and ionic contributions, while valency is the sum of bond orders of a given atom. This aims to provide quantitative results that agree with qualitative notions of chemical resonance. In contrast to the "wavefunction resonance theory", NRT uses the density matrix resonance theory, performing a superposition of density matrices to realize resonance. NRT has applications in ab initio calculations, including calculating the bond orders of intra- and intermolecular interactions and the resonance weights of radical isomers.








