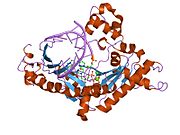
Heterochromatin is a tightly packed form of DNA or condensed DNA, which comes in multiple varieties. These varieties lie on a continuum between the two extremes of constitutive heterochromatin and facultative heterochromatin. Both play a role in the expression of genes. Because it is tightly packed, it was thought to be inaccessible to polymerases and therefore not transcribed; however, according to Volpe et al. (2002), and many other papers since, much of this DNA is in fact transcribed, but it is continuously turned over via RNA-induced transcriptional silencing (RITS). Recent studies with electron microscopy and OsO4 staining reveal that the dense packing is not due to the chromatin.

A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create two identical DNA duplexes from a single original DNA duplex. During this process, DNA polymerase "reads" the existing DNA strands to create two new strands that match the existing ones. These enzymes catalyze the chemical reaction

DNA repair is a collection of processes by which a cell identifies and corrects damage to the DNA molecules that encode its genome. In human cells, both normal metabolic activities and environmental factors such as radiation can cause DNA damage, resulting in tens of thousands of individual molecular lesions per cell per day. Many of these lesions cause structural damage to the DNA molecule and can alter or eliminate the cell's ability to transcribe the gene that the affected DNA encodes. Other lesions induce potentially harmful mutations in the cell's genome, which affect the survival of its daughter cells after it undergoes mitosis. As a consequence, the DNA repair process is constantly active as it responds to damage in the DNA structure. When normal repair processes fail, and when cellular apoptosis does not occur, irreparable DNA damage may occur. This can eventually lead to malignant tumors, or cancer as per the two-hit hypothesis.

Non-homologous end joining (NHEJ) is a pathway that repairs double-strand breaks in DNA. It is called "non-homologous" because the break ends are directly ligated without the need for a homologous template, in contrast to homology directed repair (HDR), which requires a homologous sequence to guide repair. NHEJ is active in both non-dividing and proliferating cells, while HDR is not readily accessible in non-dividing cells. The term "non-homologous end joining" was coined in 1996 by Moore and Haber.

Proliferating cell nuclear antigen (PCNA) is a DNA clamp that acts as a processivity factor for DNA polymerase δ in eukaryotic cells and is essential for replication. PCNA is a homotrimer and achieves its processivity by encircling the DNA, where it acts as a scaffold to recruit proteins involved in DNA replication, DNA repair, chromatin remodeling and epigenetics.

Pyrimidine dimers represent molecular lesions originating from thymine or cytosine bases within DNA, resulting from photochemical reactions. These lesions, commonly linked to direct DNA damage, are induced by ultraviolet light (UV), particularly UVC, result in the formation of covalent bonds between adjacent nitrogenous bases along the nucleotide chain near their carbon–carbon double bonds, the photo-coupled dimers are fluorescent. Such dimerization, which can also occur in double-stranded RNA (dsRNA) involving uracil or cytosine, leads to the creation of cyclobutane pyrimidine dimers (CPDs) and 6–4 photoproducts. These pre-mutagenic lesions modify the DNA helix structure, resulting in abnormal non-canonical base pairing and, consequently, adjacent thymines or cytosines in DNA will form a cyclobutane ring when joined together and cause a distortion in the DNA. This distortion prevents DNA replication and transcription mechanisms beyond the dimerization site.
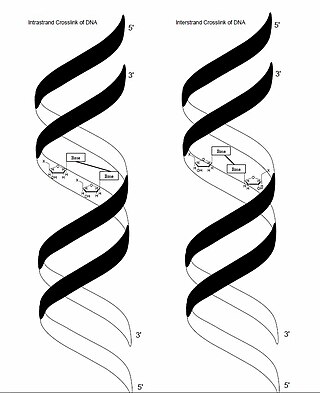
In genetics, crosslinking of DNA occurs when various exogenous or endogenous agents react with two nucleotides of DNA, forming a covalent linkage between them. This crosslink can occur within the same strand (intrastrand) or between opposite strands of double-stranded DNA (interstrand). These adducts interfere with cellular metabolism, such as DNA replication and transcription, triggering cell death. These crosslinks can, however, be repaired through excision or recombination pathways.
Richard D. Wood is an American molecular biologist specializing in research on DNA repair and mutation. He is known for pioneering studies on nucleotide excision repair (NER), particularly for reconstituting the minimum set of proteins involved in this process, identifying proliferating cell nuclear antigen (PCNA) as part of the NER complex and identifying mammalian repair polymerases.
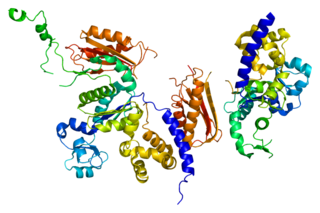
DNA polymerase kappa is a DNA polymerase that in humans is encoded by the POLK gene. It is involved in translesion synthesis.

Upstream binding transcription factor (UBTF), or upstream binding factor (UBF), is a protein that in humans is encoded by the UBTF gene.

G/T mismatch-specific thymine DNA glycosylase is an enzyme that in humans is encoded by the TDG gene. Several bacterial proteins have strong sequence homology with this protein.

DNA repair protein REV1 is a protein that in humans is encoded by the REV1 gene.

DNA polymerase theta is an enzyme that in humans is encoded by the POLQ gene. This polymerase plays a key role in one of the three major double strand break repair pathways: theta-mediated end joining (TMEJ). Most double-strand breaks are repaired by non-homologous end joining (NHEJ) or homology directed repair (HDR). However, in some contexts, NHEJ and HR are insufficient and TMEJ is the only solution to repair the break. TMEJ is often described as alternative NHEJ, but differs in that it lacks a requirement for the Ku heterodimer, and it can only act on resected DNA ends. Following annealing of short regions on the DNA overhangs, DNA polymerase theta catalyzes template-dependent DNA synthesis across the broken ends, stabilizing the paired structure.

Elongator complex protein 3, also named KAT9, is a protein that in humans is encoded by the ELP3 gene. ELP3 is the catalytic histone acetyltransferase subunit of the RNA polymerase II elongator complex, which is a component of the RNA polymerase II holoenzyme and is involved in transcriptional elongation. ELP3 supports the migration and branching of projection neurons through acetylation of alpha-tubulin in the developing cerebral cortex. In mammals, ELP3 is important for paternal DNA demethylation after fertilization. ELP3 is potentially involved in cellular redox homeostasis by mediating the acetylation of glucose-6-phosphate dehydrogenase. Besides, ELP3 may play a role in chromatin remodeling and is involved in acetylation of histones H3 and probably H4.

Protein reversionless 3-like (REV3L) also known as DNA polymerase zeta catalytic subunit (POLZ) is an enzyme that in humans is encoded by the REV3L gene.

DNA polymerase eta, is a protein that in humans is encoded by the POLH gene.

Polymerase I and transcript release factor, also known as Cavin1, Cavin-1 or PTRF, is a protein which in humans is encoded by the PTRF gene.
DNA polymerase IV is a prokaryotic polymerase that is involved in mutagenesis and is encoded by the dinB gene. It exhibits no 3′→5′ exonuclease (proofreading) activity and hence is error prone. In E. coli, DNA polymerase IV is involved in non-targeted mutagenesis. Pol IV is a Family Y polymerase expressed by the dinB gene that is switched on via SOS induction caused by stalled polymerases at the replication fork. During SOS induction, Pol IV production is increased tenfold and one of the functions during this time is to interfere with Pol III holoenzyme processivity. This creates a checkpoint, stops replication, and allows time to repair DNA lesions via the appropriate repair pathway. Another function of Pol IV is to perform translesion synthesis at the stalled replication fork like, for example, bypassing N2-deoxyguanine adducts at a faster rate than transversing undamaged DNA. Cells lacking dinB gene have a higher rate of mutagenesis caused by DNA damaging agents.

PrimPol is a protein encoded by the PRIMPOL gene in humans. PrimPol is a eukaryotic protein with both DNA polymerase and DNA Primase activities involved in translesion DNA synthesis. It is the first eukaryotic protein to be identified with priming activity using deoxyribonucleotides. It is also the first protein identified in the mitochondria to have translesion DNA synthesis activities.

Deepak Thankappan Nair is an Indian Structural Biologist and a scientist at Regional Centre for Biotechnology. He is known for his studies on DNA and RNA polymerases. Deepak was a Ramanujan fellow of the Science and Engineering Research Board (2008–2013) and a recipient of the National BioScience Award for Career Development. The Council of Scientific and Industrial Research, the apex agency of the Government of India for scientific research, awarded him the Shanti Swarup Bhatnagar Prize for Science and Technology, one of the highest Indian science awards, for his contributions to biological sciences in 2017. He was inducted as a fellow of the Indian National Science Academy in December, 2022.