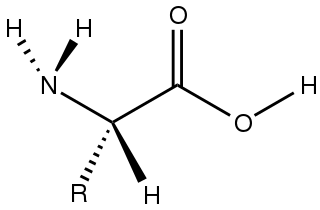
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although over 500 amino acids exist in nature, by far the most important are the 22 α-amino acids incorporated into proteins. Only these 22 appear in the genetic code of life.

Peptides are short chains of amino acids linked by peptide bonds. A polypeptide is a longer, continuous, unbranched peptide chain. Polypeptides that have a molecular mass of 10,000 Da or more are called proteins. Chains of fewer than twenty amino acids are called oligopeptides, and include dipeptides, tripeptides, and tetrapeptides.

Protein primary structure is the linear sequence of amino acids in a peptide or protein. By convention, the primary structure of a protein is reported starting from the amino-terminal (N) end to the carboxyl-terminal (C) end. Protein biosynthesis is most commonly performed by ribosomes in cells. Peptides can also be synthesized in the laboratory. Protein primary structures can be directly sequenced, or inferred from DNA sequences.

In molecular biology, post-translational modification (PTM) is the covalent process of changing proteins following protein biosynthesis. PTMs may involve enzymes or occur spontaneously. Proteins are created by ribosomes, which translate mRNA into polypeptide chains, which may then change to form the mature protein product. PTMs are important components in cell signalling, as for example when prohormones are converted to hormones.

Proteinogenic amino acids are amino acids that are incorporated biosynthetically into proteins during translation. The word "proteinogenic" means "protein creating". Throughout known life, there are 22 genetically encoded (proteinogenic) amino acids, 20 in the standard genetic code and an additional 2 that can be incorporated by special translation mechanisms.

Dehydroalanine is a dehydroamino acid. It does not exist in its free form, but it occurs naturally as a residue found in peptides of microbial origin. As an amino acid residue, it is unusual because it has an unsaturated backbone.
Native Chemical Ligation (NCL) is an important extension of the chemical ligation concept for constructing a larger polypeptide chain by the covalent condensation of two or more unprotected peptides segments. Native chemical ligation is the most effective method for synthesizing native or modified proteins of typical size.
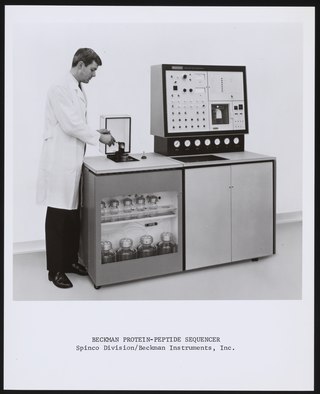
Protein sequencing is the practical process of determining the amino acid sequence of all or part of a protein or peptide. This may serve to identify the protein or characterize its post-translational modifications. Typically, partial sequencing of a protein provides sufficient information to identify it with reference to databases of protein sequences derived from the conceptual translation of genes.

Chemical biology is a scientific discipline between the fields of chemistry and biology. The discipline involves the application of chemical techniques, analysis, and often small molecules produced through synthetic chemistry, to the study and manipulation of biological systems. Although often confused with biochemistry, which studies the chemistry of biomolecules and regulation of biochemical pathways within and between cells, chemical biology remains distinct by focusing on the application of chemical tools to address biological questions.

In immunology, epitope mapping is the process of experimentally identifying the binding site, or epitope, of an antibody on its target antigen. Identification and characterization of antibody binding sites aid in the discovery and development of new therapeutics, vaccines, and diagnostics. Epitope characterization can also help elucidate the binding mechanism of an antibody and can strengthen intellectual property (patent) protection. Experimental epitope mapping data can be incorporated into robust algorithms to facilitate in silico prediction of B-cell epitopes based on sequence and/or structural data.

Citrullination or deimination is the conversion of the amino acid arginine in a protein into the amino acid citrulline. Citrulline is not one of the 20 standard amino acids encoded by DNA in the genetic code. Instead, it is the result of a post-translational modification. Citrullination is distinct from the formation of the free amino acid citrulline as part of the urea cycle or as a byproduct of enzymes of the nitric oxide synthase family.
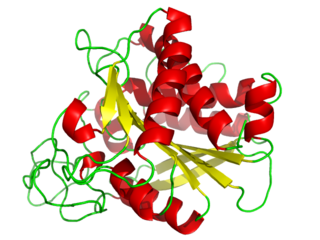
A carboxypeptidase is a protease enzyme that hydrolyzes (cleaves) a peptide bond at the carboxy-terminal (C-terminal) end of a protein or peptide. This is in contrast to an aminopeptidases, which cleave peptide bonds at the N-terminus of proteins. Humans, animals, bacteria and plants contain several types of carboxypeptidases that have diverse functions ranging from catabolism to protein maturation. At least two mechanisms have been discussed.
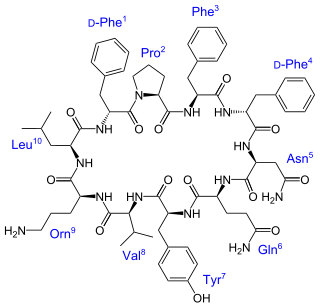
Tyrocidine is a mixture of cyclic decapeptides produced by the bacteria Brevibacillus brevis found in soil. It can be composed of 4 different amino acid sequences, giving tyrocidine A–D. Tyrocidine is the major constituent of tyrothricin, which also contains gramicidin. Tyrocidine was the first commercially available antibiotic, but has been found to be toxic toward human blood and reproductive cells. The function of tyrocidine within its host B. brevis is thought to be regulation of sporulation.
α-Bungarotoxin is one of the bungarotoxins, components of the venom of the elapid Taiwanese banded krait snake. It is a type of α-neurotoxin, a neurotoxic protein that is known to bind competitively and in a relatively irreversible manner to the nicotinic acetylcholine receptor found at the neuromuscular junction, causing paralysis, respiratory failure, and death in the victim. It has also been shown to play an antagonistic role in the binding of the α7 nicotinic acetylcholine receptor in the brain, and as such has numerous applications in neuroscience research.
Protein metabolism denotes the various biochemical processes responsible for the synthesis of proteins and amino acids (anabolism), and the breakdown of proteins by catabolism.

In enzymology, an alanine racemase is an enzyme that catalyzes the chemical reaction
Glycopeptides are peptides that contain carbohydrate moieties (glycans) covalently attached to the side chains of the amino acid residues that constitute the peptide.

In biochemistry, non-coded or non-proteinogenic amino acids are distinct from the 22 proteinogenic amino acids, which are naturally encoded in the genome of organisms for the assembly of proteins. However, over 140 non-proteinogenic amino acids occur naturally in proteins and thousands more may occur in nature or be synthesized in the laboratory. Chemically synthesized amino acids can be called unnatural amino acids. Unnatural amino acids can be synthetically prepared from their native analogs via modifications such as amine alkylation, side chain substitution, structural bond extension cyclization, and isosteric replacements within the amino acid backbone. Many non-proteinogenic amino acids are important:

Bottromycin is a macrocyclic peptide with antibiotic activity. It was first discovered in 1957 as a natural product isolated from Streptomyces bottropensis. It has been shown to inhibit methicillin-resistant Staphylococcus aureus (MRSA) and vancomycin-resistant Enterococci (VRE) among other Gram-positive bacteria and mycoplasma. Bottromycin is structurally distinct from both vancomycin, a glycopeptide antibiotic, and methicillin, a beta-lactam antibiotic.
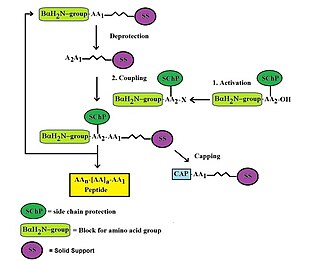
Custom peptide synthesis is the commercial production of peptides for use in biochemistry, biology, biotechnology, pharmacology and molecular medicine. Custom peptide synthesis provides synthetic peptides as valuable tools to biomedical laboratories. Synthetic oligopeptides are used extensively in research for structure-function analysis, for the development of binding assays, the study of receptor agonist/antagonists or as immunogens for the production of specific antibodies. Generally, peptides are synthesized by coupling the carboxyl group or C-terminus of one amino acid to the amino group or N-terminus of another using automated solid phase peptide synthesis chemistries. However, liquid phase synthesis may also be used for specific needs.
















