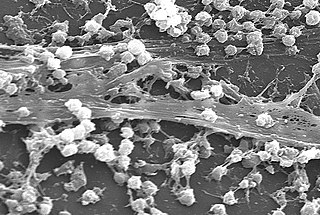
A biofilm comprises any syntrophic consortium of microorganisms in which cells stick to each other and often also to a surface. These adherent cells become embedded within a slimy extracellular matrix that is composed of extracellular polymeric substances (EPSs). The cells within the biofilm produce the EPS components, which are typically a polymeric conglomeration of extracellular polysaccharides, proteins, lipids and DNA. Because they have three-dimensional structure and represent a community lifestyle for microorganisms, they have been metaphorically described as "cities for microbes".

Pseudomonas aeruginosa is a common encapsulated, Gram-negative, aerobic–facultatively anaerobic, rod-shaped bacterium that can cause disease in plants and animals, including humans. A species of considerable medical importance, P. aeruginosa is a multidrug resistant pathogen recognized for its ubiquity, its intrinsically advanced antibiotic resistance mechanisms, and its association with serious illnesses – hospital-acquired infections such as ventilator-associated pneumonia and various sepsis syndromes.

Filamentation is the anomalous growth of certain bacteria, such as Escherichia coli, in which cells continue to elongate but do not divide. The cells that result from elongation without division have multiple chromosomal copies.

The rsmY RNA family is a set of related non-coding RNA genes, that like RsmZ, is regulated by the GacS/GacA signal transduction system in the plant-beneficial soil bacterium and biocontrol model organism Pseudomonas fluorescens CHA0. GacA/GacS target genes are translationally repressed by the small RNA binding protein RsmA. RsmY and RsmZ RNAs bind RsmA to relieve this repression and so enhance secondary metabolism and biocontrol traits.
Pseudomonas sRNA are non-coding RNAs (ncRNA) that were predicted by the bioinformatic program SRNApredict2. This program identifies putative sRNAs by searching for co-localization of genetic features commonly associated with sRNA-encoding genes and the expression of the predicted sRNAs was subsequently confirmed by Northern blot analysis. These sRNAs have been shown to be conserved across several pseudomonas species but their function is yet to be determined. Using Tet-Trap genetic approach RNAT genes post-transcriptionally regulated by temperature upshift were identified: ptxS and PA5194.
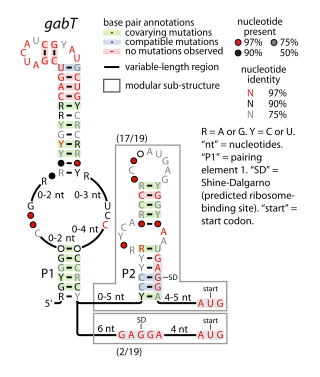
The gabT RNA motif is the name of a conserved RNA structure identified by bioinformatics whose function is unknown. The gabT motif has been detected exclusively in bacteria within the genus Pseudomonas, and is found only upstream of gabT genes, and downstream to gabD genes.

The gyrA RNA motif is a conserved RNA structure identified by bioinformatics. The RNAs are present in multiple species of bacteria within the order Pseudomonadales. This order contains the genus Pseudomonas, which includes the opportunistic human pathogen Pseudomonas aeruginosa and Pseudomonas syringae, a plant pathogen.
The Pseudomon-groES RNA motif is a conserved RNA structure identified in certain bacteria using bioinformatics. It is found in most species within the family Pseudomonadaceae, and is consistently located in the 5' untranslated regions of operons that contain groES genes. RNA transcripts of the groES genes in Pseudomonas aeruginosa where shown experimentally to be initiated at one of two start sites, from promoters called "P1" and "P2". The Pseudomon-groES RNA is in the 5' UTR of transcripts initiated from the P1 site, but is truncated in P2 transcripts. groES genes are involved in the cellular response to heat shock, but it is not thought that the Pseudomonas-groES RNA motif is involved in heat shock regulation. However, it is thought that the motif might regulate groES genes in response to other stimuli.

The rsmX gene is part of the Rsm/Csr family of non-coding RNAs (ncRNAs). Members of the Rsm/Csr family are present in a diverse range of bacteria, including Escherichia coli, Erwinia, Salmonella, Vibrio and Pseudomonas. These ncRNAs act by sequestering translational repressor proteins, called RsmA, activating expression of downstream genes that would normally be blocked by the repressors. Sequestering of target proteins is dependent upon exposed GGA motifs in the stem loops of the ncRNAs. Typically, the activated genes are involved in secondary metabolism, biofilm formation and motility.
Bacterial small RNAs (bsRNA) are small RNAs produced by bacteria; they are 50- to 500-nucleotide non-coding RNA molecules, highly structured and containing several stem-loops. Numerous sRNAs have been identified using both computational analysis and laboratory-based techniques such as Northern blotting, microarrays and RNA-Seq in a number of bacterial species including Escherichia coli, the model pathogen Salmonella, the nitrogen-fixing alphaproteobacterium Sinorhizobium meliloti, marine cyanobacteria, Francisella tularensis, Streptococcus pyogenes, the pathogen Staphylococcus aureus, and the plant pathogen Xanthomonas oryzae pathovar oryzae. Bacterial sRNAs affect how genes are expressed within bacterial cells via interaction with mRNA or protein, and thus can affect a variety of bacterial functions like metabolism, virulence, environmental stress response, and structure.

Rhamnolipids are a class of glycolipid produced by Pseudomonas aeruginosa, amongst other organisms, frequently cited as bacterial surfactants. They have a glycosyl head group, in this case a rhamnose moiety, and a 3-(hydroxyalkanoyloxy)alkanoic acid (HAA) fatty acid tail, such as 3-hydroxydecanoic acid.

CrcZ is a small RNA found in Pseudomonas bacteria, which acts as a global regulator of carbon catabolite repression. In P. aeruginosa, CrcZ is responsible for sequestering the protein Crc. Crc is an RNA-binding global regulator, which acts by inhibiting the translation of the transcriptional regulator AlkS.
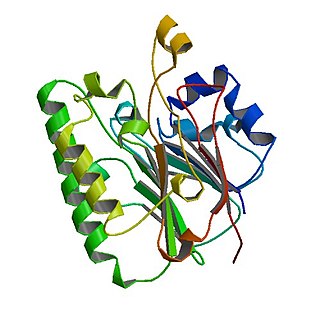
The Catabolite repression control (Crc) protein participates in suppressing expression of several genes involved in utilization of carbon sources in Pseudomonas bacteria. Presence of organic acids triggers activation of Crc and in conjunction with the Hfq protein genes that metabolize a given carbon source are downregulated until another more favorable carbon source is depleted. Crc-mediated regulation impact processes such as biofilm formation, virulence and antibiotic susceptibility.
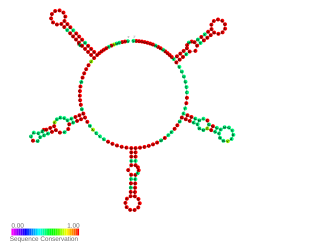
NrsZ is a bacterial small RNA found in the opportunistic pathogen Pseudomonas aeruginosa PAO1. Its transcription is induced during nitrogen limitation by the NtrB/C two-component system together with the alternative sigma factor RpoN. NrsZ by activating rhlA positively regulates the production of rhamnolipid surfactants needed for swarming motility.
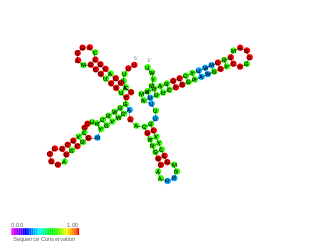
RsmW is a part of the Rsm/Csr family of non-coding RNAs (ncRNAs) discovered in Pseudomonas aeruginosa. It specifically binds to RsmA protein in vitro, restores biofilm production and partially complements the loss of RsmY and RsmZ in rsmY/rsmZ double mutant in regards to their contribution to swarming. Compared to RsmY and RsmZ its production is induced in high temperatures and rsmW is not transcriptionally activated by GacA.
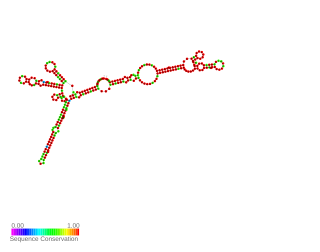
AsponA is a small asRNA transcribed antisense to the penicillin-binding protein 1A gene called ponA. It was identified by RNAseq and the expression was validated by 5' and 3' RACE experiments in Pseudomanas aeruginosa. AsponA expression was up or down regulated under different antibiotic stress. Owing to its location it may be able to prevent the transcription or translation of the opposite gene. Study by Wurtzel et al. and Ferrara et al. also detected its expression.

Twitching motility is a form of crawling bacterial motility used to move over surfaces. Twitching is mediated by the activity of hair-like filaments called type IV pili which extend from the cell's exterior, bind to surrounding solid substrates, and retract, pulling the cell forwards in a manner similar to the action of a grappling hook. The name twitching motility is derived from the characteristic jerky and irregular motions of individual cells when viewed under the microscope. It has been observed in many bacterial species, but is most well studied in Pseudomonas aeruginosa, Neisseria gonorrhoeae and Myxococcus xanthus. Active movement mediated by the twitching system has been shown to be an important component of the pathogenic mechanisms of several species.
SrbA (sRNA regulator of biofilms A) is a small regulatory non-coding RNA identified in pathogenic Pseudomonas aeruginosa. It is important for biofilm formation and pathogenicity. Bacterial strain with deleted SrbA had reduced biofilm mass. As the ability to form biofilms can contribute to the ability a pathogen to thrive within the host, the C. elegans hosts infected with the srbA deleted strain displayed significantly lower mortality rate than the wild-type strain. However, the deletion of srbA had no effect on growth or antibiotic resistance in P. aeruginosa.
Kalai Mathee is a professor at Florida International University, joint editor-in-chief of the Journal of Medical Microbiology, and an elected fellow of the American Academy of Microbiology. She is known for her research on bacterial infections caused by Pseudomonas aeruginosa.
Diffusible signal factor (DSF) is found in several gram-negative bacteria and play a role in the formation of biofilms, motility, virulence, and antibiotic resistance. Xanthomonas campestris was the first bacteria known to have DSF. The synthesis of the DSF can be seen in rpfF and rpfB enzymes. An understanding of the DSF signaling mechanism could lead to further disease control.














