
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity.

Protein biosynthesis is a core biological process, occurring inside cells, balancing the loss of cellular proteins through the production of new proteins. Proteins perform a number of critical functions as enzymes, structural proteins or hormones. Protein synthesis is a very similar process for both prokaryotes and eukaryotes but there are some distinct differences.

In biology, epigenetics is the study of stable phenotypic changes that do not involve alterations in the DNA sequence. The Greek prefix epi- in epigenetics implies features that are "on top of" or "in addition to" the traditional genetic basis for inheritance. Epigenetics most often involves changes that affect gene activity and expression, but the term can also be used to describe any heritable phenotypic change. Such effects on cellular and physiological phenotypic traits may result from external or environmental factors, or be part of normal development.

The central dogma of molecular biology is an explanation of the flow of genetic information within a biological system. It is often stated as "DNA makes RNA, and RNA makes protein", although this is not its original meaning. It was first stated by Francis Crick in 1957, then published in 1958:
The Central Dogma. This states that once "information" has passed into protein it cannot get out again. In more detail, the transfer of information from nucleic acid to nucleic acid, or from nucleic acid to protein may be possible, but transfer from protein to protein, or from protein to nucleic acid is impossible. Information means here the precise determination of sequence, either of bases in the nucleic acid or of amino acid residues in the protein.

Susan Lee Lindquist, ForMemRS was an American professor of biology at MIT specializing in molecular biology, particularly the protein folding problem within a family of molecules known as heat-shock proteins, and prions. Lindquist was a member and former director of the Whitehead Institute and was awarded the National Medal of Science in 2010.

A frameshift mutation is a genetic mutation caused by indels of a number of nucleotides in a DNA sequence that is not divisible by three. Due to the triplet nature of gene expression by codons, the insertion or deletion can change the reading frame, resulting in a completely different translation from the original. The earlier in the sequence the deletion or insertion occurs, the more altered the protein. A frameshift mutation is not the same as a single-nucleotide polymorphism in which a nucleotide is replaced, rather than inserted or deleted. A frameshift mutation will in general cause the reading of the codons after the mutation to code for different amino acids. The frameshift mutation will also alter the first stop codon encountered in the sequence. The polypeptide being created could be abnormally short or abnormally long, and will most likely not be functional.

Structural inheritance or cortical inheritance is the transmission of an epigenetic trait in a living organism by a self-perpetuating spatial structures. This is in contrast to the transmission of digital information such as is found in DNA sequences, which accounts for the vast majority of known genetic variation.
Evolvability is defined as the capacity of a system for adaptive evolution. Evolvability is the ability of a population of organisms to not merely generate genetic diversity, but to generate adaptive genetic diversity, and thereby evolve through natural selection.

Silent mutations are mutations in DNA that do not have an observable effect on the organism's phenotype. They are a specific type of neutral mutation. The phrase silent mutation is often used interchangeably with the phrase synonymous mutation; however, synonymous mutations are not always silent, nor vice versa. Synonymous mutations can affect transcription, splicing, mRNA transport, and translation, any of which could alter phenotype, rendering the synonymous mutation non-silent. The substrate specificity of the tRNA to the rare codon can affect the timing of translation, and in turn the co-translational folding of the protein. This is reflected in the codon usage bias that is observed in many species. Mutations that cause the altered codon to produce an amino acid with similar functionality are often classified as silent; if the properties of the amino acid are conserved, this mutation does not usually significantly affect protein function.
This is a list of topics in molecular biology. See also index of biochemistry articles.

Evolutionary capacitance is the storage and release of variation, just as electric capacitors store and release charge. Living systems are robust to mutations. This means that living systems accumulate genetic variation without the variation having a phenotypic effect. But when the system is disturbed, robustness breaks down, and the variation has phenotypic effects and is subject to the full force of natural selection. An evolutionary capacitor is a molecular switch mechanism that can "toggle" genetic variation between hidden and revealed states. If some subset of newly revealed variation is adaptive, it becomes fixed by genetic assimilation. After that, the rest of variation, most of which is presumably deleterious, can be switched off, leaving the population with a newly evolved advantageous trait, but no long-term handicap. For evolutionary capacitance to increase evolvability in this way, the switching rate should not be faster than the timescale of genetic assimilation.
Bacterial translation is the process by which messenger RNA is translated into proteins in bacteria.

PRNP is the human gene encoding for the major prion protein PrP, also known as CD230. Expression of the protein is most predominant in the nervous system but occurs in many other tissues throughout the body.

A fungal prion is a prion that infects fungal hosts. Fungal prions are naturally occurring proteins that can switch between multiple, structurally distinct conformations, at least one of which is self-propagating and transmissible to other prions. This transmission of protein state represents an epigenetic phenomenon where information is encoded in the protein structure itself, instead of in nucleic acids. Several prion-forming proteins have been identified in fungi, primarily in the yeast Saccharomyces cerevisiae. These fungal prions are generally considered benign, and in some cases even confer a selectable advantage to the organism.
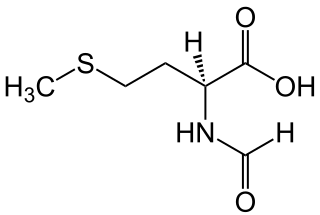
N-Formylmethionine is a derivative of the amino acid methionine in which a formyl group has been added to the amino group. It is specifically used for initiation of protein synthesis from bacterial and organellar genes, and may be removed post-translationally.

EF-Tu is a prokaryotic elongation factor responsible for catalyzing the binding of an aminoacyl-tRNA (aa-tRNA) to the ribosome. It is a G-protein, and facilitates the selection and binding of an aa-tRNA to the A-site of the ribosome. As a reflection of its crucial role in translation, EF-Tu is one of the most abundant and highly conserved proteins in prokaryotes. It is found in eukaryotic mitochrondria as TUFM.

Eukaryotic translation termination factor 1 (eRF1), also known as TB3-1, is a protein that in humans is encoded by the ETF1 gene.
Sup45p is the Saccharomyces cerevisiae eukaryotic translation termination factor. More specifically, it is the yeast eukaryotic release factor 1 (eRF1). Its job is to recognize stop codons in RNA and bind to them. It binds to the Sup35p protein and then takes on the shape of a tRNA molecule so that it can safety incorporate itself into the A site of the Ribosome to disruptits flow and "release" the protein and end translation.
NatA acetyltransferase(Nα acetyltransferase), is an enzyme that serves to catalyze the addition of acetyl groups to various proteins emerging from the ribosome. Upon translation, the NatA binds to the ribosome and then "stretches" to the front end of the forming, or nascent, polypeptide, where it adds this acetyl group. This acetyl group is added to the front end, or N-terminus of the new protein.

In biochemistry and molecular biology, SDD-AGE is short for Semi-Denaturating Detergent Agarose Gel Electrophoresis. This is a method for detecting and characterizing large protein polymers which are stable in 2% SDS at room temperature, unlike most large protein complexes. This method is very useful for studying prions and amyloids, which are characterized by the formation of proteinaceous polymers. Agarose is used for the gel since the SDS-resistant polymers are large and cannot enter a conventional polyacrylamide gel, which has small pores. Agarose on the other hand has large pores, which allows for the separation of polymers.














