Related Research Articles

In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.

Transcription is the process of copying a segment of DNA into RNA. The segments of DNA transcribed into RNA molecules that can encode proteins are said to produce messenger RNA (mRNA). Other segments of DNA are copied into RNA molecules called non-coding RNAs (ncRNAs). mRNA comprises only 1–3% of total RNA samples. Less than 2% of the human genome can be transcribed into mRNA, while at least 80% of mammalian genomic DNA can be actively transcribed, with the majority of this 80% considered to be ncRNA.

In molecular biology, RNA polymerase, or more specifically DNA-directed/dependent RNA polymerase (DdRP), is an enzyme that catalyzes the chemical reactions that synthesize RNA from a DNA template.

A ρ factor is a bacterial protein involved in the termination of transcription. Rho factor binds to the transcription terminator pause site, an exposed region of single stranded RNA after the open reading frame at C-rich/G-poor sequences that lack obvious secondary structure.
In genetics, a transcription terminator is a section of nucleic acid sequence that marks the end of a gene or operon in genomic DNA during transcription. This sequence mediates transcriptional termination by providing signals in the newly synthesized transcript RNA that trigger processes which release the transcript RNA from the transcriptional complex. These processes include the direct interaction of the mRNA secondary structure with the complex and/or the indirect activities of recruited termination factors. Release of the transcriptional complex frees RNA polymerase and related transcriptional machinery to begin transcription of new mRNAs.
Polyadenylation is the addition of a poly(A) tail to an RNA transcript, typically a messenger RNA (mRNA). The poly(A) tail consists of multiple adenosine monophosphates; in other words, it is a stretch of RNA that has only adenine bases. In eukaryotes, polyadenylation is part of the process that produces mature mRNA for translation. In many bacteria, the poly(A) tail promotes degradation of the mRNA. It, therefore, forms part of the larger process of gene expression.

Exonucleases are enzymes that work by cleaving nucleotides one at a time from the end (exo) of a polynucleotide chain. A hydrolyzing reaction that breaks phosphodiester bonds at either the 3′ or the 5′ end occurs. Its close relative is the endonuclease, which cleaves phosphodiester bonds in the middle (endo) of a polynucleotide chain. Eukaryotes and prokaryotes have three types of exonucleases involved in the normal turnover of mRNA: 5′ to 3′ exonuclease (Xrn1), which is a dependent decapping protein; 3′ to 5′ exonuclease, an independent protein; and poly(A)-specific 3′ to 5′ exonuclease.
Eukaryotic translation is the biological process by which messenger RNA is translated into proteins in eukaryotes. It consists of four phases: initiation, elongation, termination, and recapping.
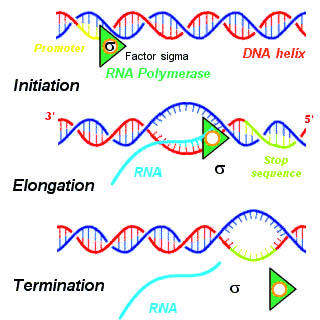
A termination signal is a sequence that signals the end of transcription or translation. Termination signals are found at the end of the part of the chromosome being transcribed during transcription of mRNA. Termination signals bring a stop to transcription, ensuring that only gene-encoding parts of the chromosome are transcribed. Transcription begins at the promoter when RNA polymerase, an enzyme that facilitates transcription of DNA into mRNA, binds to a promoter, unwinds the helical structure of the DNA, and uses the single-stranded DNA as a template to synthesize RNA. Once RNA polymerase reaches the termination signal, transcription is terminated. In bacteria, there are two main types of termination signals: intrinsic and factor-dependent terminators. In the context of translation, a termination signal is the stop codon on the mRNA that elicits the release of the growing peptide from the ribosome.
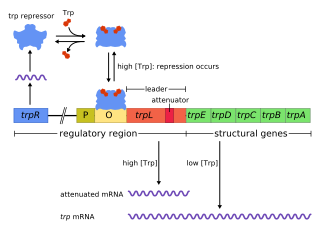
The trp operon is a group of genes that are transcribed together, encoding the enzymes that produce the amino acid tryptophan in bacteria. The trp operon was first characterized in Escherichia coli, and it has since been discovered in many other bacteria. The operon is regulated so that, when tryptophan is present in the environment, the genes for tryptophan synthesis are repressed.
Antitermination is the prokaryotic cell's aid to fix premature termination of RNA synthesis during the transcription of RNA. It occurs when the RNA polymerase ignores the termination signal and continues elongating its transcript until a second signal is reached. Antitermination provides a mechanism whereby one or more genes at the end of an operon can be switched either on or off, depending on the polymerase either recognizing or not recognizing the termination signal.

Prokaryotic DNA Replication is the process by which a prokaryote duplicates its DNA into another copy that is passed on to daughter cells. Although it is often studied in the model organism E. coli, other bacteria show many similarities. Replication is bi-directional and originates at a single origin of replication (OriC). It consists of three steps: Initiation, elongation, and termination.

Bacterial transcription is the process in which a segment of bacterial DNA is copied into a newly synthesized strand of messenger RNA (mRNA) with use of the enzyme RNA polymerase.

Eukaryotic transcription is the elaborate process that eukaryotic cells use to copy genetic information stored in DNA into units of transportable complementary RNA replica. Gene transcription occurs in both eukaryotic and prokaryotic cells. Unlike prokaryotic RNA polymerase that initiates the transcription of all different types of RNA, RNA polymerase in eukaryotes comes in three variations, each translating a different type of gene. A eukaryotic cell has a nucleus that separates the processes of transcription and translation. Eukaryotic transcription occurs within the nucleus where DNA is packaged into nucleosomes and higher order chromatin structures. The complexity of the eukaryotic genome necessitates a great variety and complexity of gene expression control.

Intrinsic, or rho-independent termination, is a process in prokaryotes to signal the end of transcription and release the newly constructed RNA molecule. In prokaryotes such as E. coli, transcription is terminated either by a rho-dependent process or rho-independent process. In the Rho-dependent process, the rho-protein locates and binds the signal sequence in the mRNA and signals for cleavage. Contrarily, intrinsic termination does not require a special protein to signal for termination and is controlled by the specific sequences of RNA. When the termination process begins, the transcribed mRNA forms a stable secondary structure hairpin loop, also known as a Stem-loop. This RNA hairpin is followed by multiple uracil nucleotides. The bonds between uracil and adenine are very weak. A protein bound to RNA polymerase (nusA) binds to the stem-loop structure tightly enough to cause the polymerase to temporarily stall. This pausing of the polymerase coincides with transcription of the poly-uracil sequence. The weak adenine-uracil bonds lower the energy of destabilization for the RNA-DNA duplex, allowing it to unwind and dissociate from the RNA polymerase. Overall, the modified RNA structure is what terminates transcription.
RNA polymerase II holoenzyme is a form of eukaryotic RNA polymerase II that is recruited to the promoters of protein-coding genes in living cells. It consists of RNA polymerase II, a subset of general transcription factors, and regulatory proteins known as SRB proteins.
Post-transcriptional regulation is the control of gene expression at the RNA level. It occurs once the RNA polymerase has been attached to the gene's promoter and is synthesizing the nucleotide sequence. Therefore, as the name indicates, it occurs between the transcription phase and the translation phase of gene expression. These controls are critical for the regulation of many genes across human tissues. It also plays a big role in cell physiology, being implicated in pathologies such as cancer and neurodegenerative diseases.
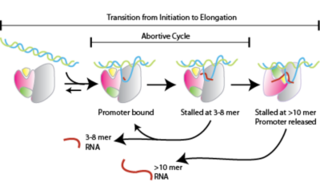
Abortive initiation, also known as abortive transcription, is an early process of genetic transcription in which RNA polymerase binds to a DNA promoter and enters into cycles of synthesis of short mRNA transcripts which are released before the transcription complex leaves the promoter. This process occurs in both eukaryotes and prokaryotes. Abortive initiation is typically studied in the T3 and T7 RNA polymerases in bacteriophages and in E. coli.
Ranjan Sen is an Indian microbiologist, biophysicist and a senior scientist as well as the head of the Laboratory of Transcription at the Centre for DNA Fingerprinting and Diagnostics. Known for his studies in the field of prokaryotic transcription, Sen is an elected fellow of the Indian National Science Academy and the National Academy of Sciences, India. The Department of Biotechnology of the Government of India awarded him the National Bioscience Award for Career Development, one of the highest Indian science awards, for his contributions to biosciences in 2007.
Transcription-translation coupling is a mechanism of gene expression regulation in which synthesis of an mRNA (transcription) is affected by its concurrent decoding (translation). In prokaryotes, mRNAs are translated while they are transcribed. This allows communication between RNA polymerase, the multisubunit enzyme that catalyzes transcription, and the ribosome, which catalyzes translation. Coupling involves both direct physical interactions between RNA polymerase and the ribosome, as well as ribosome-induced changes to the structure and accessibility of the intervening mRNA that affect transcription.
References
- 1 2 3 Lodish H, Berk A, Zipursky SL, et al. (2000). Molecular Cell Biology 4th edition. New York: W. H. Freeman.
- ↑ Boudvillain M, Figueroa-Bossi N, Bossi L (April 2013). "Terminator still moving forward: expanding roles for Rho factor". Current Opinion in Microbiology. 16 (2): 118–24. doi:10.1016/j.mib.2012.12.003. PMID 23347833.
- ↑ Richardson JP (July 2003). "Loading Rho to terminate transcription". Cell. 114 (2): 157–9. doi: 10.1016/s0092-8674(03)00554-3 . PMID 12887917.
- 1 2 Brennan CA, Dombroski AJ, Platt T (March 1987). "Transcription termination factor rho is an RNA-DNA helicase". Cell. 48 (6): 945–52. doi:10.1016/0092-8674(87)90703-3. PMID 3030561. S2CID 42011370.
- ↑ Roberts JW (April 2019). "Mechanisms of Bacterial Transcription Termination". Journal of Molecular Biology. 431 (20): 4030–4039. doi: 10.1016/j.jmb.2019.04.003 . PMID 30978344.
- ↑ Kriner MA, Sevostyanova A, Groisman EA (August 2016). "Learning from the Leaders: Gene Regulation by the Transcription Termination Factor Rho". Trends in Biochemical Sciences. 41 (8): 690–699. doi:10.1016/j.tibs.2016.05.012. PMC 4967001 . PMID 27325240.
- ↑ Qayyum MZ, Dey D, Sen R (April 2016). "Transcription Elongation Factor NusA Is a General Antagonist of Rho-dependent Termination in Escherichia coli". The Journal of Biological Chemistry. 291 (15): 8090–108. doi: 10.1074/jbc.M115.701268 . PMC 4825012 . PMID 26872975.
- ↑ Malik M, Li L, Zhao X, Kerns RJ, Berger JM, Drlica K (December 2014). "Lethal synergy involving bicyclomycin: an approach for reviving old antibiotics". The Journal of Antimicrobial Chemotherapy. 69 (12): 3227–35. doi:10.1093/jac/dku285. PMC 4228776 . PMID 25085655.
- ↑ Nielsen S, Yuzenkova Y, Zenkin N (June 2013). "Mechanism of eukaryotic RNA polymerase III transcription termination". Science. 340 (6140): 1577–80. Bibcode:2013Sci...340.1577N. doi:10.1126/science.1237934. PMC 3760304 . PMID 23812715.