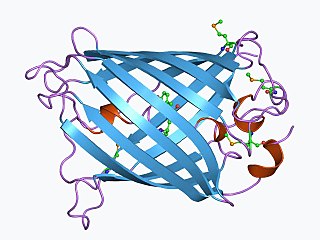
The green fluorescent protein (GFP) is a protein that exhibits green fluorescence when exposed to light in the blue to ultraviolet range. The label GFP traditionally refers to the protein first isolated from the jellyfish Aequorea victoria and is sometimes called avGFP. However, GFPs have been found in other organisms including corals, sea anemones, zoanithids, copepods and lancelets.

Bioluminescence is the production and emission of light by living organisms. It is a form of chemiluminescence. Bioluminescence occurs widely in marine vertebrates and invertebrates, as well as in some fungi, microorganisms including some bioluminescent bacteria, and terrestrial arthropods such as fireflies. In some animals, the light is bacteriogenic, produced by symbiotic bacteria such as those from the genus Vibrio; in others, it is autogenic, produced by the animals themselves.

Chemiluminescence is the emission of light (luminescence) as the result of a chemical reaction, i.e. a chemical reaction results in a flash or glow of light. A standard example of chemiluminescence in the laboratory setting is the luminol test. Here, blood is indicated by luminescence upon contact with iron in hemoglobin. When chemiluminescence takes place in living organisms, the phenomenon is called bioluminescence. A light stick emits light by chemiluminescence.

Luciferase is a generic term for the class of oxidative enzymes that produce bioluminescence, and is usually distinguished from a photoprotein. The name was first used by Raphaël Dubois who invented the words luciferin and luciferase, for the substrate and enzyme, respectively. Both words are derived from the Latin word lucifer, meaning "lightbearer", which in turn is derived from the Latin words for "light" (lux) and "to bring or carry" (ferre).
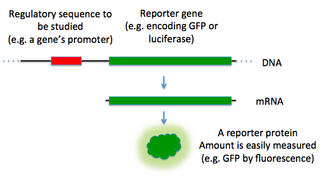
In molecular biology, a reporter gene is a gene that researchers attach to a regulatory sequence of another gene of interest in bacteria, cell culture, animals or plants. Such genes are called reporters because the characteristics they confer on organisms expressing them are easily identified and measured, or because they are selectable markers. Reporter genes are often used as an indication of whether a certain gene has been taken up by or expressed in the cell or organism population.

Aliivibrio fischeri is a Gram-negative, rod-shaped bacterium found globally in marine environments. This species has bioluminescent properties, and is found predominantly in symbiosis with various marine animals, such as the Hawaiian bobtail squid. It is heterotrophic, oxidase-positive, and motile by means of a single polar flagella. Free-living A. fischeri cells survive on decaying organic matter. The bacterium is a key research organism for examination of microbial bioluminescence, quorum sensing, and bacterial-animal symbiosis. It is named after Bernhard Fischer, a German microbiologist.

The pGLO plasmid is an engineered plasmid used in biotechnology as a vector for creating genetically modified organisms. The plasmid contains several reporter genes, most notably the green fluorescent protein (GFP) and the ampicillin resistance gene. GFP was isolated from the jelly fish Aequorea victoria. Because it shares a bidirectional promoter with a gene for metabolizing arabinose, the GFP gene is expressed in the presence of arabinose, which makes the transgenic organism express its fluorescence under UV light. GFP can be induced in bacteria containing the pGLO plasmid by growing them on +arabinose plates. pGLO is made by Bio-Rad Laboratories.

Two-hybrid screening is a molecular biology technique used to discover protein–protein interactions (PPIs) and protein–DNA interactions by testing for physical interactions between two proteins or a single protein and a DNA molecule, respectively.

Aequorin is a calcium-activated photoprotein isolated from the hydrozoan Aequorea victoria. Its bioluminescence was studied decades before the protein was isolated from the animal by Osamu Shimomura in 1962. In the animal, the protein occurs together with the green fluorescent protein to produce green light by resonant energy transfer, while aequorin by itself generates blue light.

Bimolecular fluorescence complementation is a technology typically used to validate protein interactions. It is based on the association of fluorescent protein fragments that are attached to components of the same macromolecular complex. Proteins that are postulated to interact are fused to unfolded complementary fragments of a fluorescent reporter protein and expressed in live cells. Interaction of these proteins will bring the fluorescent fragments within proximity, allowing the reporter protein to reform in its native three-dimensional structure and emit its fluorescent signal. This fluorescent signal can be detected and located within the cell using an inverted fluorescence microscope that allows imaging of fluorescence in cells. In addition, the intensity of the fluorescence emitted is proportional to the strength of the interaction, with stronger levels of fluorescence indicating close or direct interactions and lower fluorescence levels suggesting interaction within a complex. Therefore, through the visualisation and analysis of the intensity and distribution of fluorescence in these cells, one can identify both the location and interaction partners of proteins of interest.
Within the field of molecular biology, a protein-fragment complementation assay, or PCA, is a method for the identification and quantification of protein–protein interactions. In the PCA, the proteins of interest are each covalently linked to fragments of a third protein. Interaction between the bait and the prey proteins brings the fragments of the reporter protein in close proximity to allow them to form a functional reporter protein whose activity can be measured. This principle can be applied to many different reporter proteins and is also the basis for the yeast two-hybrid system, an archetypical PCA assay.

The GUS reporter system is a reporter gene system, particularly useful in plant molecular biology and microbiology. Several kinds of GUS reporter gene assay are available, depending on the substrate used. The term GUS staining refers to the most common of these, a histochemical technique.
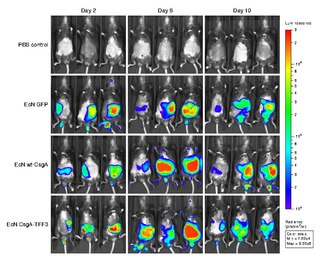
Bioluminescence imaging (BLI) is a technology developed over the past decades (1990's and onward). that allows for the noninvasive study of ongoing biological processes Recently, bioluminescence tomography (BLT) has become possible and several systems have become commercially available. In 2011, PerkinElmer acquired one of the most popular lines of optical imaging systems with bioluminescence from Caliper Life Sciences.

The GAL4-UAS system is a biochemical method used to study gene expression and function in organisms such as the fruit fly. It is based on the finding by Hitoshi Kakidani and Mark Ptashne, and Nicholas Webster and Pierre Chambon in 1988 that Gal4 binding to UAS sequences activates gene expression. The method was introduced into flies by Andrea Brand and Norbert Perrimon in 1993 and is considered a powerful technique for studying the expression of genes. The system has two parts: the Gal4 gene, encoding the yeast transcription activator protein Gal4, and the UAS, an enhancer to which GAL4 specifically binds to activate gene transcription.
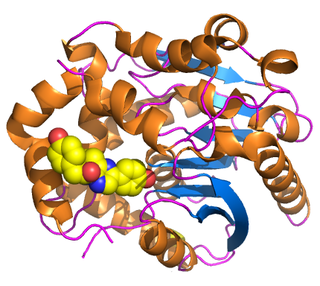
Renilla-luciferin 2-monooxygenase, Renilla luciferase, or RLuc, is a bioluminescent enzyme found in Renilla reniformis, belonging to a group of coelenterazine luciferases. Of this group of enzymes, the luciferase from Renilla reniformis has been the most extensively studied, and due to its bioluminescence requiring only molecular oxygen, has a wide range of applications, with uses as a reporter gene probe in cell culture, in vivo imaging, and various other areas of biological research. Recently, chimeras of RLuc have been developed and demonstrated to be the brightest luminescent proteins to date, and have proved effective in both noninvasive single-cell and whole body imaging.
Autoinducers are signaling molecules that are produced in response to changes in cell-population density. As the density of quorum sensing bacterial cells increases so does the concentration of the autoinducer. Detection of signal molecules by bacteria acts as stimulation which leads to altered gene expression once the minimal threshold is reached. Quorum sensing is a phenomenon that allows both Gram-negative and Gram-positive bacteria to sense one another and to regulate a wide variety of physiological activities. Such activities include symbiosis, virulence, motility, antibiotic production, and biofilm formation. Autoinducers come in a number of different forms depending on the species, but the effect that they have is similar in many cases. Autoinducers allow bacteria to communicate both within and between different species. This communication alters gene expression and allows bacteria to mount coordinated responses to their environments, in a manner that is comparable to behavior and signaling in higher organisms. Not surprisingly, it has been suggested that quorum sensing may have been an important evolutionary milestone that ultimately gave rise to multicellular life forms.
Douglas C. Prasher is an American molecular biologist. He is known for his work to clone and sequence the genes for the photoprotein aequorin and green fluorescent protein (GFP) and for his proposal to use GFP as a tracer molecule. He communicated his pioneering work to Martin Chalfie and Roger Y. Tsien, but by 1991 he was unable to obtain further research funding, and left academia. Eventually, he had to abandon science. Chalfie and Tsien were awarded the 2008 Nobel Prize in Chemistry for work that they publicly acknowledged was substantially based on Prasher's work; through their efforts and those of others, he returned to scientific research in June 2010.
Photoproteins are a type of enzyme produced by bioluminescent organisms. They add to the function of the luciferins whose usual light-producing reaction is catalyzed by the enzyme luciferase.

John Woodland "Woody" Hastings, was a leader in the field of photobiology, especially bioluminescence, and was one of the founders of the field of circadian biology. He was the Paul C. Mangelsdorf Professor of Natural Sciences and Professor of Molecular and Cellular Biology at Harvard University. He published over 400 papers and co-edited three books.

Bioluminescent bacteria are light-producing bacteria that are predominantly present in sea water, marine sediments, the surface of decomposing fish and in the gut of marine animals. While not as common, bacterial bioluminescence is also found in terrestrial and freshwater bacteria. These bacteria may be free living or in symbiosis with animals such as the Hawaiian Bobtail squid or terrestrial nematodes. The host organisms provide these bacteria a safe home and sufficient nutrition. In exchange, the hosts use the light produced by the bacteria for camouflage, prey and/or mate attraction. Bioluminescent bacteria have evolved symbiotic relationships with other organisms in which both participants benefit close to equally. Another possible reason bacteria use luminescence reaction is for quorum sensing, an ability to regulate gene expression in response to bacterial cell density.

















