An endosymbiont or endobiont is any organism that lives within the body or cells of another organism most often, though not always, in a mutualistic relationship. (The term endosymbiosis is from the Greek: ἔνδον endon "within", σύν syn "together" and βίωσις biosis "living".) Examples are nitrogen-fixing bacteria, which live in the root nodules of legumes, single-cell algae inside reef-building corals and bacterial endosymbionts that provide essential nutrients to insects.

A mycorrhiza is a symbiotic association between a fungus and a plant. The term mycorrhiza refers to the role of the fungus in the plant's rhizosphere, its root system. Mycorrhizae play important roles in plant nutrition, soil biology, and soil chemistry.
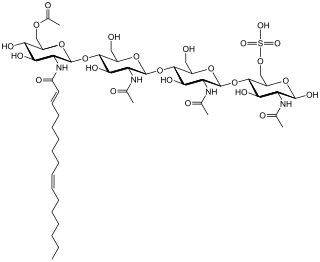
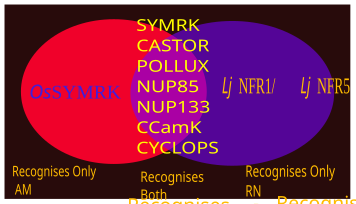
Nod factors, are signaling molecules produced by soil bacteria known as rhizobia in response to flavonoid exudation from plants under nitrogen limited conditions. Nod factors initiate the establishment of a symbiotic relationship between legumes and rhizobia by inducing nodulation. Nod factors produce the differentiation of plant tissue in root hairs into nodules where the bacteria reside and are able to fix nitrogen from the atmosphere for the plant in exchange for photosynthates and the appropriate environment for nitrogen fixation. One of the most important features provided by the plant in this symbiosis is the production of leghemoglobin, which maintains the oxygen concentration low and prevents the inhibition of nitrogenase activity.

An arbuscular mycorrhiza (AM) is a type of mycorrhiza in which the symbiont fungus penetrates the cortical cells of the roots of a vascular plant forming arbuscules. Arbuscular mycorrhiza is a type of endomycorrhiza along with ericoid mycorrhiza and orchid mycorrhiza.

Root hair, or absorbent hairs, are outgrowths of epidermal cells, specialized cells at the tip of a plant root. They are lateral extensions of a single cell and are only rarely branched. They are found in the region of maturation, of the root. Root hair cells improve plant water absorption by increasing root surface area to volume ratio which allows the root hair cell to take in more water. The large vacuole inside root hair cells makes this intake much more efficient. Root hairs are also important for nutrient uptake as they are main interface between plants and mycorrhizal fungi.
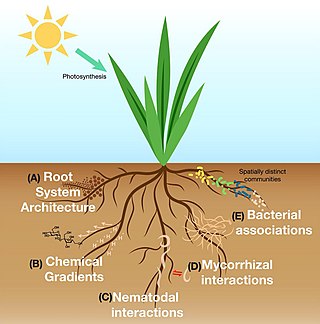
The rhizosphere is the narrow region of soil or substrate that is directly influenced by root secretions and associated soil microorganisms known as the root microbiome. Soil pores in the rhizosphere can contain many bacteria and other microorganisms that feed on sloughed-off plant cells, termed rhizodeposition, and the proteins and sugars released by roots, termed root exudates. This symbiosis leads to more complex interactions, influencing plant growth and competition for resources. Much of the nutrient cycling and disease suppression by antibiotics required by plants, occurs immediately adjacent to roots due to root exudates and metabolic products of symbiotic and pathogenic communities of microorganisms. The rhizosphere also provides space to produce allelochemicals to control neighbours and relatives.
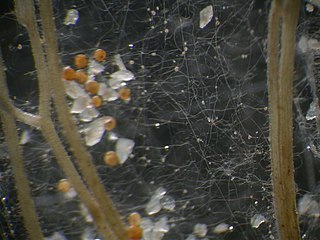
Glomeromycota are one of eight currently recognized divisions within the kingdom Fungi, with approximately 230 described species. Members of the Glomeromycota form arbuscular mycorrhizas (AMs) with the thalli of bryophytes and the roots of vascular land plants. Not all species have been shown to form AMs, and one, Geosiphon pyriformis, is known not to do so. Instead, it forms an endocytobiotic association with Nostoc cyanobacteria. The majority of evidence shows that the Glomeromycota are dependent on land plants for carbon and energy, but there is recent circumstantial evidence that some species may be able to lead an independent existence. The arbuscular mycorrhizal species are terrestrial and widely distributed in soils worldwide where they form symbioses with the roots of the majority of plant species (>80%). They can also be found in wetlands, including salt-marshes, and associated with epiphytic plants.

Glomus is a genus of arbuscular mycorrhizal (AM) fungi, and all species form symbiotic relationships (mycorrhizae) with plant roots. Glomus is the largest genus of AM fungi, with ca. 85 species described, but is currently defined as non-monophyletic.
Microbial inoculants also known as soil inoculants or bioinoculants are agricultural amendments that use beneficial rhizosphericic or endophytic microbes to promote plant health. Many of the microbes involved form symbiotic relationships with the target crops where both parties benefit (mutualism). While microbial inoculants are applied to improve plant nutrition, they can also be used to promote plant growth by stimulating plant hormone production. Although bacterial and fungal inoculants are common, inoculation with archaea to promote plant growth is being increasingly studied.
Nitrogen nutrition in the arbuscular mycorrhizal system refers to...

A mycorrhizal network is an underground network found in forests and other plant communities, created by the hyphae of mycorrhizal fungi joining with plant roots. This network connects individual plants together. Mycorrhizal relationships are most commonly mutualistic, with both partners benefiting, but can be commensal or parasitic, and a single partnership may change between any of the three types of symbiosis at different times.

An ectomycorrhiza is a form of symbiotic relationship that occurs between a fungal symbiont, or mycobiont, and the roots of various plant species. The mycobiont is often from the phyla Basidiomycota and Ascomycota, and more rarely from the Zygomycota. Ectomycorrhizas form on the roots of around 2% of plant species, usually woody plants, including species from the birch, dipterocarp, myrtle, beech, willow, pine and rose families. Research on ectomycorrhizas is increasingly important in areas such as ecosystem management and restoration, forestry and agriculture.

Rhizophagus irregularis is an arbuscular mycorrhizal fungus used as a soil inoculant in agriculture and horticulture. Rhizophagus irregularis is also commonly used in scientific studies of the effects of arbuscular mycorrhizal fungi on plant and soil improvement. Until 2001, the species was known and widely marketed as Glomus intraradices, but molecular analysis of ribosomal DNA led to the reclassification of all arbuscular fungi from Zygomycota phylum to the Glomeromycota phylum.
A phytobiome consists of a plant (phyto) situated in its specific ecological area (biome), including its environment and the associated communities of organisms which inhabit it. These organisms include all macro- and micro-organisms living in, on, or around the plant including bacteria, archaea, fungi, protists, insects, animals, and other plants. The environment includes the soil, air, and climate. Examples of ecological areas are fields, rangelands, forests. Knowledge of the interactions within a phytobiome can be used to create tools for agriculture, crop management, increased health, preservation, productivity, and sustainability of cropping and forest systems.
Martin Parniske is a German biologist with a specialisation in genetics, microbiology and biochemistry. He is university professor and head of the Institute of Genetics at the Faculty of Biology of the Ludwig Maximilian University of Munich. Parniske's scientific focus is on the molecular interaction between plants and symbiotic and pathogenic organisms including bacteria, fungi, oomycetes and insects.
Orchid mycorrhizae are endomycorrhizal fungi which develop symbiotic relationships with the roots and seeds of plants of the family Orchidaceae. Nearly all orchids are myco-heterotrophic at some point in their life cycle. Orchid mycorrhizae are critically important during orchid germination, as an orchid seed has virtually no energy reserve and obtains its carbon from the fungal symbiont.

Strigolactones are a group of chemical compounds produced by roots of plants. Due to their mechanism of action, these molecules have been classified as plant hormones or phytohormones. So far, strigolactones have been identified to be responsible for three different physiological processes: First, they promote the germination of parasitic organisms that grow in the host plant's roots, such as Strigalutea and other plants of the genus Striga. Second, strigolactones are fundamental for the recognition of the plant by symbiotic fungi, especially arbuscular mycorrhizal fungi, because they establish a mutualistic association with these plants, and provide phosphate and other soil nutrients. Third, strigolactones have been identified as branching inhibition hormones in plants; when present, these compounds prevent excess bud growing in stem terminals, stopping the branching mechanism in plants.

Mycorrhiza helper bacteria (MHB) are a group of organisms that form symbiotic associations with both ectomycorrhiza and arbuscular mycorrhiza. MHBs are diverse and belong to a wide variety of bacterial phyla including both Gram-negative and Gram-positive bacteria. Some of the most common MHBs observed in studies belong to the phylas Pseudomonas and Streptomyces. MHBs have been seen to have extremely specific interactions with their fungal hosts at times, but this specificity is lost with plants. MHBs enhance mycorrhizal function, growth, nutrient uptake to the fungus and plant, improve soil conductance, aid against certain pathogens, and help promote defense mechanisms. These bacteria are naturally present in the soil, and form these complex interactions with fungi as plant root development starts to take shape. The mechanisms through which these interactions take shape are not well-understood and needs further study.
Mycorrhizal amelioration of heavy metals or pollutants is a process by which mycorrhizal fungi in a mutualistic relationship with plants can sequester toxic compounds from the environment, as a form of bioremediation.
Dr. Mohamed Hijri is a biologist who studies arbuscular mycorrhizal fungi (AMF). He is a professor of biology and research at the Institut de recherche en biologie végétale at the University of Montreal.