
The globins are a superfamily of heme-containing globular proteins, involved in binding and/or transporting oxygen. These proteins all incorporate the globin fold, a series of eight alpha helical segments. Two prominent members include myoglobin and hemoglobin. Both of these proteins reversibly bind oxygen via a heme prosthetic group. They are widely distributed in many organisms.

A transmembrane protein is a type of integral membrane protein that spans the entirety of the cell membrane. Many transmembrane proteins function as gateways to permit the transport of specific substances across the membrane. They frequently undergo significant conformational changes to move a substance through the membrane. They are usually highly hydrophobic and aggregate and precipitate in water. They require detergents or nonpolar solvents for extraction, although some of them (beta-barrels) can be also extracted using denaturing agents.
DNA primase is an enzyme involved in the replication of DNA and is a type of RNA polymerase. Primase catalyzes the synthesis of a short RNA segment called a primer complementary to a ssDNA template. After this elongation, the RNA piece is removed by a 5' to 3' exonuclease and refilled with DNA.
DnaG is a bacterial DNA primase and is encoded by the dnaG gene. The enzyme DnaG, and any other DNA primase, synthesizes short strands of RNA known as oligonucleotides during DNA replication. These oligonucleotides are known as primers because they act as a starting point for DNA synthesis. DnaG catalyzes the synthesis of oligonucleotides that are 10 to 60 nucleotides long, however most of the oligonucleotides synthesized are 11 nucleotides. These RNA oligonucleotides serve as primers, or starting points, for DNA synthesis by bacterial DNA polymerase III. DnaG is important in bacterial DNA replication because DNA polymerase cannot initiate the synthesis of a DNA strand, but can only add nucleotides to a preexisting strand. DnaG synthesizes a single RNA primer at the origin of replication. This primer serves to prime leading strand DNA synthesis. For the other parental strand, the lagging strand, DnaG synthesizes an RNA primer every few kilobases (kb). These primers serve as substrates for the synthesis of Okazaki fragments.

The Rossmann fold is a tertiary fold found in proteins that bind nucleotides, such as enzyme cofactors FAD, NAD+, and NADP+. This fold is composed of alternating beta strands and alpha helical segments where the beta strands are hydrogen bonded to each other forming an extended beta sheet and the alpha helices surround both faces of the sheet to produce a three-layered sandwich. The classical Rossmann fold contains six beta strands whereas Rossmann-like folds, sometimes referred to as Rossmannoid folds, contain only five strands. The initial beta-alpha-beta (bab) fold is the most conserved segment of the Rossmann fold. The motif is named after Michael Rossmann who first noticed this structural motif in the enzyme lactate dehydrogenase in 1970 and who later observed that this was a frequently occurring motif in nucleotide binding proteins.

Helix-turn-helix is a DNA-binding domain (DBD). The helix-turn-helix (HTH) is a major structural motif capable of binding DNA. Each monomer incorporates two α helices, joined by a short strand of amino acids, that bind to the major groove of DNA. The HTH motif occurs in many proteins that regulate gene expression. It should not be confused with the helix–loop–helix motif.

Glutathione S-transferases (GSTs), previously known as ligandins, are a family of eukaryotic and prokaryotic phase II metabolic isozymes best known for their ability to catalyze the conjugation of the reduced form of glutathione (GSH) to xenobiotic substrates for the purpose of detoxification. The GST family consists of three superfamilies: the cytosolic, mitochondrial, and microsomal—also known as MAPEG—proteins. Members of the GST superfamily are extremely diverse in amino acid sequence, and a large fraction of the sequences deposited in public databases are of unknown function. The Enzyme Function Initiative (EFI) is using GSTs as a model superfamily to identify new GST functions.
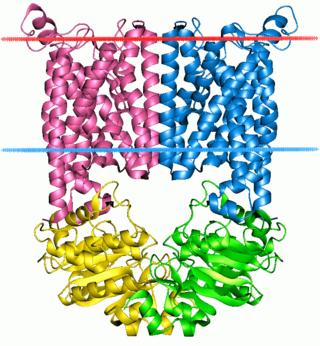
The ABC transporters, ATP synthase (ATP)-binding cassette transporters are a transport system superfamily that is one of the largest and possibly one of the oldest gene families. It is represented in all extant phyla, from prokaryotes to humans. ABC transporters belong to translocases.
A DNA-binding domain (DBD) is an independently folded protein domain that contains at least one structural motif that recognizes double- or single-stranded DNA. A DBD can recognize a specific DNA sequence or have a general affinity to DNA. Some DNA-binding domains may also include nucleic acids in their folded structure.
Phosphodiesterase 1, PDE1, EC 3.1.4.1, systematic name oligonucleotide 5′-nucleotidohydrolase) is a phosphodiesterase enzyme also known as calcium- and calmodulin-dependent phosphodiesterase. It is one of the 11 families of phosphodiesterase (PDE1-PDE11). Phosphodiesterase 1 has three subtypes, PDE1A, PDE1B and PDE1C which divide further into various isoforms. The various isoforms exhibit different affinities for cAMP and cGMP.
Exopolyphosphatase (PPX) is a phosphatase enzyme which catalyzes the hydrolysis of inorganic polyphosphate, a linear molecule composed of up to 1000 or more monomers linked by phospho-anhydride bonds. PPX is a processive exophosphatase, which means that it begins at the ends of the polyphosphate chain and cleaves the phospho-anhydride bonds to release orthophosphate as it moves along the polyphosphate molecule. PPX has several characteristics which distinguish it from other known polyphosphatases, namely that it does not act on ATP, has a strong preference for long chain polyphosphate, and has a very low affinity for polyphosphate molecules with less than 15 phosphate monomers.

Histidine kinases (HK) are multifunctional, and in non-animal kingdoms, typically transmembrane, proteins of the transferase class of enzymes that play a role in signal transduction across the cellular membrane. The vast majority of HKs are homodimers that exhibit autokinase, phosphotransfer, and phosphatase activity. HKs can act as cellular receptors for signaling molecules in a way analogous to tyrosine kinase receptors (RTK). Multifunctional receptor molecules such as HKs and RTKs typically have portions on the outside of the cell that bind to hormone- or growth factor-like molecules, portions that span the cell membrane, and portions within the cell that contain the enzymatic activity. In addition to kinase activity, the intracellular domains typically have regions that bind to a secondary effector molecule or complex of molecules that further propagate signal transduction within the cell. Distinct from other classes of protein kinases, HKs are usually parts of a two-component signal transduction mechanisms in which HK transfers a phosphate group from ATP to a histidine residue within the kinase, and then to an aspartate residue on the receiver domain of a response regulator protein. More recently, the widespread existence of protein histidine phosphorylation distinct from that of two-component histidine kinases has been recognised in human cells. In marked contrast to Ser, Thr and Tyr phosphorylation, the analysis of phosphorylated Histidine using standard biochemical and mass spectrometric approaches is much more challenging, and special procedures and separation techniques are required for their preservation alongside classical Ser, Thr and Tyr phosphorylation on proteins isolated from human cells.
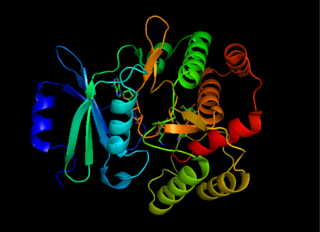
Aminoglycoside-3'-phosphotransferase, also known as aminoglycoside kinase, is an enzyme that primarily catalyzes the addition of phosphate from ATP to the 3'-hydroxyl group of a 4,6-disubstituted aminoglycoside, such as kanamycin. However, APH(3') has also been found to phosphorylate at the 5'-hydroxyl group in 4,5-disubstituted aminoglycosides, which lack a 3'-hydroxyl group, and to diphosphorylate hydroxyl groups in aminoglycosides that have both 3'- and 5'-hydroxyl groups. Primarily positively charged at biological conditions, aminoglycosides bind to the negatively charged backbone of nucleic acids to disrupt protein synthesis, effectively inhibiting bacterial cell growth. APH(3') mediated phosphorylation of aminoglycosides effectively disrupts their mechanism of action, introducing a phosphate group that reduces their binding affinity due to steric hindrances and unfavorable electrostatic interactions. APH(3') is primarily found in certain species of gram-positive bacteria.
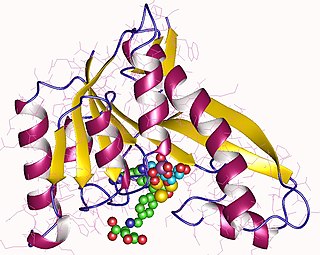
Phosphoribosylglycinamide formyltransferase (EC 2.1.2.2), also known as glycinamide ribonucleotide transformylase (GAR Tfase), is an enzyme with systematic name 10-formyltetrahydrofolate:5'-phosphoribosylglycinamide N-formyltransferase. This enzyme catalyses the following chemical reaction

Bacterial glutathione transferases are part of a superfamily of enzymes that play a crucial role in cellular detoxification. The primary role of GSTs is to catalyze the conjugation of glutathione (GSH) with the electrophilic centers of a wide variety of molecules. The most commonly known substrates of GSTs are xenobiotic synthetic chemicals. There are also classes of GSTs that utilize glutathione as a cofactor rather than a substrate. Often these GSTs are involved in reduction of reactive oxidative species toxic to the bacterium. Conjugation with glutathione receptors renders toxic substances more soluble, and therefore more readily exocytosed from the cell.

S-Adenosylmethionine synthetase, also known as methionine adenosyltransferase (MAT), is an enzyme that creates S-adenosylmethionine by reacting methionine and ATP.
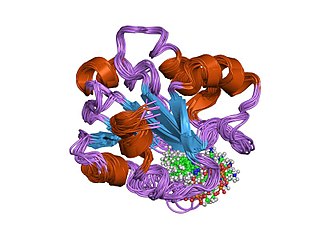
In molecular biology, the vitamin B12-binding domain is a protein domain which binds to cobalamin. It can bind two different forms of the cobalamin cofactor, with cobalt bonded either to a methyl group (methylcobalamin) or to 5'-deoxyadenosine (adenosylcobalamin). Cobalamin-binding domains are mainly found in two families of enzymes present in animals and prokaryotes, which perform distinct kinds of reactions at the cobalt-carbon bond. Enzymes that require methylcobalamin carry out methyl transfer reactions. Enzymes that require adenosylcobalamin catalyse reactions in which the first step is the cleavage of adenosylcobalamin to form cob(II)alamin and the 5'-deoxyadenosyl radical, and thus act as radical generators. In both types of enzymes the B12-binding domain uses a histidine to bind the cobalt atom of cobalamin cofactors. This histidine is embedded in a DXHXXG sequence, the most conserved primary sequence motif of the domain. Proteins containing the cobalamin-binding domain include:

The prokaryotic riboflavin biosynthesis protein is a bifunctional enzyme found in bacteria that catalyzes the phosphorylation of riboflavin into flavin mononucleotide (FMN) and the adenylylation of FMN into flavin adenine dinucleotide (FAD). It consists of a C-terminal riboflavin kinase and an N-terminal FMN-adenylyltransferase. This bacterial protein is functionally similar to the monofunctional riboflavin kinases and FMN-adenylyltransferases of eukaryotic organisms, but only the riboflavin kinases are structurally homologous.
In molecular biology, kanamycin nucleotidyltransferase EC 2.7.7.- (KNTase) is an enzyme which is involved in conferring resistance to aminoglycoside antibiotics. It catalyses the transfer of a nucleoside monophosphate group from a nucleotide to kanamycin. This enzyme is dimeric with each subunit being composed of two domains. The C-terminal domain contains five alpha helices, four of which are organised into an up-and-down alpha helical bundle. Residues found in this domain may contribute to this enzyme's active site.

Class 2 dihydroorotate dehydrogenases is an enzyme with systematic name (S)-dihydroorotate:quinone oxidoreductase. This enzyme catalyses the electron transfer from dihydroorotate to a quinone :















