
Adenylate cyclase is an enzyme with systematic name ATP diphosphate-lyase . It catalyzes the following reaction:
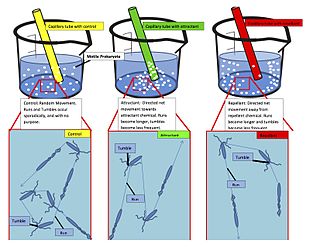
Chemotaxis is the movement of an organism or entity in response to a chemical stimulus. Somatic cells, bacteria, and other single-cell or multicellular organisms direct their movements according to certain chemicals in their environment. This is important for bacteria to find food by swimming toward the highest concentration of food molecules, or to flee from poisons. In multicellular organisms, chemotaxis is critical to early development and development as well as in normal function and health. In addition, it has been recognized that mechanisms that allow chemotaxis in animals can be subverted during cancer metastasis. The aberrant chemotaxis of leukocytes and lymphocytes also contribute to inflammatory diseases such as atherosclerosis, asthma, and arthritis. Sub-cellular components, such as the polarity patch generated by mating yeast, may also display chemotactic behavior.

Integrins are transmembrane receptors that help cell-cell and cell-extracellular matrix (ECM) adhesion. Upon ligand binding, integrins activate signal transduction pathways that mediate cellular signals such as regulation of the cell cycle, organization of the intracellular cytoskeleton, and movement of new receptors to the cell membrane. The presence of integrins allows rapid and flexible responses to events at the cell surface.

A protein kinase is a kinase which selectively modifies other proteins by covalently adding phosphates to them (phosphorylation) as opposed to kinases which modify lipids, carbohydrates, or other molecules. Phosphorylation usually results in a functional change of the target protein (substrate) by changing enzyme activity, cellular location, or association with other proteins. The human genome contains about 500 protein kinase genes and they constitute about 2% of all human genes. There are two main types of protein kinase. The great majority are serine/threonine kinases, which phosphorylate the hydroxyl groups of serines and threonines in their targets. Most of the others are tyrosine kinases, although additional types exist. Protein kinases are also found in bacteria and plants. Up to 30% of all human proteins may be modified by kinase activity, and kinases are known to regulate the majority of cellular pathways, especially those involved in signal transduction.

Signal transduction is the process by which a chemical or physical signal is transmitted through a cell as a series of molecular events. Most commonly, protein phosphorylation is catalyzed by protein kinases, ultimately resulting in a cellular response. Proteins responsible for detecting stimuli are generally termed receptors, although in some cases the term sensor is used. The changes elicited by ligand binding in a receptor give rise to a biochemical cascade, which is a chain of biochemical events known as a signaling pathway.

In biochemistry and pharmacology, receptors are chemical structures, composed of protein, that receive and transduce signals that may be integrated into biological systems. These signals are typically chemical messengers which bind to a receptor and produce physiological responses such as change in the electrical activity of a cell. For example, GABA, an inhibitory neurotransmitter, inhibits electrical activity of neurons by binding to GABAA receptors. There are three main ways the action of the receptor can be classified: relay of signal, amplification, or integration. Relaying sends the signal onward, amplification increases the effect of a single ligand, and integration allows the signal to be incorporated into another biochemical pathway.
Steroid hormone receptors are found in the nucleus, cytosol, and also on the plasma membrane of target cells. They are generally intracellular receptors and initiate signal transduction for steroid hormones which lead to changes in gene expression over a time period of hours to days. The best studied steroid hormone receptors are members of the nuclear receptor subfamily 3 (NR3) that include receptors for estrogen and 3-ketosteroids. In addition to nuclear receptors, several G protein-coupled receptors and ion channels act as cell surface receptors for certain steroid hormones.

Ligand-gated ion channels (LICs, LGIC), also commonly referred to as ionotropic receptors, are a group of transmembrane ion-channel proteins which open to allow ions such as Na+, K+, Ca2+, and/or Cl− to pass through the membrane in response to the binding of a chemical messenger (i.e. a ligand), such as a neurotransmitter.
In biology, cell signaling is the process by which a cell interacts with itself, other cells, and the environment. Cell signaling is a fundamental property of all cellular life in prokaryotes and eukaryotes.
Julius Adler is an American biochemist. He has been an Emeritus Professor of biochemistry and genetics at the University of Wisconsin–Madison since 1997.
The formyl peptide receptors (FPR) belong to a class of G protein-coupled receptors involved in chemotaxis. In humans, there are three formyl peptide receptor isoforms, each encoded by a separate gene that are named FPR1, FPR2, and FPR3. These receptors were originally identified by their ability to bind N-formyl peptides such as N-formylmethionine produced by the degradation of either bacterial or host cells. Hence formyl peptide receptors are involved in mediating immune cell response to infection. These receptors may also act to suppress the immune system under certain conditions. The close phylogenetic relation of signaling in chemotaxis and olfaction was recently proved by detection formyl peptide receptor like proteins as a distinct family of vomeronasal organ chemosensors in mice.

In enzymology, a protein-glutamate O-methyltransferase is an enzyme that catalyzes the chemical reaction

The enzyme protein-glutamate methylesterase (EC 3.1.1.61) catalyzes the reaction

Outer membrane receptors, also known as TonB-dependent receptors, are a family of beta barrel proteins named for their localization in the outer membrane of gram-negative bacteria. TonB complexes sense signals from the outside of bacterial cells and transmit them into the cytoplasm, leading to transcriptional activation of target genes. TonB-dependent receptors in gram-negative bacteria are associated with the uptake and transport of large substrates such as iron siderophore complexes and vitamin B12.

In molecular biology, a two-component regulatory system serves as a basic stimulus-response coupling mechanism to allow organisms to sense and respond to changes in many different environmental conditions. Two-component systems typically consist of a membrane-bound histidine kinase that senses a specific environmental stimulus, and a corresponding response regulator that mediates the cellular response, mostly through differential expression of target genes. Although two-component signaling systems are found in all domains of life, they are most common by far in bacteria, particularly in Gram-negative and cyanobacteria; both histidine kinases and response regulators are among the largest gene families in bacteria. They are much less common in archaea and eukaryotes; although they do appear in yeasts, filamentous fungi, and slime molds, and are common in plants, two-component systems have been described as "conspicuously absent" from animals.

Histidine kinases (HK) are multifunctional, and in non-animal kingdoms, typically transmembrane, proteins of the transferase class of enzymes that play a role in signal transduction across the cellular membrane. The vast majority of HKs are homodimers that exhibit autokinase, phosphotransfer, and phosphatase activity. HKs can act as cellular receptors for signaling molecules in a way analogous to tyrosine kinase receptors (RTK). Multifunctional receptor molecules such as HKs and RTKs typically have portions on the outside of the cell that bind to hormone- or growth factor-like molecules, portions that span the cell membrane, and portions within the cell that contain the enzymatic activity. In addition to kinase activity, the intracellular domains typically have regions that bind to a secondary effector molecule or complex of molecules that further propagate signal transduction within the cell. Distinct from other classes of protein kinases, HKs are usually parts of a two-component signal transduction mechanisms in which HK transfers a phosphate group from ATP to a histidine residue within the kinase, and then to an aspartate residue on the receiver domain of a response regulator protein. More recently, the widespread existence of protein histidine phosphorylation distinct from that of two-component histidine kinases has been recognised in human cells. In marked contrast to Ser, Thr and Tyr phosphorylation, the analysis of phosphorylated Histidine using standard biochemical and mass spectrometric approaches is much more challenging, and special procedures and separation techniques are required for their preservation alongside classical Ser, Thr and Tyr phosphorylation on proteins isolated from human cells.

Bacterial motility is the ability of bacteria to move independently using metabolic energy. Most motility mechanisms that evolved among bacteria also evolved in parallel among the archaea. Most rod-shaped bacteria can move using their own power, which allows colonization of new environments and discovery of new resources for survival. Bacterial movement depends not only on the characteristics of the medium, but also on the use of different appendages to propel. Swarming and swimming movements are both powered by rotating flagella. Whereas swarming is a multicellular 2D movement over a surface and requires the presence of surfactants, swimming is movement of individual cells in liquid environments.

A Per-Arnt-Sim (PAS) domain is a protein domain found in all kingdoms of life. Generally, the PAS domain acts as a molecular sensor, whereby small molecules and other proteins associate via binding of the PAS domain. Due to this sensing capability, the PAS domain has been shown as the key structural motif involved in protein-protein interactions of the circadian clock, and it is also a common motif found in signaling proteins, where it functions as a signaling sensor.

Phototaxis is a kind of taxis, or locomotory movement, that occurs when a whole organism moves towards or away from a stimulus of light. This is advantageous for phototrophic organisms as they can orient themselves most efficiently to receive light for photosynthesis. Phototaxis is called positive if the movement is in the direction of increasing light intensity and negative if the direction is opposite.
In molecular biology, the HAMP domain is an approximately 50-amino acid alpha-helical region that forms a dimeric, four-helical coiled coil. It is found in bacterial sensor and chemotaxis proteins and in eukaryotic histidine kinases. The bacterial proteins are usually integral membrane proteins and part of a two-component signal transduction pathway. One or several copies of the HAMP domain can be found in association with other domains, such as the histidine kinase domain, the bacterial chemotaxis sensory transducer domain, the PAS repeat, the EAL domain, the GGDEF domain, the protein phosphatase 2C-like domain, the guanylate cyclase domain, or the response regulatory domain. In its most common setting, the HAMP domain transmits conformational changes in periplasmic ligand-binding domains to cytoplasmic signalling kinase and methyl-acceptor domains and thus regulates the phosphorylation or methylation activity of homodimeric receptors.
















