Related Research Articles

The endoplasmic reticulum (ER) is a part of a transportation system of the eukaryotic cell, and has many other important functions such as protein folding. It is a type of organelle made up of two subunits – rough endoplasmic reticulum (RER), and smooth endoplasmic reticulum (SER). The endoplasmic reticulum is found in most eukaryotic cells and forms an interconnected network of flattened, membrane-enclosed sacs known as cisternae, and tubular structures in the SER. The membranes of the ER are continuous with the outer nuclear membrane. The endoplasmic reticulum is not found in red blood cells, or spermatozoa.

Protein biosynthesis is a core biological process, occurring inside cells, balancing the loss of cellular proteins through the production of new proteins. Proteins perform a number of critical functions as enzymes, structural proteins or hormones. Protein synthesis is a very similar process for both prokaryotes and eukaryotes but there are some distinct differences.
Protein targeting or protein sorting is the biological mechanism by which proteins are transported to their appropriate destinations within or outside the cell. Proteins can be targeted to the inner space of an organelle, different intracellular membranes, the plasma membrane, or to the exterior of the cell via secretion. Information contained in the protein itself directs this delivery process. Correct sorting is crucial for the cell; errors or dysfunction in sorting have been linked to multiple diseases.
A transmembrane domain (TMD) is a membrane-spanning protein domain. TMDs may consist of one or several alpha-helices or a transmembrane beta barrel. Because the interior of the lipid bilayer is hydrophobic, the amino acid residues in TMDs are often hydrophobic, although proteins such as membrane pumps and ion channels can contain polar residues. TMDs vary greatly in size and hydrophobicity; they may adopt organelle-specific properties.
The signal recognition particle (SRP) is an abundant, cytosolic, universally conserved ribonucleoprotein that recognizes and targets specific proteins to the endoplasmic reticulum in eukaryotes and the plasma membrane in prokaryotes.
The translocon is a complex of proteins associated with the translocation of polypeptides across membranes. In eukaryotes the term translocon most commonly refers to the complex that transports nascent polypeptides with a targeting signal sequence into the interior space of the endoplasmic reticulum (ER) from the cytosol. This translocation process requires the protein to cross a hydrophobic lipid bilayer. The same complex is also used to integrate nascent proteins into the membrane itself. In prokaryotes, a similar protein complex transports polypeptides across the (inner) plasma membrane or integrates membrane proteins. In either case, the protein complex is formed from Sec proteins, with the hetero-trimeric Sec61 being the channel. In prokaryotes, the homologous channel complex is known as SecYEG.
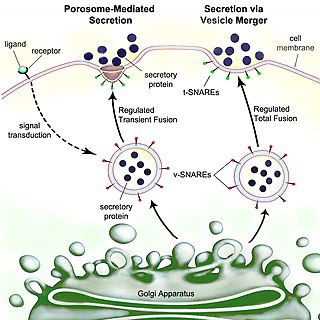
Secretion is the movement of material from one point to another, such as a secreted chemical substance from a cell or gland. In contrast, excretion is the removal of certain substances or waste products from a cell or organism. The classical mechanism of cell secretion is via secretory portals at the plasma membrane called porosomes. Porosomes are permanent cup-shaped lipoprotein structures embedded in the cell membrane, where secretory vesicles transiently dock and fuse to release intra-vesicular contents from the cell.
The N-terminus (also known as the amino-terminus, NH2-terminus, N-terminal end or amine-terminus) is the start of a protein or polypeptide, referring to the free amine group (-NH2) located at the end of a polypeptide. Within a peptide, the amine group is bonded to the carboxylic group of another amino acid, making it a chain. That leaves a free carboxylic group at one end of the peptide, called the C-terminus, and a free amine group on the other end called the N-terminus. By convention, peptide sequences are written N-terminus to C-terminus, left to right (in LTR writing systems). This correlates the translation direction to the text direction, because when a protein is translated from messenger RNA, it is created from the N-terminus to the C-terminus, as amino acids are added to the carboxyl end of the protein.
In cell biology, microsomes are heterogeneous vesicle-like artifacts re-formed from pieces of the endoplasmic reticulum (ER) when eukaryotic cells are broken-up in the laboratory; microsomes are not present in healthy, living cells.
Sec61, termed SecYEG in prokaryotes, is a membrane protein complex found in all domains of life. As the core component of the translocon, it transports proteins to the endoplasmic reticulum in eukaryotes and out of the cell in prokaryotes. It is a doughnut-shaped pore through the membrane with 3 different subunits (heterotrimeric), SecY (α), SecE (γ), and SecG (β). It has a region called the plug that blocks transport into or out of the ER. This plug is displaced when the hydrophobic region of a nascent polypeptide interacts with another region of Sec61 called the seam, allowing translocation of the polypeptide into the ER lumen.
A secretory protein is any protein, whether it be endocrine or exocrine, which is secreted by a cell. Secretory proteins include many hormones, enzymes, toxins, and antimicrobial peptides. Secretory proteins are synthesized in the endoplasmic reticulum.

Signal recognition particle (SRP) receptor, also called the docking protein, is a dimer composed of 2 different subunits that are associated exclusively with the rough ER in mammalian cells. Its main function is to identify the SRP units. SRP is a molecule that helps the ribosome-mRNA-polypeptide complexes to settle down on the membrane of the endoplasmic reticulum.
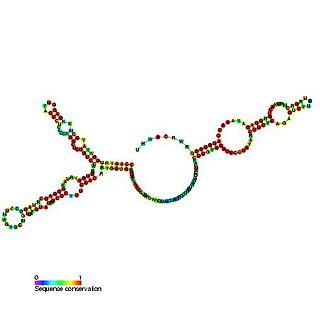

In molecular biology, the small nucleolar RNA SNORA73 belongs to the H/ACA class of small nucleolar RNAs (snoRNAs). SNORA73 has functions involved in mediating the formation of 18S rRNA, regulating chromatin function, and facilitating secretion of proteins by directing specific mRNAs to the signal recognition particle (SRP). SNORA73 has been dubbed a ternary-glue snoRNA (TAG-snoRNA) because of its ability to promote association of mRNAs encoding secreted proteins with the SRP.

The signal recognition particle RNA, is part of the signal recognition particle (SRP) ribonucleoprotein complex. SRP recognizes the signal peptide and binds to the ribosome, halting protein synthesis. SRP-receptor is a protein that is embedded in a membrane, and which contains a transmembrane pore. When the SRP-ribosome complex binds to SRP-receptor, SRP releases the ribosome and drifts away. The ribosome resumes protein synthesis, but now the protein is moving through the SRP-receptor transmembrane pore.

Ribophorins are dome shaped transmembrane glycoproteins which are located in the membrane of the rough endoplasmic reticulum, but are absent in the membrane of the smooth endoplasmic reticulum. There are two types of ribophorines: ribophorin I and II. These act in the protein complex oligosaccharyltransferase (OST) as two different subunits of the named complex. Ribophorin I and II are only present in eukaryote cells.
The SecY protein is the main transmembrane subunit of the bacterial Sec export pathway and of a protein-secreting ATPase complex, also known as a SecYEG translocon. Homologs of the SecYEG complex are found in eukaryotes and in archaea, where the subunit is known as Sec61α.
In cell biology, membrane bound polyribosomes are attached to a cell's endoplasmic reticulum. When certain proteins are synthesized by a ribosome they can become "membrane-bound". The newly produced polypeptide chains are inserted directly into the endoplasmic reticulum by the ribosome and are then transported to their destinations. Bound ribosomes usually produce proteins that are used within the cell membrane or are expelled from the cell via exocytosis.
A target peptide is a short peptide chain that directs the transport of a protein to a specific region in the cell, including the nucleus, mitochondria, endoplasmic reticulum (ER), chloroplast, apoplast, peroxisome and plasma membrane. Some target peptides are cleaved from the protein by signal peptidases after the proteins are transported.
David Domingo Sabatini is an Argentine-American cell biologist and the Frederick L. Ehrman Professor Emeritus of Cell Biology in the Department of Cell Biology at New York University School of Medicine, which he chaired from 1972 to 2011. Sabatini's major research interests have been on the mechanisms responsible for the structural complexity of the eukaryotic cell. Throughout his career, Sabatini has been recognized for his efforts in promoting science in Latin America.

Bacterial secretion systems are protein complexes present on the cell membranes of bacteria for secretion of substances. Specifically, they are the cellular devices used by pathogenic bacteria to secrete their virulence factors to invade the host cells. They can be classified into different types based on their specific structure, composition and activity. Generally, proteins can be secreted through two different processes. One process is a one-step mechanism in which proteins from the cytoplasm of bacteria are transported and delivered directly through the cell membrane into the host cell. Another involves a two-step activity in which the proteins are first transported out of the inner cell membrane, then deposited in the periplasm, and finally through the outer cell membrane into the host cell.
References
- ↑ Kapp, Katja; Schrempf, Sabrina; Lemberg, Marius K.; Dobberstein, Bernhard (2013-01-01). Post-Targeting Functions of Signal Peptides. Landes Bioscience.
- 1 2 Owji, Hajar; Nezafat, Navid; Negahdaripour, Manica; Hajiebrahimi, Ali; Ghasemi, Younes (August 2018). "A comprehensive review of signal peptides: Structure, roles, and applications". European Journal of Cell Biology. 97 (6): 422–441. doi:10.1016/j.ejcb.2018.06.003. PMID 29958716. S2CID 49612506.
- ↑ Blobel G, Dobberstein B (December 1975). "Transfer of proteins across membranes. I. Presence of proteolytically processed and unprocessed nascent immunoglobulin light chains on membrane-bound ribosomes of murine myeloma". The Journal of Cell Biology. 67 (3): 835–51. doi:10.1083/jcb.67.3.835. PMC 2111658 . PMID 811671.
- ↑ Rapoport TA (November 2007). "Protein translocation across the eukaryotic endoplasmic reticulum and bacterial plasma membranes". Nature. 450 (7170): 663–9. Bibcode:2007Natur.450..663R. doi:10.1038/nature06384. PMID 18046402. S2CID 2497138.
- ↑ Käll L, Krogh A, Sonnhammer EL (May 2004). "A combined transmembrane topology and signal peptide prediction method". Journal of Molecular Biology. 338 (5): 1027–36. doi:10.1016/j.jmb.2004.03.016. PMID 15111065.
- ↑ von Heijne G, Gavel Y (July 1988). "Topogenic signals in integral membrane proteins". European Journal of Biochemistry. 174 (4): 671–8. doi: 10.1111/j.1432-1033.1988.tb14150.x . PMID 3134198.
- ↑ Walter P, Ibrahimi I, Blobel G (November 1981). "Translocation of proteins across the endoplasmic reticulum. I. Signal recognition protein (SRP) binds to in-vitro-assembled polysomes synthesizing secretory protein". The Journal of Cell Biology. 91 (2 Pt 1): 545–50. doi:10.1083/jcb.91.2.545. PMC 2111968 . PMID 7309795.
- ↑ Gilmore R, Blobel G, Walter P (November 1982). "Protein translocation across the endoplasmic reticulum. I. Detection in the microsomal membrane of a receptor for the signal recognition particle". The Journal of Cell Biology. 95 (2 Pt 1): 463–9. doi:10.1083/jcb.95.2.463. PMC 2112970 . PMID 6292235.
- ↑ Görlich D, Prehn S, Hartmann E, Kalies KU, Rapoport TA (October 1992). "A mammalian homolog of SEC61p and SECYp is associated with ribosomes and nascent polypeptides during translocation". Cell. 71 (3): 489–503. doi:10.1016/0092-8674(92)90517-G. PMID 1423609. S2CID 19078317.
- ↑ Panzner S, Dreier L, Hartmann E, Kostka S, Rapoport TA (May 1995). "Posttranslational protein transport in yeast reconstituted with a purified complex of Sec proteins and Kar2p". Cell. 81 (4): 561–70. doi: 10.1016/0092-8674(95)90077-2 . PMID 7758110. S2CID 14398668.
- ↑ Kober L, Zehe C, Bode J (April 2013). "Optimized signal peptides for the development of high expressing CHO cell lines". Biotechnology and Bioengineering. 110 (4): 1164–73. doi:10.1002/bit.24776. PMID 23124363. S2CID 449870.
- ↑ von Heijne G (July 1985). "Signal sequences. The limits of variation". Journal of Molecular Biology. 184 (1): 99–105. doi:10.1016/0022-2836(85)90046-4. PMID 4032478.
- ↑ Molino JV, de Carvalho JC, Mayfield SP (2018-02-06). "Comparison of secretory signal peptides for heterologous protein expression in microalgae: Expanding the secretion portfolio for Chlamydomonas reinhardtii". PLOS ONE. 13 (2): e0192433. Bibcode:2018PLoSO..1392433M. doi: 10.1371/journal.pone.0192433 . PMC 5800701 . PMID 29408937.
- ↑ Palazzo AF, Springer M, Shibata Y, Lee CS, Dias AP, Rapoport TA (December 2007). "The signal sequence coding region promotes nuclear export of mRNA". PLOS Biology. 5 (12): e322. doi: 10.1371/journal.pbio.0050322 . PMC 2100149 . PMID 18052610.
- ↑ Cenik C, Chua HN, Zhang H, Tarnawsky SP, Akef A, Derti A, et al. (April 2011). Snyder M (ed.). "Genome analysis reveals interplay between 5'UTR introns and nuclear mRNA export for secretory and mitochondrial genes". PLOS Genetics. 7 (4): e1001366. doi: 10.1371/journal.pgen.1001366 . PMC 3077370 . PMID 21533221.
- ↑ Nickel W, Seedorf M (2008). "Unconventional mechanisms of protein transport to the cell surface of eukaryotic cells". Annual Review of Cell and Developmental Biology. 24: 287–308. doi:10.1146/annurev.cellbio.24.110707.175320. PMID 18590485.
- ↑ Agrawal GK, Jwa NS, Lebrun MH, Job D, Rakwal R (February 2010). "Plant secretome: unlocking secrets of the secreted proteins". Proteomics. 10 (4): 799–827. doi:10.1002/pmic.200900514. PMID 19953550. S2CID 20647387.
- ↑ Witwit, Haydar; Betancourt, Carlos Alberto; Cubitt, Beatrice; Khafaji, Roaa; Kowalski, Heinrich; Jackson, Nathaniel; Ye, Chengjin; Martinez-Sobrido, Luis; de la Torre, Juan C. (2024-08-26). "Cellular N-Myristoyl Transferases Are Required for Mammarenavirus Multiplication". Viruses. 16 (9): 1362. doi: 10.3390/v16091362 . ISSN 1999-4915.