
Nucleotides are organic molecules composed of a nitrogenous base, a pentose sugar and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules within all life-forms on Earth. Nucleotides are obtained in the diet and are also synthesized from common nutrients by the liver.

Lipid-anchored proteins are proteins located on the surface of the cell membrane that are covalently attached to lipids embedded within the cell membrane. These proteins insert and assume a place in the bilayer structure of the membrane alongside the similar fatty acid tails. The lipid-anchored protein can be located on either side of the cell membrane. Thus, the lipid serves to anchor the protein to the cell membrane. They are a type of proteolipids.

Prenylation is the addition of hydrophobic molecules to a protein or a biomolecule. It is usually assumed that prenyl groups (3-methylbut-2-en-1-yl) facilitate attachment to cell membranes, similar to lipid anchors like the GPI anchor, though direct evidence of this has not been observed. Prenyl groups have been shown to be important for protein–protein binding through specialized prenyl-binding domains.
The oxoglutarate dehydrogenase complex (OGDC) or α-ketoglutarate dehydrogenase complex is an enzyme complex, most commonly known for its role in the citric acid cycle.

In biochemistry, flavin adenine dinucleotide (FAD) is a redox-active coenzyme associated with various proteins, which is involved with several enzymatic reactions in metabolism. A flavoprotein is a protein that contains a flavin group, which may be in the form of FAD or flavin mononucleotide (FMN). Many flavoproteins are known: components of the succinate dehydrogenase complex, α-ketoglutarate dehydrogenase, and a component of the pyruvate dehydrogenase complex.
The branched-chain α-ketoacid dehydrogenase complex is a multi-subunit complex of enzymes that is found on the mitochondrial inner membrane. This enzyme complex catalyzes the oxidative decarboxylation of branched, short-chain alpha-ketoacids. BCKDC is a member of the mitochondrial α-ketoacid dehydrogenase complex family comprising pyruvate dehydrogenase and alpha-ketoglutarate dehydrogenase, key enzymes that function in the Krebs cycle.
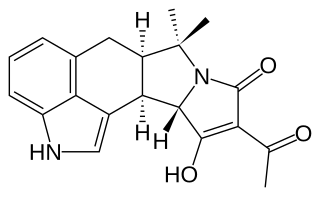
Cyclopiazonic acid (α-CPA), a mycotoxin and a fungal neurotoxin, is made by the molds Aspergillus and Penicillium. It is an indole-tetramic acid that serves as a toxin due to its ability to inhibit calcium-dependent ATPases found in the endoplasmic and sarcoplasmic reticulum. This inhibition disrupts the muscle contraction-relaxation cycle and the calcium gradient that is maintained for proper cellular activity in cells.
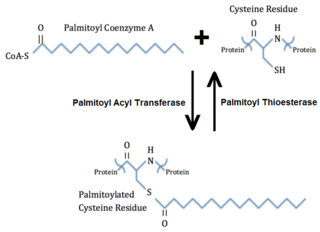
Palmitoylation is the covalent attachment of fatty acids, such as palmitic acid, to cysteine (S-palmitoylation) and less frequently to serine and threonine (O-palmitoylation) residues of proteins, which are typically membrane proteins. The precise function of palmitoylation depends on the particular protein being considered. Palmitoylation enhances the hydrophobicity of proteins and contributes to their membrane association. Palmitoylation also appears to play a significant role in subcellular trafficking of proteins between membrane compartments, as well as in modulating protein–protein interactions. In contrast to prenylation and myristoylation, palmitoylation is usually reversible (because the bond between palmitic acid and protein is often a thioester bond). The reverse reaction in mammalian cells is catalyzed by acyl-protein thioesterases (APTs) in the cytosol and palmitoyl protein thioesterases in lysosomes. Because palmitoylation is a dynamic, post-translational process, it is believed to be employed by the cell to alter the subcellular localization, protein–protein interactions, or binding capacities of a protein.
Pyruvate dehydrogenase is an enzyme that catalyzes the reaction of pyruvate and a lipoamide to give the acetylated dihydrolipoamide and carbon dioxide. The conversion requires the coenzyme thiamine pyrophosphate.
Dihydroorotate dehydrogenase (DHODH) is an enzyme that in humans is encoded by the DHODH gene on chromosome 16. The protein encoded by this gene catalyzes the fourth enzymatic step, the ubiquinone-mediated oxidation of dihydroorotate to orotate, in de novo pyrimidine biosynthesis. This protein is a mitochondrial protein located on the outer surface of the inner mitochondrial membrane (IMM). Inhibitors of this enzyme are used to treat autoimmune diseases such as rheumatoid arthritis.

In enzymology, a kynurenine 3-monooxygenase (EC 1.14.13.9) is an enzyme that catalyzes the chemical reaction

Cystathionine beta-lyase, also commonly referred to as CBL or β-cystathionase, is an enzyme that primarily catalyzes the following α,β-elimination reaction
In enzymology, a geranyltranstransferase is an enzyme that catalyzes the chemical reaction

In enzymology, a riboflavin kinase is an enzyme that catalyzes the chemical reaction

Class 2 dihydroorotate dehydrogenases is an enzyme with systematic name (S)-dihydroorotate:quinone oxidoreductase. This enzyme catalyses the electron transfer from dihydroorotate to a quinone :

Tetrahydrocannabinolic acid (THCA) synthase is an enzyme responsible for catalyzing the formation of THCA from cannabigerolic acid (CBGA). THCA is the direct precursor of tetrahydrocannabinol (THC), the principal psychoactive component of cannabis, which is produced from various strains of Cannabis sativa. Therefore, THCA synthase is considered to be a key enzyme controlling cannabis psychoactivity. Polymorphisms of THCA synthase result in varying levels of THC in Cannabis plants, resulting in "drug-type" and "fiber-type" C. sativa varieties.
Morphinone reductase is an enzyme which catalyzes the NADH-dependent saturation of the carbon-carbon double bond of morphinone and codeinone, yielding hydromorphone and hydrocodone respectively. This saturation reaction is assisted by a FMN cofactor and the enzyme is a member of the α/β-barrel flavoprotein family. The sequence of the enzyme has been obtained from bacteria Pseudomonas putida M10 and has been successfully expressed in yeast and other bacterial species. The enzyme is reported to harbor high sequence and structural similarity to the Old Yellow Enzyme, a large group of flavin-dependent redox biocatalysts of yeast species, and an oestrogen-binding protein of Candida albicans. The enzyme has demonstrated value in biosynthesis of semi-opiate drugs in microorganisms, expanding the chemical diversity of BIA biosynthesis.

UbiX is a flavin prenyltransferase, catalysing the addition of dimethylallyl-monophosphate (DMAP) onto the N5 and C6 positions of FMN culminating in the formation of the prenylated FMN (prFMN) cofactor. The enzyme is involved in the ubiquinone biosynthesis pathway in E.coli from where it gets its name UbiX is associated with the UbiD enzymes as prFMN is utilised by UbiD enzymes in their function as reversible decarboxylases. Unusually for a prenyltransferase UbiX is not metal dependent.

Ferulic acid decarboxylases (Fdc) are decarboxylase enzymes capable of the reversible decarboxylation of aromatic carboxylic acids such as ferulic acid and cinnamic acid. Fdc's are fungal homologues of the E.coli UbiD enzyme which is involved in ubiquinone biosynthesis. This places Fdc within the wider UbiD enzyme family, representing a distinct clade within the family Presence of fdc1 and the associated pad1 genes were shown to be required for the decarboxylation of phenylacrylic acids in Saccharomyces cerevisiae.
Sam Hay is a chemist from New Zealand and a Reader in the Department of Chemistry at The University of Manchester. His research in general is based on computational chemistry and theoretical chemistry, specifically on the areas of In silico Enzymology, quantum mechanics roles in biological processes, kinetic modelling of complex reactions and high pressure spectroscopy.



















