The F-plasmid allows genes to be transferred from one bacterium carrying the factor to another bacterium lacking the factor by conjugation. The F factor was the first plasmid to be discovered. Unlike other plasmids, F factor is constitutive for transfer proteins due to a mutation in the gene finO. The F plasmid belongs to F-like plasmids, a class of conjugative plasmids that control sexual functions of bacteria with a fertility inhibition (Fin) system.
Addiction modules are toxin-antitoxin systems. Each consists of a pair of genes that specify two components: a stable toxin and an unstable antitoxin that interferes with the lethal action of the toxin. Found first in Escherichia coli on low copy number plasmids, addiction modules are responsible for a process called the postsegregational killing effect. When bacteria lose these plasmid(s), the cured cells are selectively killed because the unstable antitoxin is degraded faster than the more stable toxin. The term "addiction" is used because the cell depends on the de novo synthesis of the antitoxin for cell survival. Thus, addiction modules are implicated in maintaining the stability of extrachromosomal elements.
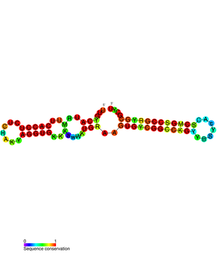
Sib RNA refers to a group of related non-coding RNA. They were originally named QUAD RNA after they were discovered as four repeat elements in Escherichia coli intergenic regions. The family was later renamed Sib when it was discovered that the number of repeats is variable in other species and in other E. coli strains.

The hok/sok system is a postsegregational killing mechanism employed by the R1 plasmid in Escherichia coli. It was the first type I toxin-antitoxin pair to be identified through characterisation of a plasmid-stabilising locus. It is a type I system because the toxin is neutralised by a complementary RNA, rather than a partnered protein.
Segrosomes are protein complexes that ensure accurate segregation (partitioning) of plasmids or chromosomes during bacterial cell division.

Yfr1 is a Cyanobacterial functional RNA that was identified by a comparative genome based screen for RNAs in cyanobacteria. Further analysis has shown that the RNA is well conserved and highly expressed in cyanobacteria. and is required for growth under several stress condition Bioinformatics research combined with follow-up experiments have shown that Yfr1 inhibits the translation of the proteins PMM1119 and PMM1121 by an antisense interaction by base pairing at the ribosomal binding site.
In a screen of the Bacillus subtilis genome for genes encoding ncRNAs, Saito et al. focused on 123 intergenic regions (IGRs) over 500 base pairs in length, the authors analyzed expression from these regions. Seven IGRs termed bsrC, bsrD, bsrE, bsrF, bsrG, bsrH and bsrI expressed RNAs smaller than 380 nt. All the small RNAs except BsrD RNA were expressed in transformed Escherichia coli cells harboring a plasmid with PCR-amplified IGRs of B. subtilis, indicating that their own promoters independently express small RNAs. Under non-stressed condition, depletion of the genes for the small RNAs did not affect growth. Although their functions are unknown, gene expression profiles at several time points showed that most of the genes except for bsrD were expressed during the vegetative phase, but undetectable during the stationary phase. Mapping the 5' ends of the 6 small RNAs revealed that the genes for BsrE, BsrF, BsrG, BsrH, and BsrI RNAs are preceded by a recognition site for RNA polymerase sigma factor σA.

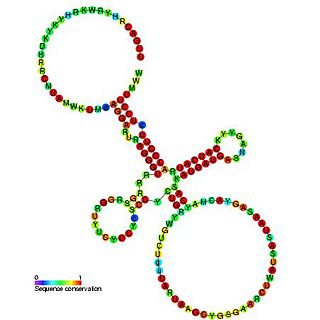
The traJ-II RNA motif is a conserved RNA structure discovered in bacteria by using bioinformatics. traJ-II RNAs appear to be in the 5' untranslated regions of protein-coding genes called traJ, which functions in the process of bacterial conjugation. A previously identified motif known as TraJ 5' UTR is also found upstream of traJ genes functions as the target of FinP antisense RNAs, so it is possible that traJ-II RNAs play a similar role as targets of an antisense RNA. However, some sequence features within the traJ-II RNA motif suggest that the biological RNA might be transcribed from the reverse-complement strand. Thus is it unclear whether traJ-II function as cis-regulatory elements. traJ-II RNAs are found in a variety of Pseudomonadota.

The TisB-IstR toxin-antitoxin system is the first known toxin-antitoxin system which is induced by the SOS response in response to DNA damage.

A toxin-antitoxin system consists of a "toxin" and a corresponding "antitoxin", usually encoded by closely linked genes. The toxin is usually a protein while the antitoxin can be a protein or an RNA. Toxin-antitoxin systems are widely distributed in prokaryotes, and organisms often have them in multiple copies. When these systems are contained on plasmids – transferable genetic elements – they ensure that only the daughter cells that inherit the plasmid survive after cell division. If the plasmid is absent in a daughter cell, the unstable antitoxin is degraded and the stable toxic protein kills the new cell; this is known as 'post-segregational killing' (PSK).

RdlD RNA is a family of small non-coding RNAs which repress the protein LdrD in a type I toxin-antitoxin system. It was discovered in Escherichia coli strain K-12 in a long direct repeat (LDR) named LDR-D. This locus encodes two products: a 35 amino acid peptide toxin (ldrD) and a 60 nucleotide RNA antitoxin. The 374nt toxin mRNA has a half-life of around 30 minutes while rdlD RNA has a half-life of only 2 minutes. This is in keeping with other type I toxin-antitoxin systems.

The SymE-SymR toxin-antitoxin system consists of a small symbiotic endonuclease toxin, SymE, and a non-coding RNA symbiotic RNA antitoxin, SymR, which inhibits SymE translation. SymE-SymR is a type I toxin-antitoxin system, and is under regulation by the antitoxin, SymR. The SymE-SymR complex is believed to play an important role in recycling damaged RNA and DNA. The relationship and corresponding structures of SymE and SymR provide insight into the mechanism of toxicity and overall role in prokaryotic systems.
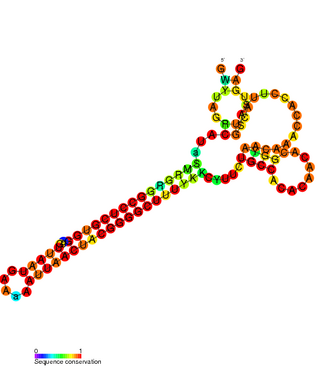
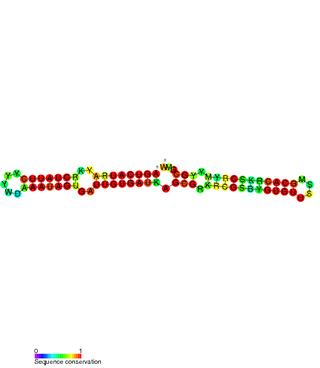
The FlmA-FlmB toxin-antitoxin system consists of FlmB RNA, a family of non-coding RNAs and the protein toxin FlmA. The FlmB RNA transcript is 100 nucleotides in length and is homologous to sok RNA from the hok/sok system and fulfills the identical function as a post-segregational killing (PSK) mechanism.
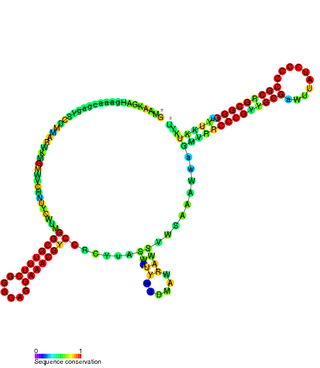
CrfA RNA is a family of non-coding RNAs found in Caulobacter crescentus. CrfA is expressed upon carbon starvation and is thought to activate 27 genes. It was originally identified along with 26 other non-coding RNAs using a tiled Caulobacter microarray protocol specifically aimed at detecting small RNAs.

The TxpA/RatA toxin-antitoxin system was first identified in Bacillus subtilis. It consists of a non-coding 222nt sRNA called RatA and a protein toxin named TxpA.

The par stability determinant is a 400 bp locus of the pAD1 plasmid which encodes a type I toxin-antitoxin system in Enterococcus faecalis. It was the first such plasmid addiction module to be found in gram-positive bacteria.
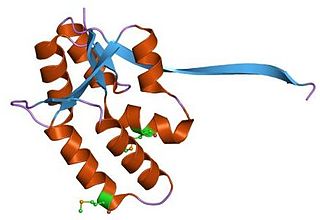
VapBC is the largest family of type II toxin-antitoxin system genetic loci in prokaryotes. VapBC operons consist of two genes: VapC encodes a toxic PilT N-terminus (PIN) domain, and VapB encodes a matching antitoxin. The toxins in this family are thought to perform RNA cleavage, which is inhibited by the co-expression of the antitoxin, in a manner analogous to a poison and antidote.
In molecular biology, trans-activating crispr RNA (tracrRNA) is a small trans-encoded RNA. It was first discovered by Emmanuelle Charpentier in her study of human pathogen Streptococcus pyogenes, a type of bacteria that causes harm to humanity. In bacteria and archaea; CRISPR-Cas constitute an RNA-mediated defense system which protects against viruses and plasmids. This defensive pathway has three steps. First a copy of the invading nucleic acid is integrated into the CRISPR locus. Next, CRISPR RNAs (crRNAs) are transcribed from this CRISPR locus. The crRNAs are then incorporated into effector complexes, where the crRNA guides the complex to the invading nucleic acid and the Cas proteins degrade this nucleic acid. There are several CRISPR system subtypes.
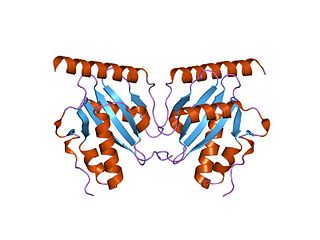
The parDE type II toxin-antitoxin system is one example of the bacterial toxin-antitoxin (TA) systems that encode two proteins, one a potent inhibitor of cell proliferation (toxin) and the other its specific antidote (antitoxin). These systems preferentially guarantee growth of plasmid-carrying daughter cells in a bacterial population by killing newborn bacteria that have not inherited a plasmid copy at cell division.