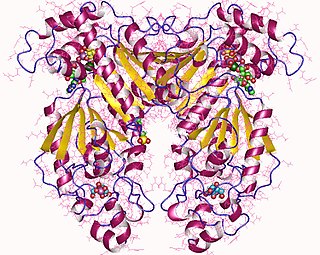
Phosphofructokinase-2 (6-phosphofructo-2-kinase, PFK-2) or fructose bisphosphatase-2 (FBPase-2), is an enzyme indirectly responsible for regulating the rates of glycolysis and gluconeogenesis in cells. It catalyzes formation and degradation of a significant allosteric regulator, fructose-2,6-bisphosphate (Fru-2,6-P2) from substrate fructose-6-phosphate. Fru-2,6-P2 contributes to the rate-determining step of glycolysis as it activates enzyme phosphofructokinase 1 in the glycolysis pathway, and inhibits fructose-1,6-bisphosphatase 1 in gluconeogenesis. Since Fru-2,6-P2 differentially regulates glycolysis and gluconeogenesis, it can act as a key signal to switch between the opposing pathways. Because PFK-2 produces Fru-2,6-P2 in response to hormonal signaling, metabolism can be more sensitively and efficiently controlled to align with the organism's glycolytic needs. This enzyme participates in fructose and mannose metabolism. The enzyme is important in the regulation of hepatic carbohydrate metabolism and is found in greatest quantities in the liver, kidney and heart. In mammals, several genes often encode different isoforms, each of which differs in its tissue distribution and enzymatic activity. The family described here bears a resemblance to the ATP-driven phospho-fructokinases, however, they share little sequence similarity, although a few residues seem key to their interaction with fructose 6-phosphate.
The crotonase family comprises mechanistically diverse proteins that share a conserved trimeric quaternary structure, the core of which consists of 4 turns of a (beta/beta/alpha)n superhelix.
In enzymology, a camphor 1,2-monooxygenase (EC 1.14.15.2) is an enzyme that catalyzes the chemical reaction

In enzymology, a steroid Δ5-isomerase is an enzyme that catalyzes the chemical reaction

In enzymology, a kynurenine-oxoglutarate transaminase is an enzyme that catalyzes the chemical reaction

Iodotyrosine deiodinase, also known as iodotyrosine dehalogenase 1, is a type of deiodinase enzyme that scavenges iodide by removing it from iodinated tyrosine residues in the thyroid gland. These iodinated tyrosines are produced during thyroid hormone biosynthesis. The iodide that is scavenged by iodotyrosine deiodinase is necessary to again synthesize the thyroid hormones. After synthesis, the thyroid hormones circulate through the body to regulate metabolic rate, protein expression, and body temperature. Iodotyrosine deiodinase is thus necessary to keep levels of both iodide and thyroid hormones in balance.

The haloacid dehydrogenase superfamily is a superfamily of enzymes that include phosphatases, phosphonatases, P-type ATPases, beta-phosphoglucomutases, phosphomannomutases, and dehalogenases, and are involved in a variety of cellular processes ranging from amino acid biosynthesis to detoxification.
1,8-Cineole 2-endo-monooxygenase (EC 1.14.14.133, Formerly EC 1.14.13.156, P450cin, CYP176A, CYP176A1) is an enzyme with systematic name 1,8-cineole,NADPH:oxygen oxidoreductase (2-endo-hydroxylating). This enzyme catalyses the following chemical reaction
2,5-diketocamphane 1,2-monooxygenase (EC 1.14.14.108, 2,5-diketocamphane lactonizing enzyme, ketolactonase I, 2,5-diketocamphane 1,2-monooxygenase oxygenating component, 2,5-DKCMO, camphor 1,2-monooxygenase, camphor ketolactonase I) is an enzyme with systematic name (+)-bornane-2,5-dione,NADH:oxygen oxidoreductase (1,2-lactonizing). This enzyme catalyses the following chemical reaction
3-oxo-5,6-dehydrosuberyl-CoA semialdehyde dehydrogenase (EC 1.17.1.7, paaZ (gene)) is an enzyme with systematic name 3-oxo-5,6-dehydrosuberyl-CoA semialdehyde:NADP+ oxidoreductase. This enzyme catalyses the following chemical reaction
Deoxyhypusine synthase (EC 2.5.1.46, spermidine:eIF5A-lysine 4-aminobutyltransferase (propane-1,3-diamine-forming)) is an enzyme with systematic name (eIF5A-precursor)-lysine:spermidine 4-aminobutyltransferase (propane-1,3-diamine-forming). This enzyme catalyses the following chemical reaction

Protein O-GlcNAcase (EC 3.2.1.169, OGA, glycoside hydrolase O-GlcNAcase, O-GlcNAcase, BtGH84, O-GlcNAc hydrolase) is an enzyme with systematic name (protein)-3-O-(N-acetyl-D-glucosaminyl)-L-serine/threonine N-acetylglucosaminyl hydrolase. OGA is encoded by the OGA gene. This enzyme catalyses the removal of the O-GlcNAc post-translational modification in the following chemical reaction:
- [protein]-3-O-(N-acetyl-β-D-glucosaminyl)-L-serine + H2O ⇌ [protein]-L-serine + N-acetyl-D-glucosamine
- [protein]-3-O-(N-acetyl-β-D-glucosaminyl)-L-threonine + H2O ⇌ [protein]-L-threonine + N-acetyl-D-glucosamine
Glutathione hydrolase (EC 3.4.19.13, glutathionase, GGT, gamma-glutamyltranspeptidase) is an enzyme. This enzyme catalyses the following chemical reaction
UDP-2,3-diacylglucosamine diphosphatase (EC 3.6.1.54, UDP-2,3-diacylglucosamine hydrolase, UDP-2,3-diacylglucosamine pyrophosphatase, ybbF (gene), lpxH (gene)) is an enzyme with systematic name UDP-2,3-bis((3R)-3-hydroxymyristoyl)-alpha-D-glucosamine 2,3-bis((3R)-3-hydroxymyristoyl)-beta-D-glucosaminyl 1-phosphate phosphohydrolase. This enzyme catalyses the following chemical reaction
2-hydroxy-dATP diphosphatase is an enzyme that in humans is encoded by the NUDT1 gene. During DNA repair, the enzyme hydrolyses oxidized purines and prevents their addition onto the DNA chain. As such it has important role in aging and cancer development.
Oxepin-CoA hydrolase (EC 3.7.1.16, paaZ (gene)) is an enzyme with systematic name 2-oxepin-2(3H)-ylideneacetyl-CoA hydrolyase. This enzyme catalyses the following chemical reaction
(S)-hydroxynitrile lyase (EC 4.1.2.47, (S)-cyanohydrin producing hydroxynitrile lyase, (S)-oxynitrilase, (S)-HbHNL, (S)-MeHNL, hydroxynitrile lyase, oxynitrilase, HbHNL, MeHNL, (S)-selective hydroxynitrile lyase, (S)-cyanohydrin carbonyl-lyase (cyanide forming), hydroxynitrilase) is an enzyme with systematic name (S)-cyanohydrin lyase (cyanide forming). This enzyme catalyses the interconversion between cyanohydrins and the carbonyl compounds derived from the cyanohydrin with free cyanide, as in the following two chemical reactions:
TDP-4-oxo-6-deoxy-alpha-D-glucose-3,4-oxoisomerase (dTDP-3-dehydro-6-deoxy-alpha-D-galactopyranose-forming) is an enzyme with systematic name dTDP-4-dehydro-6-deoxy-alpha-D-glucopyranose:dTDP-3-dehydro-6-deoxy-alpha-D-galactopyranose isomerase. This enzyme catalyses the following chemical reaction

Acyl-CoA thioesterase 9 is a protein that is encoded by the human ACOT9 gene. It is a member of the acyl-CoA thioesterase superfamily, which is a group of enzymes that hydrolyze Coenzyme A esters. There is no known function, however it has been shown to act as a long-chain thioesterase at low concentrations, and a short-chain thioesterase at high concentrations.

Acyl-CoA thioesterase 13 is a protein that in humans is encoded by the ACOT13 gene. This gene encodes a member of the thioesterase superfamily. In humans, the protein co-localizes with microtubules and is essential for sustained cell proliferation.







