
Bioinformatics is an interdisciplinary field of science that develops methods and software tools for understanding biological data, especially when the data sets are large and complex. Bioinformatics uses biology, chemistry, physics, computer science, computer programming, information engineering, mathematics and statistics to analyze and interpret biological data. The subsequent process of analyzing and interpreting data is referred to as computational biology.
Cell biology is a branch of biology that studies the structure, function, and behavior of cells. All living organisms are made of cells. A cell is the basic unit of life that is responsible for the living and functioning of organisms. Cell biology is the study of the structural and functional units of cells. Cell biology encompasses both prokaryotic and eukaryotic cells and has many subtopics which may include the study of cell metabolism, cell communication, cell cycle, biochemistry, and cell composition. The study of cells is performed using several microscopy techniques, cell culture, and cell fractionation. These have allowed for and are currently being used for discoveries and research pertaining to how cells function, ultimately giving insight into understanding larger organisms. Knowing the components of cells and how cells work is fundamental to all biological sciences while also being essential for research in biomedical fields such as cancer, and other diseases. Research in cell biology is interconnected to other fields such as genetics, molecular genetics, molecular biology, medical microbiology, immunology, and cytochemistry.

In vitro studies are performed with microorganisms, cells, or biological molecules outside their normal biological context. Colloquially called "test-tube experiments", these studies in biology and its subdisciplines are traditionally done in labware such as test tubes, flasks, Petri dishes, and microtiter plates. Studies conducted using components of an organism that have been isolated from their usual biological surroundings permit a more detailed or more convenient analysis than can be done with whole organisms; however, results obtained from in vitro experiments may not fully or accurately predict the effects on a whole organism. In contrast to in vitro experiments, in vivo studies are those conducted in living organisms, including humans, known as clinical trials, and whole plants.
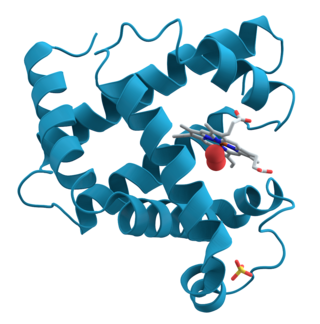
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity.

The proteome is the entire set of proteins that is, or can be, expressed by a genome, cell, tissue, or organism at a certain time. It is the set of expressed proteins in a given type of cell or organism, at a given time, under defined conditions. Proteomics is the study of the proteome.

Proteomics is the large-scale study of proteins. Proteins are vital parts of living organisms, with many functions such as the formation of structural fibers of muscle tissue, enzymatic digestion of food, or synthesis and replication of DNA. In addition, other kinds of proteins include antibodies that protect an organism from infection, and hormones that send important signals throughout the body.
Glycomics is the comprehensive study of glycomes, including genetic, physiologic, pathologic, and other aspects. Glycomics "is the systematic study of all glycan structures of a given cell type or organism" and is a subset of glycobiology. The term glycomics is derived from the chemical prefix for sweetness or a sugar, "glyco-", and was formed to follow the omics naming convention established by genomics and proteomics.
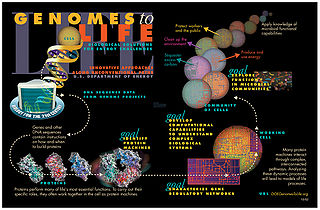
Systems biology is the computational and mathematical analysis and modeling of complex biological systems. It is a biology-based interdisciplinary field of study that focuses on complex interactions within biological systems, using a holistic approach to biological research.
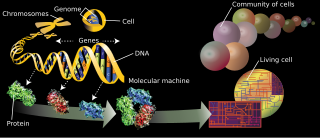
The branches of science known informally as omics are various disciplines in biology whose names end in the suffix -omics, such as genomics, proteomics, metabolomics, metagenomics, phenomics and transcriptomics. Omics aims at the collective characterization and quantification of pools of biological molecules that translate into the structure, function, and dynamics of an organism or organisms.

Cell death is the event of a biological cell ceasing to carry out its functions. This may be the result of the natural process of old cells dying and being replaced by new ones, as in programmed cell death, or may result from factors such as diseases, localized injury, or the death of the organism of which the cells are part. Apoptosis or Type I cell-death, and autophagy or Type II cell-death are both forms of programmed cell death, while necrosis is a non-physiological process that occurs as a result of infection or injury.
Regulome refers to the whole set of regulatory components in a cell. Those components can be regulatory elements, genes, mRNAs, proteins, and metabolites. The description includes the interplay of regulatory effects between these components, and their dependence on variables such as subcellular localization, tissue, developmental stage, and pathological state.

In pharmacology, the term mechanism of action (MOA) refers to the specific biochemical interaction through which a drug substance produces its pharmacological effect. A mechanism of action usually includes mention of the specific molecular targets to which the drug binds, such as an enzyme or receptor. Receptor sites have specific affinities for drugs based on the chemical structure of the drug, as well as the specific action that occurs there.
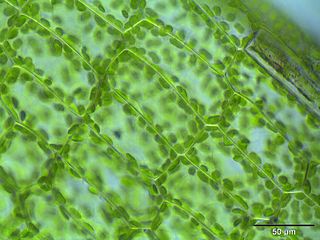

The following outline is provided as an overview of and topical guide to cell biology:
The secretome is the set of proteins expressed by an organism and secreted into the extracellular space. In humans, this subset of the proteome encompasses 13-20% of all proteins, including cytokines, growth factors, extracellular matrix proteins and regulators, and shed receptors. The secretome of a specific tissue can be measured by mass spectrometry and its analysis constitutes a type of proteomics known as secretomics.
Flow cytometry bioinformatics is the application of bioinformatics to flow cytometry data, which involves storing, retrieving, organizing and analyzing flow cytometry data using extensive computational resources and tools. Flow cytometry bioinformatics requires extensive use of and contributes to the development of techniques from computational statistics and machine learning. Flow cytometry and related methods allow the quantification of multiple independent biomarkers on large numbers of single cells. The rapid growth in the multidimensionality and throughput of flow cytometry data, particularly in the 2000s, has led to the creation of a variety of computational analysis methods, data standards, and public databases for the sharing of results.
Toponomics is a discipline in systems biology, molecular cell biology, and histology concerning the study of the toponome of organisms. It is the field of study that purposes to decode the complete toponome in health and disease —which is the next big challenge in human biotechnology after having decoded the human genome.
An imaging cycler microscope (ICM) is a fully automated (epi)fluorescence microscope which overcomes the spectral resolution limit resulting in parameter- and dimension-unlimited fluorescence imaging. The principle and robotic device was described by Walter Schubert in 1997 and has been further developed with his co-workers within the human toponome project. The ICM runs robotically controlled repetitive incubation-imaging-bleaching cycles with dye-conjugated probe libraries recognizing target structures in situ (biomolecules in fixed cells or tissue sections). This results in the transmission of a randomly large number of distinct biological informations by re-using the same fluorescence channel after bleaching for the transmission of another biological information using the same dye which is conjugated to another specific probe, a.s.o. Thereby noise-reduced quasi-multichannel fluorescence images with reproducible physical, geometrical, and biophysical stabilities are generated. The resulting power of combinatorial molecular discrimination (PCMD) per data point is given by 65,536k, where 65,536 is the number of grey value levels (output of a 16-bit CCD camera), and k is the number of co-mapped biomolecules and/or subdomains per biomolecule(s). High PCMD has been shown for k = 100, and in principle can be expanded for much higher numbers of k. In contrast to traditional multichannel–few-parameter fluorescence microscopy (panel a in the figure) high PCMDs in an ICM lead to high functional and spatial resolution (panel b in the figure). Systematic ICM analysis of biological systems reveals the supramolecular segregation law that describes the principle of order of large, hierarchically organized biomolecular networks in situ (toponome). The ICM is the core technology for the systematic mapping of the complete protein network code in tissues (human toponome project). The original ICM method includes any modification of the bleaching step. Corresponding modifications have been reported for antibody retrieval and chemical dye-quenching debated recently. The Toponome Imaging Systems (TIS) and multi-epitope-ligand cartographs (MELC) represent different stages of the ICM technological development. Imaging cycler microscopy received the American ISAC best paper award in 2008 for the three symbol code of organized proteomes.

Cytometry Part A is a peer-reviewed scientific journal covering all aspects of the study of cytometry that was established in 1980. It is the official journal of the International Society for Advancement of Cytometry.
Minimum information standards are sets of guidelines and formats for reporting data derived by specific high-throughput methods. Their purpose is to ensure the data generated by these methods can be easily verified, analysed and interpreted by the wider scientific community. Ultimately, they facilitate the transfer of data from journal articles into databases in a form that enables data to be mined across multiple data sets. Minimal information standards are available for a vast variety of experiment types including microarray (MIAME), RNAseq (MINSEQE), metabolomics (MSI) and proteomics (MIAPE).
Tissue image cytometry or tissue cytometry is a method of digital histopathology and combines classical digital pathology and computational pathology into one integrated approach with solutions for all kinds of diseases, tissue and cell types as well as molecular markers and corresponding staining methods to visualize these markers. Tissue cytometry uses virtual slides as they can be generated by multiple, commercially available slide scanners, as well as dedicated image analysis software – preferentially including machine and deep learning algorithms. Tissue cytometry enables cellular analysis within thick tissues, retaining morphological and contextual information, including spatial information on defined cellular subpopulations. In this process, a tissue sample, either formalin-fixed paraffin-embedded (FFPE) or frozen tissue section, also referred to as “cryocut”, is labelled with either immunohistochemistry(IHC) or immunofluorescent markers, scanned with high-throughput slide scanners and the data gathered from virtual slides is processed and analyzed using software that is able to identify individual cells in tissue context automatically and distinguish between nucleus and cytoplasm for each cell. Additional algorithms can identify cellular membranes, subcellular structures and/or multicellular tissue structures.











