
In machine learning, supervised learning (SL) is a paradigm where a model is trained using input objects and desired output values, which are often human-made labels. The training process builds a function that maps new data to expected output values. An optimal scenario will allow for the algorithm to accurately determine output values for unseen instances. This requires the learning algorithm to generalize from the training data to unseen situations in a "reasonable" way. This statistical quality of an algorithm is measured via a generalization error.

Numerical analysis is the study of algorithms that use numerical approximation for the problems of mathematical analysis. It is the study of numerical methods that attempt to find approximate solutions of problems rather than the exact ones. Numerical analysis finds application in all fields of engineering and the physical sciences, and in the 21st century also the life and social sciences like economics, medicine, business and even the arts. Current growth in computing power has enabled the use of more complex numerical analysis, providing detailed and realistic mathematical models in science and engineering. Examples of numerical analysis include: ordinary differential equations as found in celestial mechanics, numerical linear algebra in data analysis, and stochastic differential equations and Markov chains for simulating living cells in medicine and biology.
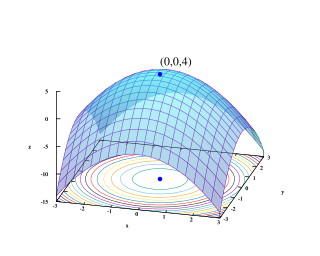
Mathematical optimization or mathematical programming is the selection of a best element, with regard to some criteria, from some set of available alternatives. It is generally divided into two subfields: discrete optimization and continuous optimization. Optimization problems arise in all quantitative disciplines from computer science and engineering to operations research and economics, and the development of solution methods has been of interest in mathematics for centuries.
Minimum Description Length (MDL) is a model selection principle where the shortest description of the data is the best model. MDL methods learn through a data compression perspective and are sometimes described as mathematical applications of Occam's razor. The MDL principle can be extended to other forms of inductive inference and learning, for example to estimation and sequential prediction, without explicitly identifying a single model of the data.
In mathematics and computer algebra, automatic differentiation, also called algorithmic differentiation, computational differentiation, and differentiation arithmetic is a set of techniques to evaluate the partial derivative of a function specified by a computer program. Automatic differentiation is a subtle and central tool to automatize the simultaneous computation of the numerical values of arbitrarily complex functions and their derivatives with no need for the symbolic representation of the derivative, only the function rule or an algorithm thereof is required. Auto-differentiation is thus neither numeric nor symbolic, nor is it a combination of both. It is also preferable to ordinary numerical methods: In contrast to the more traditional numerical methods based on finite differences, auto-differentiation is 'in theory' exact, and in comparison to symbolic algorithms, it is computationally inexpensive.
Computational science, also known as scientific computing, technical computing or scientific computation (SC), is a division of science, and more specifically the Computer Sciences, which uses advanced computing capabilities to understand and solve complex physical problems. While this discussion typically extenuates into Visual Computation, this research field of study will typically include the following research categorizations.
In statistics, a generalized additive model (GAM) is a generalized linear model in which the linear response variable depends linearly on unknown smooth functions of some predictor variables, and interest focuses on inference about these smooth functions.
In statistics, M-estimators are a broad class of extremum estimators for which the objective function is a sample average. Both non-linear least squares and maximum likelihood estimation are special cases of M-estimators. The definition of M-estimators was motivated by robust statistics, which contributed new types of M-estimators. However, M-estimators are not inherently robust, as is clear from the fact that they include maximum likelihood estimators, which are in general not robust. The statistical procedure of evaluating an M-estimator on a data set is called M-estimation. The "M" initial stands for "maximum likelihood-type".
BALL is a C++ class framework and set of algorithms and data structures for molecular modelling and computational structural bioinformatics, a Python interface to this library, and a graphical user interface to BALL, the molecule viewer BALLView.
As applied in the field of computer vision, graph cut optimization can be employed to efficiently solve a wide variety of low-level computer vision problems, such as image smoothing, the stereo correspondence problem, image segmentation, object co-segmentation, and many other computer vision problems that can be formulated in terms of energy minimization.
In control theory a self-tuning system is capable of optimizing its own internal running parameters in order to maximize or minimize the fulfilment of an objective function; typically the maximization of efficiency or error minimization.
Approximate Bayesian computation (ABC) constitutes a class of computational methods rooted in Bayesian statistics that can be used to estimate the posterior distributions of model parameters.

Computational statistics, or statistical computing, is the study which is the intersection of statistics and computer science, and refers to the statistical methods that are enabled by using computational methods. It is the area of computational science specific to the mathematical science of statistics. This area is fast developing. The view that the broader concept of computing must be taught as part of general statistical education is gaining momentum.
In applied mathematics, Hessian automatic differentiation are techniques based on automatic differentiation (AD) that calculate the second derivative of an -dimensional function, known as the Hessian matrix.

Stan is a probabilistic programming language for statistical inference written in C++. The Stan language is used to specify a (Bayesian) statistical model with an imperative program calculating the log probability density function.

Adept is a combined automatic differentiation and array software library for the C++ programming language. The automatic differentiation capability facilitates the development of applications involving mathematical optimization. Adept is notable for having applied the template metaprogramming technique of expression templates to speed-up the differentiation of mathematical statements. Along with the efficient way that it stores the differential information, this makes it significantly faster than most other C++ tools that provide similar functionality, although comparable performance has been reported for Stan and in some cases Sacado. Differentiation may be in forward mode, reverse mode, or the full Jacobian matrix may be computed.






