Related Research Articles
Cladistics is an approach to biological classification in which organisms are categorized in groups ("clades") based on hypotheses of most recent common ancestry. The evidence for hypothesized relationships is typically shared derived characteristics (synapomorphies) that are not present in more distant groups and ancestors. However, from an empirical perspective, common ancestors are inferences based on a cladistic hypothesis of relationships of taxa whose character states can be observed. Theoretically, a last common ancestor and all its descendants constitute a (minimal) clade. Importantly, all descendants stay in their overarching ancestral clade. For example, if the terms worms or fishes were used within a strict cladistic framework, these terms would include humans. Many of these terms are normally used paraphyletically, outside of cladistics, e.g. as a 'grade', which are fruitless to precisely delineate, especially when including extinct species. Radiation results in the generation of new subclades by bifurcation, but in practice sexual hybridization may blur very closely related groupings.
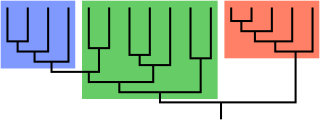
In biological phylogenetics, a clade, also known as a monophyletic group or natural group, is a grouping of organisms that are monophyletic – that is, composed of a common ancestor and all its lineal descendants – on a phylogenetic tree. In the taxonomical literature, sometimes the Latin form cladus is used rather than the English form. Clades are the fundamental unit of cladistics, a modern approach to taxonomy adopted by most biological fields.

In biological cladistics for the classification of organisms, monophyly is the condition of a taxonomic grouping being a clade – that is, a grouping of taxa which meets these criteria:
- the grouping contains its own most recent common ancestor, i.e. excludes non-descendants of that common ancestor
- the grouping contains all the descendants of that common ancestor, without exception
In biology, phylogenetics is the study of the evolutionary history and relationships among or within groups of organisms. These relationships are determined by phylogenetic inference methods that focus on observed heritable traits, such as DNA sequences, protein amino acid sequences, or morphology. The result of such an analysis is a phylogenetic tree—a diagram containing a hypothesis of relationships that reflects the evolutionary history of a group of organisms.

Paraphyly is a taxonomic term describing a grouping that consists of the grouping's last common ancestor and most of its descendants, but excludes one or more subgroups. The grouping is said to be paraphyletic with respect to the excluded subgroups. In contrast, a monophyletic grouping includes a common ancestor and all of its descendants.
In biology, taxonomy is the scientific study of naming, defining (circumscribing) and classifying groups of biological organisms based on shared characteristics. Organisms are grouped into taxa and these groups are given a taxonomic rank; groups of a given rank can be aggregated to form a more inclusive group of higher rank, thus creating a taxonomic hierarchy. The principal ranks in modern use are domain, kingdom, phylum, class, order, family, genus, and species. The Swedish botanist Carl Linnaeus is regarded as the founder of the current system of taxonomy, as he developed a ranked system known as Linnaean taxonomy for categorizing organisms and binomial nomenclature for naming organisms.

The dicotyledons, also known as dicots, are one of the two groups into which all the flowering plants (angiosperms) were formerly divided. The name refers to one of the typical characteristics of the group: namely, that the seed has two embryonic leaves or cotyledons. There are around 200,000 species within this group. The other group of flowering plants were called monocotyledons, typically each having one cotyledon. Historically, these two groups formed the two divisions of the flowering plants.

A phylogenetic tree, phylogeny or evolutionary tree is a graphical representation which shows the evolutionary history between a set of species or taxa during a specific time. In other words, it is a branching diagram or a tree showing the evolutionary relationships among various biological species or other entities based upon similarities and differences in their physical or genetic characteristics. In evolutionary biology, all life on Earth is theoretically part of a single phylogenetic tree, indicating common ancestry. Phylogenetics is the study of phylogenetic trees. The main challenge is to find a phylogenetic tree representing optimal evolutionary ancestry between a set of species or taxa. Computational phylogenetics focuses on the algorithms involved in finding optimal phylogenetic tree in the phylogenetic landscape.
Molecular phylogenetics is the branch of phylogeny that analyzes genetic, hereditary molecular differences, predominantly in DNA sequences, to gain information on an organism's evolutionary relationships. From these analyses, it is possible to determine the processes by which diversity among species has been achieved. The result of a molecular phylogenetic analysis is expressed in a phylogenetic tree. Molecular phylogenetics is one aspect of molecular systematics, a broader term that also includes the use of molecular data in taxonomy and biogeography.

In biology, a taxon is a group of one or more populations of an organism or organisms seen by taxonomists to form a unit. Although neither is required, a taxon is usually known by a particular name and given a particular ranking, especially if and when it is accepted or becomes established. It is very common, however, for taxonomists to remain at odds over what belongs to a taxon and the criteria used for inclusion, especially in the context of rank-based ("Linnaean") nomenclature. If a taxon is given a formal scientific name, its use is then governed by one of the nomenclature codes specifying which scientific name is correct for a particular grouping.

A polyphyletic group is an assemblage that includes organisms with mixed evolutionary origin but does not include their most recent common ancestor. The term is often applied to groups that share similar features known as homoplasies, which are explained as a result of convergent evolution. The arrangement of the members of a polyphyletic group is called a polyphyly. It is contrasted with monophyly and paraphyly.
Evolutionary taxonomy, evolutionary systematics or Darwinian classification is a branch of biological classification that seeks to classify organisms using a combination of phylogenetic relationship, progenitor-descendant relationship, and degree of evolutionary change. This type of taxonomy may consider whole taxa rather than single species, so that groups of species can be inferred as giving rise to new groups. The concept found its most well-known form in the modern evolutionary synthesis of the early 1940s.
Wastebasket taxon is a term used by some taxonomists to refer to a taxon that has the purpose of classifying organisms that do not fit anywhere else. They are typically defined by either their designated members' often superficial similarity to each other, or their lack of one or more distinct character states or by their not belonging to one or more other taxa. Wastebasket taxa are by definition either paraphyletic or polyphyletic, and are therefore not considered valid taxa under strict cladistic rules of taxonomy. The name of a wastebasket taxon may in some cases be retained as the designation of an evolutionary grade, however.

A grade is a taxon united by a level of morphological or physiological complexity. The term was coined by British biologist Julian Huxley, to contrast with clade, a strictly phylogenetic unit.
A phylogenetic network is any graph used to visualize evolutionary relationships between nucleotide sequences, genes, chromosomes, genomes, or species. They are employed when reticulation events such as hybridization, horizontal gene transfer, recombination, or gene duplication and loss are believed to be involved. They differ from phylogenetic trees by the explicit modeling of richly linked networks, by means of the addition of hybrid nodes instead of only tree nodes. Phylogenetic trees are a subset of phylogenetic networks. Phylogenetic networks can be inferred and visualised with software such as SplitsTree, the R-package, phangorn, and, more recently, Dendroscope. A standard format for representing phylogenetic networks is a variant of Newick format which is extended to support networks as well as trees.
Computational phylogenetics, phylogeny inference, or phylogenetic inference focuses on computational and optimization algorithms, heuristics, and approaches involved in phylogenetic analyses. The goal is to find a phylogenetic tree representing optimal evolutionary ancestry between a set of genes, species, or taxa. Maximum likelihood, parsimony, Bayesian, and minimum evolution are typical optimality criteria used to assess how well a phylogenetic tree topology describes the sequence data. Nearest Neighbour Interchange (NNI), Subtree Prune and Regraft (SPR), and Tree Bisection and Reconnection (TBR), known as tree rearrangements, are deterministic algorithms to search for optimal or the best phylogenetic tree. The space and the landscape of searching for the optimal phylogenetic tree is known as phylogeny search space.
In mathematics, Newick tree format is a way of representing graph-theoretical trees with edge lengths using parentheses and commas. It was adopted by James Archie, William H. E. Day, Joseph Felsenstein, Wayne Maddison, Christopher Meacham, F. James Rohlf, and David Swofford, at two meetings in 1986, the second of which was at Newick's restaurant in Dover, New Hampshire, US. The adopted format is a generalization of the format developed by Meacham in 1984 for the first tree-drawing programs in Felsenstein's PHYLIP package.
Phylogenetic nomenclature is a method of nomenclature for taxa in biology that uses phylogenetic definitions for taxon names as explained below. This contrasts with the traditional approach, in which taxon names are defined by a type, which can be a specimen or a taxon of lower rank, and a description in words. Phylogenetic nomenclature is currently regulated by the International Code of Phylogenetic Nomenclature (PhyloCode).

The Cichorioideae are a subfamily of the family Asteraceae of flowering plants. Familiar members of Cichorioideae include lettuce, dandelions, chicory and Gazania species. The subfamily comprises about 240 genera and about 2900 species. It is heterogeneous and hard to characterize except with molecular characters.
T-REX is a freely available web server, developed at the department of Computer Science of the Université du Québec à Montréal, dedicated to the inference, validation and visualization of phylogenetic trees and phylogenetic networks. The T-REX web server allows the users to perform several popular methods of phylogenetic analysis as well as some new phylogenetic applications for inferring, drawing and validating phylogenetic trees and networks.
References
- ↑ Hörandl E; Stuessy T.F. (2010). "Paraphyletic groups as natural units of biological classification". Taxon . 59 (2): 345–350. doi: 10.1002/tax.592001 .
- ↑ Stuessy T.F.; König C (May 2008). "Patrocladistic classification". Taxon . 57 (2): 594–601. doi: 10.2307/25066026 . JSTOR 25066026.
- ↑ Achigan-Dako, Enoch G. (2008). Phylogenetic and Genetic Variation Analyses in Cucurbit Species (Cucurbitaceae) from West Africa: definition of Conservation strategies. Cuvillier Verlag. pp. 1–145. ISBN 978-3867277853.
- ↑ Hörandl E. (April 2010). "Beyond cladistics: Extending evolutionary classifications into deeper time levels". Taxon . 59 (2): 345–350. doi: 10.1002/tax.592001 .
- ↑ Wiley E.O. (February 2009). "Patrocladistics, nothing new". Taxon . 58 (1): 2–6. doi: 10.1002/tax.581002 .
- ↑ Fourment M.; Gibbs M. J. (2006). "PATRISTIC: a program for calculating patristic distances and graphically comparing the components of genetic change". BMC Evolutionary Biology. 6: 1–5. doi: 10.1186/1471-2148-6-1 . PMC 1352388 . PMID 16388682.
- ↑ Pommier T.; Canbäck B.; Lundberg P.; Hagström A.; Tunlid A. (2009). "RAMI: a tool for identification and characterization of phylogenetic clusters in microbial communities". Bioinformatics . 25 (6): 736–742. doi:10.1093/bioinformatics/btp051. PMC 2654800 . PMID 19223450.