Related Research Articles

Escherichia coli, also known as E. coli, is a Gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus Escherichia that is commonly found in the lower intestine of warm-blooded organisms (endotherms). Most E. coli strains are harmless, but some serotypes can cause serious food poisoning in their hosts, and are occasionally responsible for product recalls due to food contamination. The harmless strains are part of the normal microbiota of the gut, and can benefit their hosts by producing vitamin K2, (which helps blood to clot when you have a cut to form a scab) and preventing colonisation of the intestine with pathogenic bacteria, having a symbiotic relationship. E. coli is expelled into the environment within fecal matter. The bacterium grows massively in fresh fecal matter under aerobic conditions for 3 days, but its numbers decline slowly afterwards.
A sigma factor is a protein needed for initiation of transcription in bacteria. It is a bacterial transcription initiation factor that enables specific binding of RNA polymerase (RNAP) to gene promoters. It is homologous to archaeal transcription factor B and to eukaryotic factor TFIIB. The specific sigma factor used to initiate transcription of a given gene will vary, depending on the gene and on the environmental signals needed to initiate transcription of that gene. Selection of promoters by RNA polymerase is dependent on the sigma factor that associates with it. They are also found in plant chloroplasts as a part of the bacteria-like plastid-encoded polymerase (PEP).
Virulence factors are molecules produced by bacteria, viruses, fungi, and protozoa that add to their effectiveness and enable them to achieve the following:
The gene rpoS encodes the sigma factor sigma-38, a 37.8 kD protein in Escherichia coli. Sigma factors are proteins that regulate transcription in bacteria. Sigma factors can be activated in response to different environmental conditions. rpoS is transcribed in late exponential phase, and RpoS is the primary regulator of stationary phase genes. RpoS is a central regulator of the general stress response and operates in both a retroactive and a proactive manner: it not only allows the cell to survive environmental challenges, but it also prepares the cell for subsequent stresses (cross-protection). The transcriptional regulator CsgD is central to biofilm formation, controlling the expression of the curli structural and export proteins, and the diguanylate cyclase, adrA, which indirectly activates cellulose production. The rpoS gene most likely originated in the gammaproteobacteria.
In molecular genetics, a regulon is a group of genes that are regulated as a unit, generally controlled by the same regulatory gene that expresses a protein acting as a repressor or activator. This terminology is generally, although not exclusively, used in reference to prokaryotes, whose genomes are often organized into operons; the genes contained within a regulon are usually organized into more than one operon at disparate locations on the chromosome. Applied to eukaryotes, the term refers to any group of non-contiguous genes controlled by the same regulatory gene.
The gene rpoF encodes the sigma factor sigma-28, a protein in Escherichia coli and other species of bacteria. Depending on the bacterial species, this gene may be referred to as sigD or fliA. The protein encoded by this gene has been found to be necessary for flagellum formation.

DicF RNA is a non-coding RNA that is an antisense inhibitor of cell division gene ftsZ. DicF is bound by the Hfq protein which enhances its interaction with its targets. Pathogenic E. coli strains possess multiple copies of sRNA DicF in their genomes, while no-pathogenic strains do not.DicF and Hfq are both necessary to reduce FtsZ protein levels, leading to cell filamentation under anaerobic conditions.

The DnaX ribosomal frameshifting element is a RNA element found in the mRNA of the dnaX gene in E. coli. The dnaX gene has two encoded products, tau and gamma, which are produced in a 1:1 ratio. The gamma protein is synthesised due to programmed frameshifting and is shorter than tau. The two products of the dnaX gene are DNA polymerase III subunits.

DsrA RNA is a non-coding RNA that regulates both transcription, by overcoming transcriptional silencing by the nucleoid-associated H-NS protein, and translation, by promoting efficient translation of the stress sigma factor, RpoS. These two activities of DsrA can be separated by mutation: the first of three stem-loops of the 85 nucleotide RNA is necessary for RpoS translation but not for anti-H-NS action, while the second stem-loop is essential for antisilencing and less critical for RpoS translation. The third stem-loop, which behaves as a transcription terminator, can be substituted by the trp transcription terminator without loss of either DsrA function. The sequence of the first stem-loop of DsrA is complementary with the upstream leader portion of RpoS messenger RNA, suggesting that pairing of DsrA with the RpoS message might be important for translational regulation. The structures of DsrA and DsrA/rpoS complex were studied by NMR. The study concluded that the sRNA contains a dynamic conformational equilibrium for its second stem–loop which might be an important mechanism for DsrA to regulate the translations of its multiple target mRNAs.

The gcvB RNA gene encodes a small non-coding RNA involved in the regulation of a number of amino acid transport systems as well as amino acid biosynthetic genes. The GcvB gene is found in enteric bacteria such as Escherichia coli. GcvB regulates genes by acting as an antisense binding partner of the mRNAs for each regulated gene. This binding is dependent on binding to a protein called Hfq. Transcription of the GcvB RNA is activated by the adjacent GcvA gene and repressed by the GcvR gene. A deletion of GcvB RNA from Y. pestis changed colony shape as well as reducing growth. It has been shown by gene deletion that GcvB is a regulator of acid resistance in E. coli. GcvB enhances the ability of the bacterium to survive low pH by upregulating the levels of the alternate sigma factor RpoS. A polymeric form of GcvB has recently been identified. Interaction of GcvB with small RNA SroC triggers the degradation of GcvB by RNase E, lifting the GcvB-mediated mRNA repression of its target genes.
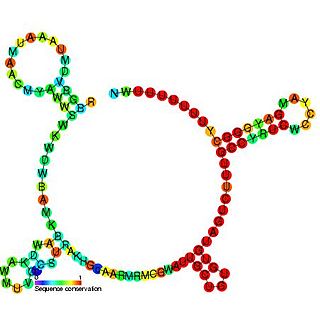
The RprA RNA gene encodes a 106 nucleotide regulatory non-coding RNA. Translational regulation of the stationary phase sigma factor RpoS is mediated by the formation of a double-stranded RNA stem-loop structure in the upstream region of the rpoS messenger RNA, occluding the translation initiation site. Clones carrying rprA increased the translation of RpoS. As with DsrA, RprA is predicted to form three stem-loops. Thus, at least two small RNAs, DsrA and RprA, participate in the positive regulation of RpoS translation. RprA also appears to bind to the RpoS leader. RprA is non-essential. Wasserman et al. demonstrated that this RNA is bound by the Hfq protein. Binding to Hfq alters the conformation of RprA. In the presence of Hfq the stability of RprA is influenced by the osmolarity of the cell, this is dependent on the endoribonuclease RNase E.

Spot 42 (spf) RNA is a regulatory non-coding bacterial small RNA encoded by the spf gene. Spf is found in gammaproteobacteria and the majority of experimental work on Spot42 has been performed in Escherichia coli and recently in Aliivibrio salmonicida. In the cell Spot42 plays essential roles as a regulator in carbohydrate metabolism and uptake, and its expression is activated by glucose, and inhibited by the cAMP-CRP complex.

The Hfq protein encoded by the hfq gene was discovered in 1968 as an Escherichia coli host factor that was essential for replication of the bacteriophage Qβ. It is now clear that Hfq is an abundant bacterial RNA binding protein which has many important physiological roles that are usually mediated by interacting with Hfq binding sRNA.
In enzymology, a phosphoglucosamine mutase is an enzyme that catalyzes the chemical reaction

A toxin-antitoxin system is a set of two or more closely linked genes that together encode both a "toxin" protein and a corresponding "antitoxin". Toxin-antitoxin systems are widely distributed in prokaryotes, and organisms often have them in multiple copies. When these systems are contained on plasmids – transferable genetic elements – they ensure that only the daughter cells that inherit the plasmid survive after cell division. If the plasmid is absent in a daughter cell, the unstable antitoxin is degraded and the stable toxic protein kills the new cell; this is known as 'post-segregational killing' (PSK).
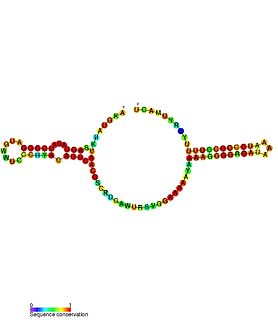
Escherichia coli contains a number of small RNAs located in intergenic regions of its genome. The presence of at least 55 of these has been verified experimentally. 275 potential sRNA-encoding loci were identified computationally using the QRNA program. These loci will include false positives, so the number of sRNA genes in E. coli is likely to be less than 275. A computational screen based on promoter sequences recognised by the sigma factor sigma 70 and on Rho-independent terminators predicted 24 putative sRNA genes, 14 of these were verified experimentally by northern blotting. The experimentally verified sRNAs included the well characterised sRNAs RprA and RyhB. Many of the sRNAs identified in this screen, including RprA, RyhB, SraB and SraL, are only expressed in the stationary phase of bacterial cell growth. A screen for sRNA genes based on homology to Salmonella and Klebsiella identified 59 candidate sRNA genes. From this set of candidate genes, microarray analysis and northern blotting confirmed the existence of 17 previously undescribed sRNAs, many of which bind to the chaperone protein Hfq and regulate the translation of RpoS. UptR sRNA transcribed from the uptR gene is implicated in suppressing extracytoplasmic toxicity by reducing the amount of membrane-bound toxic hybrid protein.
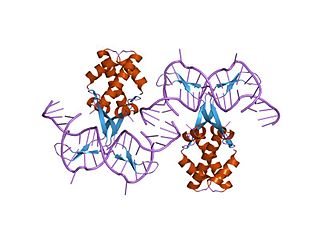
In molecular biology, bacterial DNA binding proteins are a family of small, usually basic proteins of about 90 residues that bind DNA and are known as histone-like proteins. Since bacterial binding proteins have a diversity of functions, it has been difficult to develop a common function for all of them. They are commonly referred to as histone-like and have many similar traits with the eukaryotic histone proteins. Eukaryotic histones package DNA to help it to fit in the nucleus, and they are known to be the most conserved proteins in nature. Examples include the HU protein in Escherichia coli, a dimer of closely related alpha and beta chains and in other bacteria can be a dimer of identical chains. HU-type proteins have been found in a variety of eubacteria and archaebacteria, and are also encoded in the chloroplast genome of some algae. The integration host factor (IHF), a dimer of closely related chains which is suggested to function in genetic recombination as well as in translational and transcriptional control is found in Enterobacteria and viral proteins including the African swine fever virus protein A104R.

The gab operon is responsible for the conversion of γ-aminobutyrate (GABA) to succinate. The gab operon comprises three structural genes – gabD, gabT and gabP – that encode for a succinate semialdehyde dehydrogenase, GABA transaminase and a GABA permease respectively. There is a regulatory gene csiR, downstream of the operon, that codes for a putative transcriptional repressor and is activated when nitrogen is limiting.
The gene rpoN encodes the sigma factor sigma-54, a protein in Escherichia coli and other species of bacteria. RpoN antagonizes RpoS sigma factors.
The Monovalent Cation:Proton Antiporter-2 (CPA2) Family is a moderately large family of transporters belonging to the CPA superfamily. Members of the CPA2 family have been found in bacteria, archaea and eukaryotes. The proteins of the CPA2 family consist of between 333 and 900 amino acyl residues and exhibit 10-14 transmembrane α-helical spanners (TMSs).
References
- ↑ Rouvière, PE; De Las Peñas, A; Mecsas, J; Lu, CZ; Rudd, KE; Gross, CA (1995). "rpoE, the gene encoding the second heat-shock sigma factor, sigma E, in Escherichia coli". The EMBO Journal. 14 (5): 1032–42. doi:10.1002/j.1460-2075.1995.tb07084.x. PMC 398175 . PMID 7889934.
- ↑ Jensen-Cain, DM; Quinn, FD (2001). "Differential expression of sigE by Mycobacterium tuberculosis during intracellular growth". Microbial Pathogenesis. 30 (5): 271–8. doi:10.1006/mpat.2001.0431. PMID 11373121.
- ↑ Hiratsu, K; Amemura, M; Nashimoto, H; Shinagawa, H; Makino, K (1995). "The rpoE gene of Escherichia coli, which encodes sigma E, is essential for bacterial growth at high temperature". Journal of Bacteriology. 177 (10): 2918–22. doi:10.1128/jb.177.10.2918-2922.1995. PMC 176969 . PMID 7751307.
- ↑ De Las Peñas, A; Connolly, L; Gross, CA (Nov 1997). "SigmaE is an essential sigma factor in Escherichia coli". Journal of Bacteriology. 179 (21): 6862–4. doi:10.1128/jb.179.21.6862-6864.1997. PMC 179621 . PMID 9352942.
- ↑ Kirk, DG; Zhang, Z; Korkeala, H; Lindström, M (2014). "Alternative sigma factors SigF, SigE, and SigG are essential for sporulation in Clostridium botulinum ATCC 3502". Applied and Environmental Microbiology. 80 (16): 5141–50. doi:10.1128/aem.01015-14. PMC 4135750 . PMID 24928875.
- ↑ De Las Peñas, A; Connolly, L; Gross, CA (Nov 1997). "SigmaE is an essential sigma factor in Escherichia coli". Journal of Bacteriology. 179 (21): 6862–4. doi:10.1128/jb.179.21.6862-6864.1997. PMC 179621 . PMID 9352942.
| This gene article is a stub. You can help Wikipedia by expanding it. |