Related Research Articles

In molecular biology, DNA replication is the biological process of producing two identical replicas of DNA from one original DNA molecule. DNA replication occurs in all living organisms acting as the most essential part for biological inheritance. This is essential for cell division during growth and repair of damaged tissues, while it also ensures that each of the new cells receives its own copy of the DNA. The cell possesses the distinctive property of division, which makes replication of DNA essential.
Semiconservative replication describes the mechanism of DNA replication in all known cells. DNA replication occurs on multiple origins of replication along the DNA template strand. As the DNA double helix is unwound by helicase, replication occurs separately on each template strand in antiparallel directions. This process is known as semi-conservative replication because two copies of the original DNA molecule are produced. Each copy contains one original strand and one newly-synthesized strand. The structure of DNA suggested that each strand of the double helix would serve as a template for synthesis of a new strand. It was not known how newly synthesized strands combined with template strands to form two double helical DNA molecules.
Topoisomerases are enzymes that participate in the overwinding or underwinding of DNA. The winding problem of DNA arises due to the intertwined nature of its double-helical structure. During DNA replication and transcription, DNA becomes overwound ahead of a replication fork. If left unabated, this torsion would eventually stop the ability of DNA or RNA polymerases involved in these processes to continue down the DNA strand.

The nucleoid is an irregularly shaped region within the prokaryotic cell that contains all or most of the genetic material. The chromosome of a prokaryote is circular, and its length is very large compared to the cell dimensions needing it to be compacted in order to fit. In contrast to the nucleus of a eukaryotic cell, it is not surrounded by a nuclear membrane. Instead, the nucleoid forms by condensation and functional arrangement with the help of chromosomal architectural proteins and RNA molecules as well as DNA supercoiling. The length of a genome widely varies and a cell may contain multiple copies of it.
DNA gyrase, or simply gyrase, is an enzyme within the class of topoisomerase and is a subclass of Type II topoisomerases that reduces topological strain in an ATP dependent manner while double-stranded DNA is being unwound by elongating RNA-polymerase or by helicase in front of the progressing replication fork. The enzyme causes negative supercoiling of the DNA or relaxes positive supercoils. It does so by looping the template so as to form a crossing, then cutting one of the double helices and passing the other through it before releasing the break, changing the linking number by two in each enzymatic step. This process occurs in bacteria, whose single circular DNA is cut by DNA gyrase and the two ends are then twisted around each other to form supercoils. Gyrase is also found in eukaryotic plastids: it has been found in the apicoplast of the malarial parasite Plasmodium falciparum and in chloroplasts of several plants. Bacterial DNA gyrase is the target of many antibiotics, including nalidixic acid, novobiocin, and ciprofloxacin.

In molecular biology, the term double helix refers to the structure formed by double-stranded molecules of nucleic acids such as DNA. The double helical structure of a nucleic acid complex arises as a consequence of its secondary structure, and is a fundamental component in determining its tertiary structure. The term entered popular culture with the publication in 1968 of The Double Helix: A Personal Account of the Discovery of the Structure of DNA by James Watson, while the structure itself was originally discovered by Rosalind Franklin.
A nick is a discontinuity in a double stranded DNA molecule where there is no phosphodiester bond between adjacent nucleotides of one strand typically through damage or enzyme action. Nicks allow DNA strands to untwist during replication, and are also thought to play a role in the DNA mismatch repair mechanisms that fix errors on both the leading and lagging daughter strands.

The replisome is a complex molecular machine that carries out replication of DNA. The replisome first unwinds double stranded DNA into two single strands. For each of the resulting single strands, a new complementary sequence of DNA is synthesized. The net result is formation of two new double stranded DNA sequences that are exact copies of the original double stranded DNA sequence.
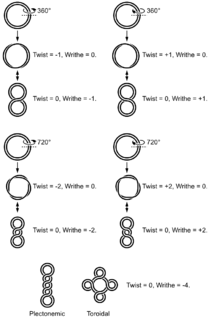
DNA supercoiling refers to the over- or under-winding of a DNA strand, and is an expression of the strain on that strand. Supercoiling is important in a number of biological processes, such as compacting DNA, and by regulating access to the genetic code, DNA supercoiling strongly affects DNA metabolism and possibly gene expression. Additionally, certain enzymes such as topoisomerases are able to change DNA topology to facilitate functions such as DNA replication or transcription. Mathematical expressions are used to describe supercoiling by comparing different coiled states to relaxed B-form DNA.
Topoisomerase inhibitors are chemical compounds that block the action of topoisomerases, which are broken into two broad subtypes: type I topoisomerases (TopI) and type II topoisomerases (TopII). Topoisomerase plays important roles in cellular reproduction and DNA organization, as they mediate the cleavage of single and double stranded DNA to relax supercoils, untangle catenanes, and condense chromosomes in eukaryotic cells. Topoisomerase inhibitors influence these essential cellular processes. Some topoisomerase inhibitors prevent topoisomerases from performing DNA strand breaks while others, deemed topoisomerase poisons, associate with topoisomerase-DNA complexes and prevent the re-ligation step of the topoisomerase mechanism. These topoisomerase-DNA-inhibitor complexes are cytotoxic agents, as the un-repaired single and double stranded DNA breaks that they cause can lead to apoptosis and cell death. Because of this ability to induce apoptosis, topoisomerase inhibitors have gained interest as therapeutics against infectious and cancerous cells.
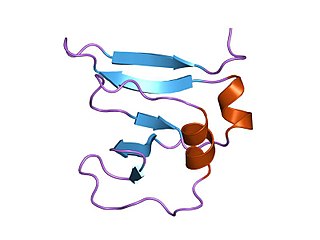
In molecular biology Type I topoisomerases are enzymes that cut one of the two strands of double-stranded DNA, relax the strand, and reanneal the strand. They are further subdivided into two structurally and mechanistically distinct topoisomerases: type IA and type IB.

Type II topoisomerases are topoisomerases that cut both strands of the DNA helix simultaneously in order to manage DNA tangles and supercoils. They use the hydrolysis of ATP, unlike Type I topoisomerase. In this process, these enzymes change the linking number of circular DNA by ±2.
In enzymology, glutamate racemase is an enzyme that catalyzes the chemical reaction
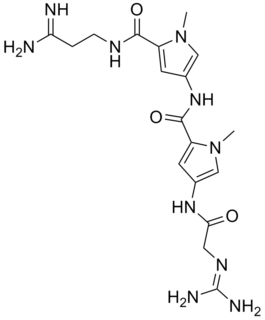
Netropsin is a polyamide with antibiotic and antiviral activity. Netropsin was discovered by Finlay et al., and first isolated from the actinobacterium Streptomyces netropsis. It belongs to the class of pyrrole-amidine antibiotics.

A circular chromosome is a chromosome in bacteria, archaea, mitochondria, and chloroplasts, in the form of a molecule of circular DNA, unlike the linear chromosome of most eukaryotes.

Nucleic acid structure refers to the structure of nucleic acids such as DNA and RNA. Chemically speaking, DNA and RNA are very similar. Nucleic acid structure is often divided into four different levels: primary, secondary, tertiary, and quaternary.

In addition to the variety of verified DNA structures, there have been a range of obsolete models that have either been disproven, or lack evidence.

Neisseria gonorrhoeae, the bacterium that causes the sexually transmitted infection gonorrhea, has developed antibiotic resistance to many antibiotics. The bacteria was first identified in 1879.
Saccharolobus solfataricus is a species of thermophilic archaeon. It was transferred from the genus Sulfolobus to the new genus Saccharolobus with the description of Saccharolobus caldissimus in 2018.
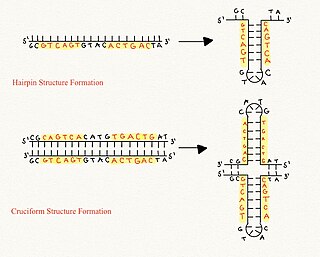
Cruciform DNA is a form of non-B DNA, or an alternative DNA structure. The formation of cruciform DNA requires the presence of palindromes called inverted repeat sequences. These inverted repeats contain a sequence of DNA in one strand that is repeated in the opposite direction on the other strand. As a result, inverted repeats are self-complementary and can give rise to structures such as hairpins and cruciforms. Cruciform DNA structures require at least a six nucleotide sequence of inverted repeats to form a structure consisting of a stem, branch point and loop in the shape of a cruciform, stabilized by negative DNA supercoiling.
References
- ↑ Kato, Jun-ichi; Suzuki, Hideho; Ikeda, Hideo (25 December 1992). "Purification and characterization of DNA topoisomerase IV in Escherichia coli". Journal of Biological Chemistry . 267 (36): 25676–25684. doi: 10.1016/S0021-9258(18)35660-6 . PMID 1334483.
- ↑ Rawdon, Eric; Dorier, Julien; Račko, Dušan; Millet, Kenneth C.; Stasiak, Andrzej (22 April 2016). "How topoisomerase IV can efficiently unknot and decatenate negatively supercoiled DNA molecules without causing their torsional relaxation". Nucleic Acids Research . 44 (10): 4528–4538. doi:10.1093/nar/gkw311. PMC 4889953 . PMID 27106058.