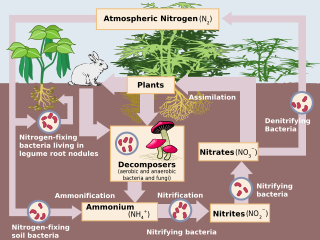
Nitrification is the biological oxidation of ammonia to nitrate via the intermediary nitrite. Nitrification is an important step in the nitrogen cycle in soil. The process of complete nitrification may occur through separate organisms or entirely within one organism, as in comammox bacteria. The transformation of ammonia to nitrite is usually the rate limiting step of nitrification. Nitrification is an aerobic process performed by small groups of autotrophic bacteria and archaea.
The Aquificota phylum is a diverse collection of bacteria that live in harsh environmental settings. The name Aquificota was given to this phylum based on an early genus identified within this group, Aquifex, which is able to produce water by oxidizing hydrogen. They have been found in springs, pools, and oceans. They are autotrophs, and are the primary carbon fixers in their environments. These bacteria are Gram-negative, non-spore-forming rods. They are true bacteria as opposed to the other inhabitants of extreme environments, the Archaea.
The Thermomicrobia is a group of thermophilic green non-sulfur bacteria. Based on species Thermomicrobium roseum and Sphaerobacter thermophilus, this bacteria class has the following description:

Anammox, an abbreviation for "anaerobic ammonium oxidation", is a globally important microbial process of the nitrogen cycle that takes place in many natural environments. The bacteria mediating this process were identified in 1999, and were a great surprise for the scientific community. In the anammox reaction, nitrite and ammonium ions are converted directly into diatomic nitrogen and water.
Nitrosomonas europaea is a Gram-negative obligate chemolithoautotroph that can derive all its energy and reductant for growth from the oxidation of ammonia to nitrite and lives in several places such as soil, sewage, freshwater, the walls of buildings and on the surface of monuments especially in polluted areas where the air contains high levels of nitrogen compounds.

Nitrosomonas is a genus of Gram-negative bacteria, belonging to the Betaproteobacteria. It is one of the five genera of ammonia-oxidizing bacteria and, as an obligate chemolithoautotroph, uses ammonia as an energy source and carbon dioxide as a carbon source in the presence of oxygen. Nitrosomonas are important in the global biogeochemical nitrogen cycle, since they increase the bioavailability of nitrogen to plants and in the denitrification, which is important for the release of nitrous oxide, a powerful greenhouse gas. This microbe is photophobic, and usually generate a biofilm matrix, or form clumps with other microbes, to avoid light. Nitrosomonas can be divided into six lineages: the first one includes the species Nitrosomonas europea, Nitrosomonas eutropha, Nitrosomonas halophila, and Nitrosomonas mobilis. The second lineage presents the species Nitrosomonas communis, N. sp. I and N. sp. II. The third lineage includes only Nitrosomonas nitrosa. The fourth lineage includes the species Nitrosomonas ureae and Nitrosomonas oligotropha. The fifth and sixth lineages include the species Nitrosomonas marina, N. sp. III, Nitrosomonas estuarii, and Nitrosomonas cryotolerans.
Nitrobacter is a genus comprising rod-shaped, gram-negative, and chemoautotrophic bacteria. The name Nitrobacter derives from the Latin neuter gender noun nitrum, nitri, alkalis; the Ancient Greek noun βακτηρία, βακτηρίᾱς, rod. They are non-motile and reproduce via budding or binary fission. Nitrobacter cells are obligate aerobes and have a doubling time of about 13 hours.
Microbial metabolism is the means by which a microbe obtains the energy and nutrients it needs to live and reproduce. Microbes use many different types of metabolic strategies and species can often be differentiated from each other based on metabolic characteristics. The specific metabolic properties of a microbe are the major factors in determining that microbe's ecological niche, and often allow for that microbe to be useful in industrial processes or responsible for biogeochemical cycles.
Nitrifying bacteria are chemolithotrophic organisms that include species of genera such as Nitrosomonas, Nitrosococcus, Nitrobacter, Nitrospina, Nitrospira and Nitrococcus. These bacteria get their energy from the oxidation of inorganic nitrogen compounds. Types include ammonia-oxidizing bacteria (AOB) and nitrite-oxidizing bacteria (NOB). Many species of nitrifying bacteria have complex internal membrane systems that are the location for key enzymes in nitrification: ammonia monooxygenase, hydroxylamine oxidoreductase, and nitrite oxidoreductase.
Nitrospira translate into “a nitrate spiral” is a genus of bacteria within the monophyletic clade of the Nitrospirota phylum. The first member of this genus was described 1986 by Watson et al., isolated from the Gulf of Maine. The bacterium was named Nitrospira marina. Populations were initially thought to be limited to marine ecosystems, but it was later discovered to be well-suited for numerous habitats, including activated sludge of wastewater treatment systems, natural biological marine settings, water circulation biofilters in aquarium tanks, terrestrial systems, fresh and salt water ecosystems, agricultural lands and hot springs. Nitrospira is a ubiquitous bacterium that plays a role in the nitrogen cycle by performing nitrite oxidation in the second step of nitrification. Nitrospira live in a wide array of environments including but not limited to, drinking water systems, waste treatment plants, rice paddies, forest soils, geothermal springs, and sponge tissue. Despite being abundant in many natural and engineered ecosystems Nitrospira are difficult to culture, so most knowledge of them is from molecular and genomic data. However, due to their difficulty to be cultivated in laboratory settings, the entire genome was only sequenced in one species, Nitrospira defluvii. In addition, Nitrospira bacteria's 16S rRNA sequences are too dissimilar to use for PCR primers, thus some members go unnoticed. In addition, members of Nitrospira with the capabilities to perform complete nitrification has also been discovered and cultivated.
Nitrite oxidoreductase is an enzyme involved in nitrification. It is the last step in the process of aerobic ammonia oxidation, which is carried out by two groups of nitrifying bacteria: ammonia oxidizers such as Nitrosospira, Nitrosomonas, and Nitrosococcus convert ammonia to nitrite, while nitrite oxidizers such as Nitrobacter and Nitrospira oxidize nitrite to nitrate. NXR is responsible for producing almost all nitrate found in nature.
Anaerobic oxidation of methane (AOM) is a methane-consuming microbial process occurring in anoxic marine and freshwater sediments. AOM is known to occur among mesophiles, but also in psychrophiles, thermophiles, halophiles, acidophiles, and alkophiles. During AOM, methane is oxidized with different terminal electron acceptors such as sulfate, nitrate, nitrite and metals, either alone or in syntrophy with a partner organism.

Cytochrome c nitrite reductase (ccNiR) is a bacterial enzyme that catalyzes the six electron reduction of nitrite to ammonia; an important step in the biological nitrogen cycle. The enzyme catalyses the second step in the two step conversion of nitrate to ammonia, which allows certain bacteria to use nitrite as a terminal electron acceptor, rather than oxygen, during anaerobic conditions. During this process, ccNiR draws electrons from the quinol pool, which are ultimately provided by a dehydrogenase such as formate dehydrogenase or hydrogenase. These dehydrogenases are responsible for generating a proton motive force.
Nitrospirota is a phylum of bacteria. It includes multiple genera, such as Nitrospira, the largest. The first member of this phylum, Nitrospira marina, was discovered in 1985. The second member, Nitrospira moscoviensis, was discovered in 1995.
Nitrososphaera is a mesophilic genus of ammonia-oxidizing Crenarchaeota. The first Nitrososphaera organism was discovered in garden soils at the University of Vienna leading to the categorization of a new genus, family, order and class of Archaea. This genus is contains three distinct species: N. viennensis, Ca. N. gargensis, and Ca N. evergladensis. Nitrososphaera are chemolithoautotrophs and have important biogeochemical roles as nitrifying organisms.
Comammox is the name attributed to an organism that can convert ammonia into nitrite and then into nitrate through the process of nitrification. Nitrification has traditionally been thought to be a two-step process, where ammonia-oxidizing bacteria and archaea oxidize ammonia to nitrite and then nitrite-oxidizing bacteria convert to nitrate. Complete conversion of ammonia into nitrate by a single microorganism was first predicted in 2006. In 2015 the presence of microorganisms that could carry out both conversion processes was discovered within the genus Nitrospira, and the nitrogen cycle was updated. Within the genus Nitrospira, the major ecosystems comammox are primarily found in natural aquifers and engineered ecosystems.
Nitrososphaera gargensis is a non-pathogenic, small coccus measuring 0.9 ± 0.3 μm in diameter. N. gargensis is observed in small abnormal cocci groupings and uses its archaella to move via chemotaxis. Being an Archaeon, Nitrososphaera gargensis has a cell membrane composed of crenarchaeol, its isomer, and a distinct glycerol dialkyl glycerol tetraether (GDGT), which is significant in identifying ammonia-oxidizing archaea (AOA). The organism plays a role in influencing ocean communities and food production.
Nitrospira inopinata is a bacterium from the phylum Nitrospirota. This phylum contains nitrite-oxidizing bacteria playing role in nitrification. However N. inopinata was shown to perform complete ammonia oxidation to nitrate thus being the first comammox bacterium to be discovered.
Nitrospinota is a bacterial phylum. Despite only few described species, members of this phylum are major nitrite-oxidizing bacteria in surface waters in oceans. By oxidation of nitrite to nitrate they are important in the process of nitrification in marine environments.
Nitrosococcus is a genus of Gram-negative bacteria. It is an ammonia-oxidizing bacterium which oxidizes ammonia into nitrite. Nitrosococcus can be found in marine environments and salt lakes.




