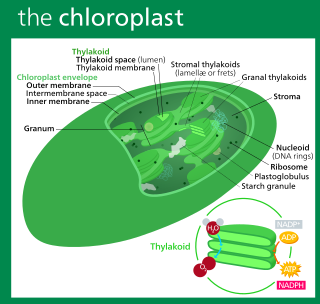
A chloroplast is a type of organelle known as a plastid that conducts photosynthesis mostly in plant and algal cells. Chloroplasts have a high concentration of chlorophyll pigments which capture the energy from sunlight and convert it to chemical energy and release oxygen. The chemical energy created is then used to make sugar and other organic molecules from carbon dioxide in a process called the Calvin cycle. Chloroplasts carry out a number of other functions, including fatty acid synthesis, amino acid synthesis, and the immune response in plants. The number of chloroplasts per cell varies from one, in some unicellular algae, up to 100 in plants like Arabidopsis and wheat.

Chlorophyta is a division of green algae informally called chlorophytes.

A genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA. The nuclear genome includes protein-coding genes and non-coding genes, other functional regions of the genome such as regulatory sequences, and often a substantial fraction of junk DNA with no evident function. Almost all eukaryotes have mitochondria and a small mitochondrial genome. Algae and plants also contain chloroplasts with a chloroplast genome.
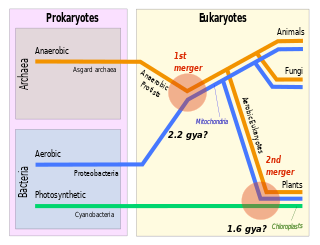
Symbiogenesis is the leading evolutionary theory of the origin of eukaryotic cells from prokaryotic organisms. The theory holds that mitochondria, plastids such as chloroplasts, and possibly other organelles of eukaryotic cells are descended from formerly free-living prokaryotes taken one inside the other in endosymbiosis. Mitochondria appear to be phylogenetically related to Rickettsiales bacteria, while chloroplasts are thought to be related to cyanobacteria.

The green algae are a group of chlorophyll-containing autotrophic eukaryotes consisting of the phylum Prasinodermophyta and its unnamed sister group that contains the Chlorophyta and Charophyta/Streptophyta. The land plants (Embryophytes) have emerged deep within the charophytes as a sister of the Zygnematophyceae. Since the realization that the Embryophytes emerged within the green algae, some authors are starting to include them. The completed clade that includes both green algae and embryophytes is monophyletic and is referred to as the clade Viridiplantae and as the kingdom Plantae. The green algae include unicellular and colonial flagellates, most with two flagella per cell, as well as various colonial, coccoid (spherical), and filamentous forms, and macroscopic, multicellular seaweeds. There are about 22,000 species of green algae, many of which live most of their lives as single cells, while other species form coenobia (colonies), long filaments, or highly differentiated macroscopic seaweeds.

Chlamydomonas reinhardtii is a single-cell green alga about 10 micrometres in diameter that swims with two flagella. It has a cell wall made of hydroxyproline-rich glycoproteins, a large cup-shaped chloroplast, a large pyrenoid, and an eyespot apparatus that senses light.

Streptophyta, informally the streptophytes, is a clade of plants. The composition of the clade varies considerably between authors, but the definition employed here includes land plants and all green algae except the Chlorophyta and the more basal Prasinodermophyta.

Charophyta is a group of freshwater green algae, called charophytes, sometimes treated as a division, yet also as a superdivision or an unranked clade. The terrestrial plants, the Embryophyta emerged deep within Charophyta, possibly from terrestrial unicellular charophytes, with the class Zygnematophyceae as a sister group.

Sphaeropleales is an order of green algae that used to be called Chlorococcales. The order includes some of the most common freshwater planktonic algae such as Scenedesmus and Pediastrum. The Sphaeropleales includes vegetatively non-motile unicellular, colonial, or filamentous taxa. They have biflagellate zoospores with flagella that are directly opposed in direction : Sphaeroplea, Atractomorpha, Neochloris, Hydrodictyon, and Pediastrum. All of these taxa have basal body core connections. Motile cells generally lack cell walls or have only a very fine layer surrounding the cell membrane. Other common characteristics include a robust vegetative cell wall, cup-shaped chloroplasts with large pyrenoids, and relatively large nuclei.

Viridiplantae is a clade of around 450,000–500,000 species of eukaryotic organisms, most of which obtain their energy by photosynthesis. The green plants are chloroplast-bearing autotrophs that play important primary production roles in both terrestrial and aquatic ecosystems. They include green algae, which are primarily aquatic, and the land plants, which emerged within freshwater green algae. Green algae traditionally excludes the land plants, rendering them a paraphyletic group, however it is cladistically accurate to think of land plants as a special clade of green algae that evolved to thrive on dry land. Since the realization that the embryophytes emerged from within the green algae, some authors are starting to include them.

The Archaeplastida are a major group of eukaryotes, comprising the photoautotrophic red algae (Rhodophyta), green algae, land plants, and the minor group glaucophytes. It also includes the non-photosynthetic lineage Rhodelphidia, a predatorial (eukaryotrophic) flagellate that is sister to the Rhodophyta, and probably the microscopic picozoans. The Archaeplastida have chloroplasts that are surrounded by two membranes, suggesting that they were acquired directly through a single endosymbiosis event by phagocytosis of a cyanobacterium. All other groups which have chloroplasts, besides the amoeboid genus Paulinella, have chloroplasts surrounded by three or four membranes, suggesting they were acquired secondarily from red or green algae. Unlike red and green algae, glaucophytes have never been involved in secondary endosymbiosis events.

The prasinophytes are a group of unicellular green algae. Prasinophytes mainly include marine planktonic species, as well as some freshwater representatives. The prasinophytes are morphologically diverse, including flagellates with one to eight flagella and non-motile (coccoid) unicells. The cells of many species are covered with organic body scales; others are naked. Well studied genera include Ostreococcus, considered to be the smallest free-living eukaryote, and Micromonas, both of which are found in marine waters worldwide. Prasinophytes have simple cellular structures, containing a single chloroplast and a single mitochondrion. The genomes are relatively small compared to other eukaryotes . At least one species, the Antarctic form Pyramimonas gelidicola, is capable of phagocytosis and is therefore a mixotrophic algae.

The Trebouxiophyceae, also known as trebouxiophytes, are a class of green algae, in the division Chlorophyta. Members of this class are single-celled, colonial, or multicellular and are found in freshwater or terrestrial habitats worldwide. Many taxa in the Trebouxiophyceae form symbiotic relationships with other organisms; in particular, the majority of phycobionts within lichens are trebouxiophytes. A number of taxa have also lost the ability to photosynthesize, and have evolved to become parasitic; examples include Prototheca and Helicosporidium.

Prototheca is a genus of algae in the family Chlorellaceae. While this genus is a member of the green algae, all Prototheca no longer have chloroplasts and therefore their photosynthetic ability. Some species can cause protothecosis in humans and various vertebrates.
Pseudomuriella is a genus of green algae, specifically of the class Chlorophyceae. It is the sole genus of the family Pseudomuriellaceae. It is a terrestrial alga that inhabits soils.

Zygnematophyceae is a class of green algae in the paraphylum streptophyte algae, also referred to as Charophyta, consisting of more than 4000 described species. The Zygnematophyceae are the sister clade of the Embryophyta.
The CoRR hypothesis states that the location of genetic information in cytoplasmic organelles permits regulation of its expression by the reduction-oxidation ("redox") state of its gene products.
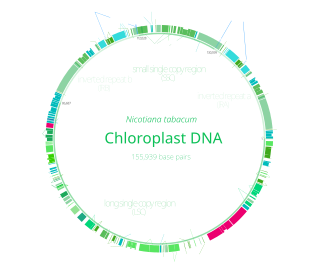
Chloroplast DNA (cpDNA), also known as plastid DNA (ptDNA) is the DNA located in chloroplasts, which are photosynthetic organelles located within the cells of some eukaryotic organisms. Chloroplasts, like other types of plastid, contain a genome separate from that in the cell nucleus. The existence of chloroplast DNA was identified biochemically in 1959, and confirmed by electron microscopy in 1962. The discoveries that the chloroplast contains ribosomes and performs protein synthesis revealed that the chloroplast is genetically semi-autonomous. The first complete chloroplast genome sequences were published in 1986, Nicotiana tabacum (tobacco) by Sugiura and colleagues and Marchantia polymorpha (liverwort) by Ozeki et al. Since then, tens of thousands of chloroplast genomes from various species have been sequenced.

Cyanidioschyzon merolae is a small (2μm), club-shaped, unicellular haploid red alga adapted to high sulfur acidic hot spring environments. The cellular architecture of C. merolae is extremely simple, containing only a single chloroplast and a single mitochondrion and lacking a vacuole and cell wall. In addition, the cellular and organelle divisions can be synchronized. For these reasons, C. merolae is considered an excellent model system for study of cellular and organelle division processes, as well as biochemistry and structural biology. The organism's genome was the first full algal genome to be sequenced in 2004; its plastid was sequenced in 2000 and 2003, and its mitochondrion in 1998. The organism has been considered the simplest of eukaryotic cells for its minimalist cellular organization.

















