
A retrovirus is a type of virus that inserts a DNA copy of its RNA genome into the DNA of a host cell that it invades, thus changing the genome of that cell. After invading a host cell's cytoplasm, the virus uses its own reverse transcriptase enzyme to produce DNA from its RNA genome, the reverse of the usual pattern, thus retro (backwards). The new DNA is then incorporated into the host cell genome by an integrase enzyme, at which point the retroviral DNA is referred to as a provirus. The host cell then treats the viral DNA as part of its own genome, transcribing and translating the viral genes along with the cell's own genes, producing the proteins required to assemble new copies of the virus. Many retroviruses cause serious diseases in humans, other mammals, and birds.

Endogenous retroviruses (ERVs) are endogenous viral elements in the genome that closely resemble and can be derived from retroviruses. They are abundant in the genomes of jawed vertebrates, and they comprise up to 5–8% of the human genome.

Furin is a protease, a proteolytic enzyme that in humans and other animals is encoded by the FURIN gene. Some proteins are inactive when they are first synthesized, and must have sections removed in order to become active. Furin cleaves these sections and activates the proteins. It was named furin because it was in the upstream region of an oncogene known as FES. The gene was known as FUR and therefore the protein was named furin. Furin is also known as PACE. A member of family S8, furin is a subtilisin-like peptidase.
Simian foamy virus (SFV) is a species of the genus Spumavirus that belongs to the family of Retroviridae. It has been identified in a wide variety of primates, including prosimians, New World and Old World monkeys, as well as apes, and each species has been shown to harbor a unique (species-specific) strain of SFV, including African green monkeys, baboons, macaques, and chimpanzees. As it is related to the more well-known retrovirus human immunodeficiency virus (HIV), its discovery in primates has led to some speculation that HIV may have been spread to the human species in Africa through contact with blood from apes, monkeys, and other primates, most likely through bushmeat-hunting practices.
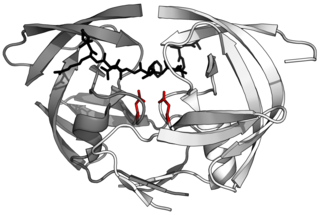
Aspartic proteases are a catalytic type of protease enzymes that use an activated water molecule bound to one or more aspartate residues for catalysis of their peptide substrates. In general, they have two highly conserved aspartates in the active site and are optimally active at acidic pH. Nearly all known aspartyl proteases are inhibited by pepstatin.
Group-specific antigen, or gag, is the polyprotein that contains the core structural proteins of an Ortervirus. It was named as such because scientists used to believe it was antigenic. Now it is known that it makes up the inner shell, not the envelope exposed outside. It makes up all the structural units of viral conformation and provides supportive framework for mature virion.

HIV-1 protease (PR) is a retroviral aspartyl protease (retropepsin), an enzyme involved with peptide bond hydrolysis in retroviruses, that is essential for the life-cycle of HIV, the retrovirus that causes AIDS. HIV protease cleaves newly synthesized polyproteins at nine cleavage sites to create the mature protein components of an HIV virion, the infectious form of a virus outside of the host cell. Without effective HIV protease, HIV virions remain uninfectious.

Syncytin-1 also known as enverin is a protein found in humans and other primates that is encoded by the ERVW-1 gene. Syncytin-1 is a cell-cell fusion protein whose function is best characterized in placental development. The placenta in turn aids in embryo attachment to the uterus and establishment of a nutrient supply.

HERV-R_7q21.2 provirus ancestral envelope (Env) polyprotein is a protein that in humans is encoded by the ERV3 gene.
HERV-K_19q12 provirus ancestral Pol protein is a protein that in humans is encoded by the ERVK6 gene.

Legumain is a protein that in humans is encoded by the LGMN gene.

Syncytin-2 also known as endogenous retrovirus group FRD member 1 is a protein that in humans is encoded by the ERVFRD-1 gene. This protein plays a key role in the implantation of human embryos in the womb.
Many major physiological processes depend on regulation of proteolytic enzyme activity and there can be dramatic consequences when equilibrium between an enzyme and its substrates is disturbed. In this prospective, the discovery of small-molecule ligands, like protease inhibitors, that can modulate catalytic activities has an enormous therapeutic effect. Hence, inhibition of the HIV protease is one of the most important approaches for the therapeutic intervention in HIV infection and their development is regarded as major success of structure-based drug design. They are highly effective against HIV and have, since the 1990s, been a key component of anti-retroviral therapies for HIV/AIDS.
Leumorphin, also known as dynorphin B1–29, is a naturally occurring endogenous opioid peptide. Derived as a proteolytic cleavage product of residues 226-254 of prodynorphin, leumorphin is a nonacosapeptide and has the sequence Tyr-Gly-Gly-Phe-Leu-Arg-Arg-Gln-Phe-Lys-Val-Val-Thr-Arg-Ser-Gln-Glu-Asp-Pro-Asn-Ala-Tyr-Ser-Gly-Glu-Leu-Phe-Asp-Ala. It can be further reduced to dynorphin B and dynorphin B-14 by pitrilysin metallopeptidase 1, an enzyme of the endopeptidase family. Leumorphin behaves as a potent and selective κ-opioid receptor agonist, similarly to other endogenous opioid peptide derivatives of prodynorphin.
Alpha-lytic endopeptidase or Alpha-lytic protease is an enzyme isolated from the myxobacterium Lysobacter enzymogenes. This enzyme is a serine protease that catalyses the breakage of peptide bonds using a hydrolysis chemical reaction. Alpha-lytic protease was named based on the observed cleavage specificity for the α position of the tetrapeptide component in gram-positive bacterial cell walls (alanine). Alpha-lytic protease is also capable of digesting elastin and other proteins.
Pestivirus NS3 polyprotein peptidase is an enzyme. This enzyme catalyses the following chemical reaction
Calicivirin is an enzyme. This enzyme catalyses the following chemical reaction
HIV-2 retropepsin is an enzyme. This enzyme catalyses the following chemical reaction
Human endogenous retrovirus K (HERV-K) or Human teratocarcinoma-derived virus (HDTV) is a family of human endogenous retroviruses associated with malignant tumors of the testes. Phylogenetically, the HERV-K group belongs to the ERV2 or Class II or Betaretrovirus-like supergroup. Over the past several years, it has been found that this group of ERVs play an important role in embryogenesis, but their expression is silenced in most cell types in healthy adults. The HERV-K family, and particularly its subgroup HML-2, is the youngest and most transcriptionally active group and hence, it is the best studied among other ERVs. Reactivation of it or anomalous expression of HML-2 in adult tissues has been associated with various types of cancer and with neurodegenerative diseases such as amytrophic lateral sclerosis (ALS). Endogenous retrovirus K (HERV-K) is related to mammary tumor virus in mice. It exists in the human and cercopithecoid genomes. Human genome contains hundreds of copies of HERV-K and many of them possess complete open reading frames (ORFs) that are transcribed and translated, especially in early embryogenesis and in malignancies. HERV-K is also found in apes and Old World monkeys. It is uncertain how long ago in primate evolution the full-length HERV-K proviruses which are in the human genome today were created.
Human Endogenous Retrovirus-W (HERV-W) is the coding for a protein that would normally be part of the envelope of one family of Human Endogenous Retro-Viruses, or HERVs.







