
Hexosaminidase is an enzyme involved in the hydrolysis of terminal N-acetyl-D-hexosamine residues in N-acetyl-β-D-hexosaminides.

In enzymology, an aspartate-semialdehyde dehydrogenase is an enzyme that is very important in the biosynthesis of amino acids in prokaryotes, fungi, and some higher plants. It forms an early branch point in the metabolic pathway forming lysine, methionine, leucine and isoleucine from aspartate. This pathway also produces diaminopimelate which plays an essential role in bacterial cell wall formation. There is particular interest in ASADH as disabling this enzyme proves fatal to the organism giving rise to the possibility of a new class of antibiotics, fungicides, and herbicides aimed at inhibiting it.
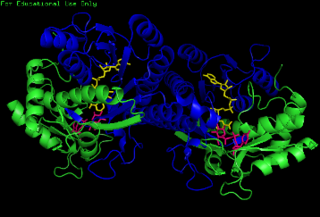
The enzyme UDP-glucose 4-epimerase, also known as UDP-galactose 4-epimerase or GALE, is a homodimeric epimerase found in bacterial, fungal, plant, and mammalian cells. This enzyme performs the final step in the Leloir pathway of galactose metabolism, catalyzing the reversible conversion of UDP-galactose to UDP-glucose. GALE tightly binds nicotinamide adenine dinucleotide (NAD+), a co-factor required for catalytic activity.
In enzymology, an aspartate—tRNA ligase is an enzyme that catalyzes the chemical reaction
Carnosine synthase is an enzyme that catalyzes the chemical reaction

In enzymology, a D-alanine—D-alanine ligase is an enzyme that catalyzes the chemical reaction

In enzymology, a dihydrofolate synthase is an enzyme that catalyzes the chemical reaction

In enzymology, a formate—tetrahydrofolate ligase is an enzyme that catalyzes the chemical reaction
In enzymology, a glutamate-putrescine ligase is an enzyme that catalyzes the chemical reaction
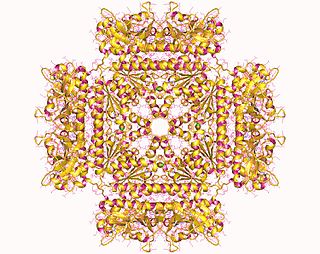

In molecular biology, the protein domain SAICAR synthase is an enzyme which catalyses a reaction to create SAICAR. In enzymology, this enzyme is also known as phosphoribosylaminoimidazolesuccinocarboxamide synthase. It is an enzyme that catalyzes the chemical reaction
In enzymology, a tetrahydrofolate synthase is an enzyme that catalyzes the chemical reaction
In enzymology, a UDP-N-acetylmuramate—L-alanine ligase is an enzyme that catalyzes the chemical reaction
In enzymology, a UDP-N-acetylmuramoyl-L-alanine—D-glutamate ligase is an enzyme that catalyzes the chemical reaction
In enzymology, a UDP-N-acetylmuramoyl-L-alanyl-D-glutamate—L-lysine ligase is an enzyme that catalyzes the chemical reaction
In enzymology, a UDP-N-acetylmuramoyl-tripeptide—D-alanyl-D-alanine ligase is an enzyme that catalyzes the chemical reaction
Peptidoglycan glycosyltransferase is an enzyme used in the biosynthesis of peptidoglycan. It transfers a disaccharide-peptide from a donor substrate to synthesize a glycan chain.
In enzymology, an undecaprenyldiphospho-muramoylpentapeptide beta-N-acetylglucosaminyltransferase is an enzyme that catalyzes the chemical reaction
In enzymology, a phospho-N-acetylmuramoyl-pentapeptide-transferase is an enzyme that catalyzes the chemical reaction
The bacterial cell wall provides strength and rigidity to counteract internal osmotic pressure, and protection against the environment. The peptidoglycan layer gives the cell wall its strength, and helps maintain the overall shape of the cell. The basic peptidoglycan structure of both Gram-positive and Gram-negative bacteria comprises a sheet of glycan chains connected by short cross-linking polypeptides. Biosynthesis of peptidoglycan is a multi-step process comprising three main stages:
- formation of UDP-N-acetylmuramic acid (UDPMurNAc) from N-acetylglucosamine (GlcNAc).
- addition of a short polypeptide chain to the UDPMurNAc.
- addition of a second GlcNAc to the disaccharide-pentapeptide building block and transport of this unit through the cytoplasmic membrane and incorporation into the growing peptidoglycan layer.
UDP-N-acetylmuramoyl-L-alanyl-D-glutamate—D-lysine ligase is an enzyme with systematic name UDP-N-acetylmuramoyl-L-alanyl-D-glutamate:D-lysine alpha-ligase (ADP-forming). This enzyme catalyses the following chemical reaction







