In molecular biology, biosynthesis is a multi-step, enzyme-catalyzed process where substrates are converted into more complex products in living organisms. In biosynthesis, simple compounds are modified, converted into other compounds, or joined to form macromolecules. This process often consists of metabolic pathways. Some of these biosynthetic pathways are located within a single cellular organelle, while others involve enzymes that are located within multiple cellular organelles. Examples of these biosynthetic pathways include the production of lipid membrane components and nucleotides. Biosynthesis is usually synonymous with anabolism.
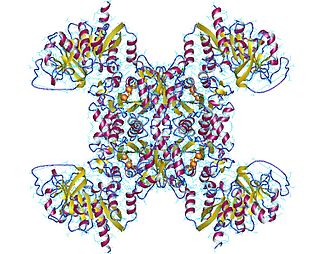
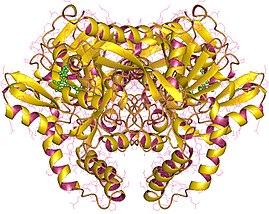
CTP synthase is an enzyme involved in pyrimidine biosynthesis that interconverts UTP and CTP.

The acetolactate synthase (ALS) enzyme is a protein found in plants and micro-organisms. ALS catalyzes the first step in the synthesis of the branched-chain amino acids.
Purine metabolism refers to the metabolic pathways to synthesize and break down purines that are present in many organisms.

Guanosine monophosphate synthetase, also known as GMPS is an enzyme that converts xanthosine monophosphate to guanosine monophosphate.
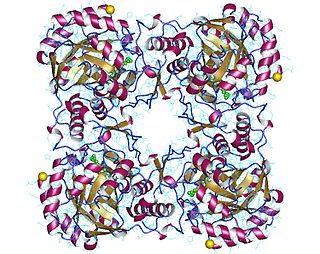
GMP reductase EC 1.7.1.7 is an enzyme that catalyzes the irreversible and NADPH-dependent reductive deamination of GMP into IMP.
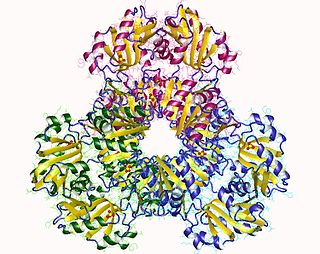
Ribose-phosphate diphosphokinase is an enzyme that converts ribose 5-phosphate into phosphoribosyl pyrophosphate (PRPP). It is classified under EC 2.7.6.1.

In enzymology, a shikimate dehydrogenase (EC 1.1.1.25) is an enzyme that catalyzes the chemical reaction

12-oxophytodienoate reductase (OPRs) is an enzyme of the family of Old Yellow Enzymes (OYE). OPRs are grouped into two groups: OPRI and OPRII – the second group is the focus of this article, as the function of the first group is unknown, but is the subject of current research. The OPR enzyme utilizes the cofactor flavin mononucleotide (FMN) and catalyzes the following reaction in the jasmonic acid synthesis pathway:

Cystathionine beta-lyase, also commonly referred to as CBL or β-cystathionase, is an enzyme that primarily catalyzes the following α,β-elimination reaction
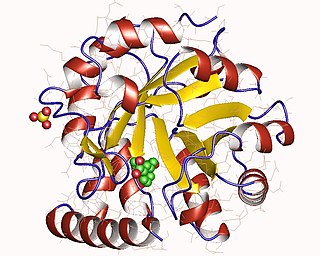
The enzyme indole-3-glycerol-phosphate synthase (IGPS) (EC 4.1.1.48) catalyzes the chemical reaction

In enzymology, an aminodeoxychorismate synthase is an enzyme that catalyzes the chemical reaction

In enzymology, a phenylalanine—tRNA ligase is an enzyme that catalyzes the chemical reaction
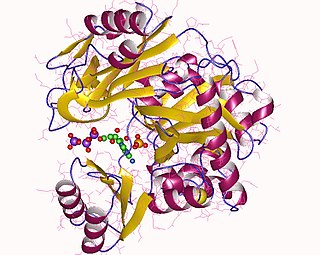
Phosphoribosylamine—glycine ligase, also known as glycinamide ribonucleotide synthetase (GARS), (EC 6.3.4.13) is an enzyme that catalyzes the chemical reaction
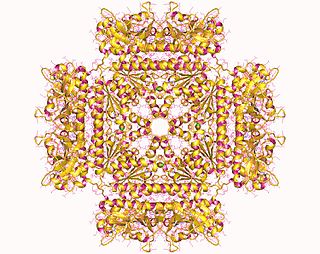

In molecular biology, the protein domain SAICAR synthase is an enzyme which catalyses a reaction to create SAICAR. In enzymology, this enzyme is also known as phosphoribosylaminoimidazolesuccinocarboxamide synthase. It is an enzyme that catalyzes the chemical reaction
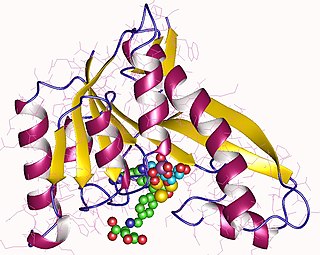
Phosphoribosylglycinamide formyltransferase (EC 2.1.2.2, 2-amino-N-ribosylacetamide 5'-phosphate transformylase, GAR formyltransferase, GAR transformylase, glycinamide ribonucleotide transformylase, GAR TFase, 5,10-methenyltetrahydrofolate:2-amino-N-ribosylacetamide ribonucleotide transformylase) is an enzyme with systematic name 10-formyltetrahydrofolate:5'-phosphoribosylglycinamide N-formyltransferase. This enzyme catalyses the following chemical reaction

S-Adenosylmethionine synthetase, also known as methionine adenosyltransferase (MAT), is an enzyme that creates S-adenosylmethionine by reacting methionine and ATP.

In molecular biology, the amino acid kinase domain is a protein domain. It is found in protein kinases with various specificities, including the aspartate, glutamate and uridylate kinase families. In prokaryotes and plants the synthesis of the essential amino acids lysine and threonine is predominantly regulated by feed-back inhibition of aspartate kinase (AK) and dihydrodipicolinate synthase (DHPS). In Escherichia coli, thrA, metLM, and lysC encode aspartokinase isozymes that show feedback inhibition by threonine, methionine, and lysine, respectively. The lysine-sensitive isoenzyme of aspartate kinase from spinach leaves has a subunit composition of 4 large and 4 small subunits.

The prokaryotic riboflavin biosynthesis protein is a bifunctional enzyme found in bacteria that catalyzes the phosphorylation of riboflavin into flavin mononucleotide (FMN) and the adenylylation of FMN into flavin adenine dinucleotide (FAD). It consists of a C-terminal riboflavin kinase and an N-terminal FMN-adenylyltransferase. This bacterial protein is functionally similar to the monofunctional riboflavin kinases and FMN-adenylyltransferases of eukaryotic organisms, but only the riboflavin kinases are structurally homologous.
The gua operon is responsible for regulating the synthesis of guanosine mono phosphate (GMP), a purine nucleotide, from inosine monophosphate. It consists of two structural genes guaB (encodes for IMP dehydrogenase or and guaA apart from the promoter and operator region.